Abstract
Knowledge of connectivity in the nervous system is essential to understanding its function. Here we describe connectomes for both adult sexes of the nematode Caenorhabditis elegans, an important model organism for neuroscience research. We present quantitative connectivity matrices that encompass all connections from sensory input to end-organ output across the entire animal, information that is necessary to model behaviour. Serial electron microscopy reconstructions that are based on the analysis of both new and previously published electron micrographs update previous results and include data on the male head. The nervous system differs between sexes at multiple levels. Several sex-shared neurons that function in circuits for sexual behaviour are sexually dimorphic in structure and connectivity. Inputs from sex-specific circuitry to central circuitry reveal points at which sexual and non-sexual pathways converge. In sex-shared central pathways, a substantial number of connections differ in strength between the sexes. Quantitative connectomes that include all connections serve as the basis for understanding how complex, adaptive behavior is generated.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The datasets generated and/or analysed during the current study are available at http://wormwiring.org/. The collection of C. elegans nervous system electron micrographs are also available at https://www.wormatlas.org/ and https://wormimage.org. The datasets generated and/or analysed during the current study are also available from the corresponding author on reasonable request.
References
White, J. G., Southgate, E., Thomson, J. N. & Brenner, S. The structure of the nervous system of the nematode Caenorhabditis elegans. Phil. Trans. R. Soc. Lond. B 314, 1–340 (1986).
Albertson, D. G. & Thomson, J. N. The pharynx of Caenorhabditis elegans. Phil. Trans. R. Soc. Lond. B 275, 299–325 (1976).
Jarrell, T. A. et al. The connectome of a decision-making neural network. Science 337, 437–444 (2012).
Hall, D. H. & Russell, R. L. The posterior nervous system of the nematode Caenorhabditis elegans: serial reconstruction of identified neurons and complete pattern of synaptic interactions. J. Neurosci. 11, 1–22 (1991).
Bumbarger, D. J., Riebesell, M., Rödelsperger, C. & Sommer, R. J. System-wide rewiring underlies behavioral differences in predatory and bacterial-feeding nematodes. Cell 152, 109–119 (2013).
Helmstaedter, M. et al. Connectomic reconstruction of the inner plexiform layer in the mouse retina. Nature 500, 168–174 (2013).
Bock, D. D. et al. Network anatomy and in vivo physiology of visual cortical neurons. Nature 471, 177–182 (2011).
Briggman, K. L., Helmstaedter, M. & Denk, W. Wiring specificity in the direction-selectivity circuit of the retina. Nature 471, 183–188 (2011).
Kasthuri, N. et al. Saturated reconstruction of a volume of neocortex. Cell 162, 648–661 (2015).
Lee, W.-C. A. et al. Anatomy and function of an excitatory network in the visual cortex. Nature 532, 370–374 (2016).
Ohyama, T. et al. A multilevel multimodal circuit enhances action selection in Drosophila. Nature 520, 633–639 (2015).
Fushiki, A. et al. A circuit mechanism for the propagation of waves of muscle contraction in Drosophila. eLife 5, e13253 (2016).
Takemura, S. Y. et al. The comprehensive connectome of a neural substrate for ‘ON’ motion detection in Drosophila. eLife 6, e24394 (2017).
Eichler, K. et al. The complete connectome of a learning and memory centre in an insect brain. Nature 548, 175–182 (2017).
Larderet, I. et al. Organization of the Drosophila larval visual circuit. eLife 6, e28387 (2017).
Randel, N. et al. Neuronal connectome of a sensory-motor circuit for visual navigation. eLife 3, e02730 (2014).
Ryan, K., Lu, Z. & Meinertzhagen, I. A. The CNS connectome of a tadpole larva of Ciona intestinalis (L.) highlights sidedness in the brain of a chordate sibling. eLife 5, e16962 (2016).
Varshney, L. R., Chen, B. L., Paniagua, E., Hall, D. H. & Chklovskii, D. B. Structural properties of the Caenorhabditis elegans neuronal network. PLOS Comput. Biol. 7, e1001066 (2011).
Ahn, Y.-Y., Jeong, H. & Kim, B. J. Wiring cost in the organization of a biological neuronal network. Physica A 367, 531–537 (2006).
Chen, B. L., Hall, D. H. & Chklovskii, D. B. Wiring optimization can relate neuronal structure and function. Proc. Natl Acad. Sci. USA 103, 4723–4728 (2006).
Wang, I. E. & Clandinin, T. R. The influence of wiring economy on nervous system evolution. Curr. Biol. 26, R1101–R1108 (2016).
Gushchin, A. & Tang, A. Total wiring length minimization of C. elegans neural network: a constrained optimization approach. PLoS ONE 10, e0145029 (2015).
Carmel, L., Harel, D. & Koren, Y. Combining hierarchy and energy for drawing directed graphs. IEEE Trans. Vis. Comput. Graph. 10, 46–57 (2004).
Gray, J. M., Hill, J. J. & Bargmann, C. I. A circuit for navigation in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 102, 3184–3191 (2005).
Zou, W. et al. Polymodal responses in C. elegans phasmid neurons rely on multiple intracellular and intercellular signaling pathways. Sci. Rep. 7, 42295 (2017).
Hart, A. C., Kass, J., Shapiro, J. E. & Kaplan, J. M. Distinct signaling pathways mediate touch and osmosensory responses in a polymodal sensory neuron. J. Neurosci. 19, 1952–1958 (1999).
Pereira, L. et al. A cellular and regulatory map of the cholinergic nervous system of C. elegans. eLife 4, e12432 (2015).
Zhao, H. & Nonet, M. L. A retrograde signal is involved in activity-dependent remodeling at a C. elegans neuromuscular junction. Development 127, 1253–1266 (2000).
Kratsios, P. et al. Transcriptional coordination of synaptogenesis and neurotransmitter signaling. Curr. Biol. 25, 1282–1295 (2015).
Schwarz, J. & Bringmann, H. Analysis of the NK2 homeobox gene ceh-24 reveals sublateral motor neuron control of left–right turning during sleep. eLife 6, e24846 (2017).
Stephens, G. J., Bueno de Mesquita, M., Ryu, W. S. & Bialek, W. Emergence of long timescales and stereotyped behaviors in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 108, 7286–7289 (2011).
Stephens, G. J., Johnson-Kerner, B., Bialek, W. & Ryu, W. S. Dimensionality and dynamics in the behavior of C. elegans. PLOS Comput. Biol. 4, e1000028 (2008).
Kato, S. et al. Global brain dynamics embed the motor command sequence of Caenorhabditis elegans. Cell 163, 656–669 (2015).
Pierce-Shimomura, J. T., Faumont, S., Gaston, M. R., Pearson, B. J. & Lockery, S. R. The homeobox gene lim-6 is required for distinct chemosensory representations in C. elegans. Nature 410, 694–698 (2001).
Johnston, R. J. Jr, Chang, S., Etchberger, J. F., Ortiz, C. O. & Hobert, O. MicroRNAs acting in a double-negative feedback loop to control a neuronal cell fate decision. Proc. Natl Acad. Sci. USA 102, 12449–12454 (2005).
Emmons, S. W. Neural circuits of sexual behavior in Caenorhabditis elegans. Annu. Rev. Neurosci. 41, 349–369 (2018).
Sherlekar, A. L. et al. The C. elegans male exercises directional control during mating through cholinergic regulation of sex-shared command interneurons. PLoS ONE 8, e60597 (2013).
Koo, P. K., Bian, X., Sherlekar, A. L., Bunkers, M. R. & Lints, R. The robustness of Caenorhabditis elegans male mating behavior depends on the distributed properties of ray sensory neurons and their output through core and male-specific targets. J. Neurosci. 31, 7497–7510 (2011).
Ryan, D. A. et al. Sex, age, and hunger regulate behavioral prioritization through dynamic modulation of chemoreceptor expression. Curr. Biol. 24, 2509–2517 (2014).
Hilbert, Z. A. & Kim, D. H. Sexually dimorphic control of gene expression in sensory neurons regulates decision-making behavior in C. elegans. eLife 6, e21166 (2017).
Serrano-Saiz, E. et al. A neurotransmitter atlas of the Caenorhabditis elegans male nervous system reveals sexually dimorphic neurotransmitter usage. Genetics 206, 1251–1269 (2017).
Oren-Suissa, M., Bayer, E. A. & Hobert, O. Sex-specific pruning of neuronal synapses in Caenorhabditis elegans. Nature 533, 206–211 (2016).
Barrios, A., Ghosh, R., Fang, C., Emmons, S. W. & Barr, M. M. PDF-1 neuropeptide signaling modulates a neural circuit for mate-searching behavior in C. elegans. Nat. Neurosci. 15, 1675–1682 (2012).
Jang, H. et al. Neuromodulatory state and sex specify alternative behaviors through antagonistic synaptic pathways in C. elegans. Neuron 75, 585–592 (2012).
Hart, M. P. & Hobert, O. Neurexin controls plasticity of a mature, sexually dimorphic neuron. Nature 553, 165–170 (2018).
Toth, M. L. et al. Neurite sprouting and synapse deterioration in the aging Caenorhabditis elegans nervous system. J. Neurosci. 32, 8778–8790 (2012).
Stern, S., Kirst, C. & Bargmann, C. I. Neuromodulatory control of long-term behavioral patterns and individuality across development. Cell 171, 1649–1662 (2017).
Goodman, M. B., Hall, D. H., Avery, L. & Lockery, S. R. Active currents regulate sensitivity and dynamic range in C. elegans neurons. Neuron 20, 763–772 (1998).
Hendricks, M., Ha, H., Maffey, N. & Zhang, Y. Compartmentalized calcium dynamics in a C. elegans interneuron encode head movement. Nature 487, 99–103 (2012).
LeBoeuf, B. & Garcia, L. R. Caenorhabditis elegans male copulation circuitry incorporates sex-shared defecation components to promote intromission and sperm transfer. G3 (Bethesda) 7, 647–662 (2017).
Chalfie, M. et al. The neural circuit for touch sensitivity in Caenorhabditis elegans. J. Neurosci. 5, 956–964 (1985).
Jee, C., Goncalves, J. F., LeBoeuf, B. & Garcia, L. R. CRF-like receptor SEB-3 in sex-common interneurons potentiates stress handling and reproductive drive in C. elegans. Nat. Commun. 7, 11957 (2016).
Hardaker, L. A., Singer, E., Kerr, R., Zhou, G. & Schafer, W. R. Serotonin modulates locomotory behavior and coordinates egg-laying and movement in Caenorhabditis elegans. J. Neurobiol. 49, 303–313 (2001).
Garrison, J. L. et al. Oxytocin/vasopressin-related peptides have an ancient role in reproductive behavior. Science 338, 540–543 (2012).
Barrios, A., Nurrish, S. & Emmons, S. W. Sensory regulation of C. elegans male mate-searching behavior. Curr. Biol. 18, 1865–1871 (2008).
Sammut, M. et al. Glia-derived neurons are required for sex-specific learning in C. elegans. Nature 526, 385–390 (2015).
Sakai, N. et al. A sexually conditioned switch of chemosensory behavior in C. elegans. PLoS ONE 8, e68676 (2013).
Narayan, A. et al. Contrasting responses within a single neuron class enable sex-specific attraction in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 113, E1392–E1401 (2016).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Sulston, J. E., Albertson, D. G. & Thomson, J. N. The Caenorhabditis elegans male: postembryonic development of nongonadal structures. Dev. Biol. 78, 542–576 (1980).
Peachey, L. D. Thin sections. I. A study of section thickness and physical distortion produced during microtomy. J. Biophys. Biochem. Cytol. 4, 233–242 (1958).
Xu, M. et al. Computer assisted assembly of connectomes from electron micrographs: application to Caenorhabditis elegans. PLoS ONE 8, e54050 (2013).
Duerr, J. S., Gaskin, J. & Rand, J. B. Identified neurons in C. elegans coexpress vesicular transporters for acetylcholine and monoamines. Am. J. Physiol. Cell Physiol. 280, C1616–C1622 (2001).
Desbois, M., Cook, S. J., Emmons, S. W. & Bülow, H. E. Directional trans-synaptic labeling of specific neuronal connections in live animals. Genetics 200, 697–705 (2015).
Acknowledgements
We thank A. Barrios and B. Kim for comments on the manuscript; J. Koehler for comments on the statistical analysis; J. Hodgkin and J. White for their help in lending the electron microscopy archives from MRC/LMB for long-term curation at the Hall laboratory. M. Bernstein tested the fertility of males for the N930 reconstruction. M. Xu computationally aligned EM images. C. Crocker created the interactive version of Fig. 1 and formatted figures throughout. This work was supported by NIH grants from NINDS (F31NS096863 to E.A.B.; R01NS096672 to H.E.B.; R37NS039996 to O.H.), NIHD (P30HD071593 to S.W.E.), NIMH (R01MH112689 to S.W.E.), NIGMS (T32GM00749 1 to S.J.C.; R01GM066897 to S.W.E.), NINCDS (R15N548916 to J.S.D) and the Office of the NIH Director (OD 010943 to D.H.H.), and by the G. Harold and Leila Y. Mathers Charitable Foundation (S.W.E.). L.T.-H.T. is a Croucher Foundation Research Fellow. H.E.B. is an Irma T. Hirschl/Monique Weill-Caullier Research Fellow. O.H. is an investigator of the Howard Hughes Medical Institute. Some nematode strains were provided by the CGC, which is funded by NIH Office of Research Infrastructure Programs (P40 0D010440).
Reviewer information
Nature thanks Albert-László Barabási, Dan Bumbarger, Douglas Portman, Paul W. Sternberg, Emma Towlson and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
S.J.C., T.A.J., C.A.B., Y.W., A.E.B. and M.A.Y. generated and analysed the data, including performing electron microscopy experiments, annotating electron micrographs, compilation of data and curation of neuron maps; A.E.B. performed the statistical analysis for left versus right and hermaphrodite versus male comparisons; K.C.Q.N. carried out sample preparation and serial sectioning for electron microscopy; S.J.C., L.T.-H.T. and E.A.B. performed the experiments involving fluorescent labelling of synapses; J.S.D. carried out analysis of acetylcholine expression in the sublateral cords; H.E.B. and O.H. conceived and supervised the experiments involving fluorescent labelling of synapses; D.H.H. and S.W.E. conceived, planned and supervised the experiments; all co-authors contributed to preparation of the manuscript; S.W.E. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Comparison of previous reconstructions to this work.
Many more synapses are scored primarily because the reconstruction method using Elegance enabled the marking of synapses on images, facilitating an exhaustive annotation (Methods). a, Maps of AIML and AIMR from reconstruction of the hermaphrodite series N2U as presented previously1 (left) and the present study (right). The map from the present reconstruction, as given at http://wormwiring.org/, has been redrawn to have the same projection as the previous study (laying out the portion that extends around the nerve ring). Furthermore, as in the previous study, only the target cells at polyadic chemical synapses at which AIML and AIMR are presynaptic are shown. However, co-recipients at synapses in which AIML or AIMR are postsynaptic may be obtained by moving over the synapse on the online map at http://wormwiring.org/. The small branches on the skeleton backbone are not neurite branches or spines, but rather are locations where a protrusion or bulging out of the neurite resulted in EM profiles separated from the main profile for a few sections. In the scoring method, all profiles are marked by a point object. The skeleton map is created by joining these objects, resulting in the short extensions from the backbone at the bulges. b, Comparison of the number of edges in the adjacency matrices and the number of synapses scored in the present and previous studies1,18. c, The edges in the present study that are not present in the previous study are mostly small.
Extended Data Fig. 2 Properties of synapses.
a, Electron micrographs of a representative chemical synapse (left) and a gap junction synapse (right), marked by an asterisk. The triadic chemical synapse of AIYR to RIAR, AIZR, and RIBR (synapse number N2U391) is recognized by the presynaptic density and the presence of presynaptic vesicles within AIYR. The postsynaptic cells are entered into the database in a clockwise order around the presynaptic density. As electron micrographs give a consistent cross-sectional view from the posterior, the location of the neuronal processes is oriented in the worm. The gap junction (synapse number N2U3875) is recognized as a straightened or slightly curving region of apposed membranes with increased staining and a uniform small gap. Below the micrographs is shown how each of these two synapses is listed in the table of synapses (Supplementary Information 3). b, Number of postsynaptic partners at chemical synapses. The lower number of polyads in the male data is probably not a true difference between the sexes, but rather a result of the less-complete nature of the male reconstruction, which was due to the lower quality of the electron micrographs. c, Size distributions of synapses. d, Larger edges are composed of larger synapses as well as more synapses.
Extended Data Fig. 3 Properties of the graphs.
a, Distributions of weights for chemical (left) and gap junction (right) edges (number of serial EM sections). b, Degree distributions. c, Distribution of the presumptive load (cumulative physical sizes of the edges) as a function of edge weight. A large fraction of the load is carried by the many small edges. d, The fraction of connectivity due to gap junctions for the classes of interneurons in the hermaphrodite.
Extended Data Fig. 4 Similar arrangement of neurite processes in the two sexes.
The transverse section of the ventral ganglion in the hermaphrodite (N2U series) depicted in figure 16 of the previous study1 is compared to the corresponding section in the male (series N930, section 1100). Some of the processes in the male were not identified.
Extended Data Fig. 5 Output of the algorithm for finding hierarchy in a network.
The previously published algorithm18,23 was used to analyse the hierarchical structure of the nervous system networks (Fig. 2). Hierarchical position is shown on the y axis; the adjacency of neurons (roughly, anterior to left, posterior to right) is shown on the x axis. For Fig. 2, some small adjustments to the node positions were made, primarily in the horizontal direction, to clarify the data by removing overlaps. Two adjustments were made in the vertical direction to make neuron positions reflect the preponderance of their output. RIA was moved down to the level of the other neurons that have a majority of output onto motor neurons and muscles. Notably, 88% of RIA chemical output is onto motor neurons and muscles, making it seem that RIA should be considered one of the premotor interneurons. It is probably placed at a higher level of the hierarchy by the algorithm because of its large number of inputs from sensory neurons (10) and layer 3 interneurons (3). It receives input from only a single layer 2 interneuron, RIB (see below). (It has negligible gap junctional connectivity.) In the second case, RIB was moved up, to the next higher layer (interneuron layer 2). Only 10% of RIB chemical output is onto motor neurons and muscles, whereas 43% is onto layer 1 interneurons (including RIA). In addition, 40% of RIB total output is through gap junctions. Of these, 37% of the load is with motor neurons (possibly influencing its placement by the algorithm), 15% with layer 1 interneurons and 47% with layer 2 and above. These were the only two neuron classes that seemed to be placed by the algorithm at a position that did not well represent the preponderance of their output.
Extended Data Fig. 6 Chemical and gap junction networks.
a–d, The hierarchical networks of Fig. 2 are shown with chemical and gap junction edges separated. e, Left, gap junction output of sensory neuron classes. The touch neurons (SN3) have a higher proportion of gap junctional connectivity than the other classes. Middle, gap junctions are distributed fairly uniformly throughout the network. The category 4 interneurons, which cannot be placed at a particular level in the otherwise hierarchical structure, are also distinguished by making remarkably few gap junctions. (The number of category 4 interneurons is a quarter to a half the number of neurons in each of the other categories.) Right, the fraction of connectivity due to gap junctions is similar throughout the network.
Extended Data Fig. 7 Analysis of doublet and triplet motifs in the somatic nervous system.
The frequency (blue) of doublets and triplets in the unweighted hermaphrodite and male networks compared to 1,000 degree-conserved randomized networks (red crosses). a, b, Hermaphrodite and male gap junction doublets. c, d, Hermaphrodite and male chemical synapse doublets. e, f, Hermaphrodite and male chemical synapse triplets. We determined statistically significant deviations from our randomized data using a MATLAB script based on a previously published study18. We used the single step min p procedure and multiple hypothesis testing; *P < 0.05.
Extended Data Fig. 8 Connectivity in the hermaphrodite to non-muscle end organs.
There is substantial connectivity (>3 EM serial sections) to five cells or organs: the intestine (int), hypodermis (hyp), head mesodermal cell (hmc), CAN cell (a neuron-like cell with unusual properties) and the excretory gland (exc_gl) cell. In the male, DVB and AVL are repurposed to have a role in ejaculation. Many of the neurons that innervate the hypodermis have processes running through the animal in the ventral cord. The head mesodermal cell, a cell of unknown function, has extensive gap junctions to body-wall muscles. This provides an indirect connection between the body-wall muscles and the excretory gland cell.
Extended Data Fig. 9 Innervation of body-wall muscle and definition of motor neurons.
a, Fraction of synaptic output that is onto muscle for the neuron classes that make neuromuscular junctions. With the exception of IL1, classes for which greater than 30% of their chemical output is onto muscle are classified as motor neurons (mn). IL1 neurons—which are classified as sensory neurons (sn)—have clear sensory endings and sensory function; they are sensory–motor neurons, but we do not use that classification here. VC and HSN—hermaphrodite-specific neurons—innervate egg-laying muscles and have traditionally been considered motor neurons. The remaining neurons are classified as interneurons. Neurons that have previously been categorized as interneurons are indicated by a dagger. Neurons that have previously been categorized as motor neurons are indicated by an asterisk. b, Fraction of the input to the body-wall muscles of the head region and the remainder of the body that comes from neurons of various classes.
Extended Data Fig. 10 Five classes of sublateral motor neurons (SAB, SMD, SMB, SIB and SIA) are contained in sublateral nerves and innervate body-wall muscles.
a, Schematic transverse view of anterior midbody to show the relative positions of the four posterior sublateral nerves (large arrows) in relation to the body-wall muscles (green), the lateral nerves (Lat), ventral nerve cord (VC) and dorsal nerve cord (DC). All nerves are shown in red. Anterior to the nerve ring, the anterior sublateral nerves adopt the same relative positions with respect to more anterior body-wall muscles, tending to lie in a cleft at the margins of two adjoining rows of muscles in each quadrant. Intestine (pink), germline tissues (dark blue), pseudocoelom (yellow), hypodermis (tan), excretory canal (purple) and seam (orange). Cartoon is adapted from the WormAtlas. b, Adult head showing Cy3-anti-mouse antibody staining of VAChT (monoclonal antibody1403; secondary antibody from Jackson ImmunoResearch) inside neuronal processes in the nerve ring (NR), dorsal cord, and anterior and posterior sublateral nerves. The members of the sublateral nerves show periodic swellings that are brightly stained, whereas the entire nerve ring and dorsal cord are extremely bright. Lateral view, anterior to the left, dorsal to the top. Scale bar, 100 μm. n > 50 animals. c, d, Transmission electron micrographs from adult wild-type animals obtained as transverse cross-sections. Note that each sublateral nerve contains 4–5 axons lying in an indentation of the outer edge of the body-wall muscle quadrant, close to the cell border between two adjoining muscles. White asterisks mark swollen axons of interest. Synaptic vesicles cluster among electron-dense ground substance near the active zone. Other sublateral axons tend to be very narrow, contain a few microtubules and often a narrow tube of smooth endoplasmic reticulum but no ground substance. Electron-dense material is sometimes seen at the base of the plasma membrane to form a small dark presynaptic bar. Diffuse basal lamina fills the extracellular space between the muscles and the underlying layer of hypodermis and its sublateral nerve. The cuticle is shown along the bottom edge of each panel. c, Adult sublateral nerve in which the axon on the far left forms a presynaptic density (small electron-dense patch is the centre of the active zone) pointing leftward to BWM and hypodermis. Animal N2U, E4, image 014. d, Adult sublateral nerve in which one swollen axon contains vesicles and ground substance, but shows no evidence of a presynaptic density. The thin hypodermal layer is locally swollen to accommodate a mitochondrion just to the left of the nerve. Some swellings show only vesicles and extra ground substance, whereas others include a small presynaptic dense bar that can point towards muscle, hypodermis or other members of the nerve. Animal N501C, image Y382. c, d, Scale bar, 0.5 μm. Transmission electron micrographs of 6 animals show similar synaptic features (J.S.D. et al., manuscript in preparation).
Extended Data Fig. 11 Estimated s.d. as a function of edge weight for the left–right comparison.
Analysis related to data shown in the scatter plots of Fig. 5a. Values are the number of serial EM sections.
Extended Data Fig. 12 Comparison of connectivity across sex using transsynaptic fluorescent labelling.
a–h, Fluorescent micrographs and quantification of immunofluorescence data. a, iBLINC labelling of AFD to AIY synapses. n = 18 male animals and n = 23 hermaphrodite animals, P = 0.99. b, iBLINC labelling of PHA to PHB synapses. n = 14 male animals and n = 15 hermaphrodite animals, P = 0.43. c, iBLINC labelling of URX and AQR to AVA, RIG, and AIY synapses. n = 15 male animals and n = 15 hermaphrodite animals, P = 1.00. d, iBLINC labelling of ASI to AFD synapses. n = 22 male animals and n = 22 hermaphrodite animals, P = 0.0000059. e, GRASP labelling of IL2 to RIB synapses. n = 19 male animals and n = 23 hermaphrodite animals, P = 0.0114. f, iBLINC labelling of IL1 to RIB. n = 17 male animals and n = 26 hermaphrodite animals, P = 0.0000099. g, GRASP labelling of AIB to RIM synapses. n = 20 male animals and n = 29 hermaphrodite animals, P = 0.053. h, GRASP labelling of ASH to AVA. n = 14 male animals and n = 16 hermaphrodite animals, P < 0.0001. For all fluorescence comparisons a two-sided Wilcoxon rank-sum test was used. Each dot represents the number of synaptic puncta in one animal, the middle black line is the median, the box shows the interquartile range, whiskers show the outer quartiles.
Extended Data Fig. 13 Possible sex differences in connectivity between shared neurons are distributed throughout the network.
Separately for the chemical and gap junction networks, the hermaphrodite and male networks for cell classes (Fig. 2) were added together and edges that come mainly from just one sex are highlighted: green, stronger in the hermaphrodite, blue, stronger in the male. Selection of edges to highlight was done as follows. Using the data in Supplementary Information 9, for the chemical matrix the highest 100 z-scores for each sex (out of 1,823 comparisons) were selected; for the gap junction matrix the highest 35 z-scores out of 356 comparisons were selected. Thus approximately 20% of edges are highlighted. Note, differences between the two reconstructions can arise for many reasons. Whether the edges highlighted here are true sex differences needs to be tested by methodology that allows many animals to be examined.
Supplementary information
Supplementary Table 1
Transgenic strains used in this study.
Supplementary Figure
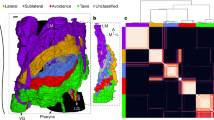
This file contains high-resolution PDFs of the C. elegans adult nervous system, neuroanatomy and connectivity, shown in Fig. 1 (adult hermaphrodite and pharynx on page 1, and adult male on page 2). The worm diagrams show the locations of the neuronal cell nuclei and outlines of the muscle cells (the insets show the sex muscles). The nodes of the network diagrams show the entire set of neurons and end organs (muscles and other tissues such as the gut and skin) and the connections between them—black arrows, chemical connections; red lines, gap junction connections. The node layout was generated algorithmically to reflect the amount of connectivity between the cells. Similarity of the layout to the worm’s neuroanatomy reflects economical wiring. Node shapes show neuron types: triangles, sensory neurons; hexagons, interneurons; ovals or circles, motor neurons; rectangles, muscles. Colours define various categories. Small worm cartoons show the major nerve tracts and ganglia: CG, cloacal ganglion; DRG, dorsorectal ganglion; LG, lumbar ganglion; PAG, pre-anal ganglion; RVG, retrovesicular ganglion; VG, ventral ganglion.
Supplementary Figure
(Interactive) The C. elegans adult nervous system, neuroanatomy and connectome. Interactive version of Fig. 1 with active links for further information. Top diagrams: the major nerve tracts and ganglia (anterior to left). Not shown are lateral nerves containing the processes of three neurons associated with the canal cell and processes of lateral touch neurons. Hermaphrodite (top) and male (bottom) have the same overall layout. The major difference is a larger number of neurons and muscles in the male tail that sub-serve copulation (see insets). The primary centers of complex connectivity are the nerve ring and, in the male, the pre-anal ganglion (PAG). VG, ventral ganglion; RVG, retrovesicular ganglion; PAG, pre-anal ganglion; DRG, dorsorectal ganglion; LG, lumbar ganglion; CG, cloacal ganglion. Lower diagrams: the nervous system—neuroanatomy and network graph. Top, adult hermaphrodite; bottom, adult male. It is possible to magnify the figure to view the details; mouse over or click on nuclei or nodes to relate them and for pop-up labels and links to information about each cell. (After navigating away from the figure to a website, on return click first on the background before closing the box.) The worm diagrams show the locations of cell nuclei (left side and center only, right-side homologues of left–right pairs are not shown). In the graph representations, the layout of the vertices is determined by an algorithm that clusters more-heavily connected cell pairs (AllegroViva, force-directed strong clustering algorithm). The display is by Cytoscape (cytoscape.org). Directed edges (black arrows) represent chemical synapses; undirected edges (red lines) represent gap junction connections. The widths and transparencies of the lines represent the edge weights. Triangles, sensory neurons; hexagons, interneurons; ovals or circles, motor neurons; rectangles, muscles. Colours define various categories: various reds indicate categories of sensory neurons defined by modality and similarity of connectivity (see also Fig. 3); various blues indicate interneuron categories according to their assignment to a layer (or lack of assignment in the case of IN4) (see also Fig. 2); motor neuron classes (various yellows/oranges) are described in the text; non-muscle end organs are white/grey/black. Sex-specific neurons are pink or purple, with numerous additional colours used for the male-specific network in the male tail, delineating the modules described in ref 3.
Supplementary Information 1
Significant updates and corrections to previous data
Supplementary Information 2
Number of scored synapses between pairs of cells: synapse adjacency matrices, hermaphrodite and male, chemical synapses and gap junctions For chemical connections, rows: presynaptic cell; columns: post-synaptic cell. The body of the table gives the number of synapses scored between each pair of cells. NB: This matrix differs from the Connectome Adjacency Matrices (Supplementary Information 5) in not taking into account the sizes of synapses and in showing only connections scored on electron micrographs. It does not include any connections inserted by extrapolation. Thus there are cells showing no connection here that are connected in the Connectome Adjacency Matrices. Also note, there are more edges in this table than the total number of synapses scored (synapse lists, Supplementary Information 3). This is because polyads are listed here more than once (once for each postsynaptic cell).
Supplementary Information 3
List of synapses, hermaphrodite and male, chemical and gap junctional These files give the properties of all the synapses scored. The contin number ("contin" = a connected set of database objects representing a structure that may "continue" across several sections) together with the EM series designation give each synapse a unique identifier. They may be found on a neuron map at WormWiring.org by using this identifier. For chemical polyads, the order of the postsynaptic cells given in column D and H-K is the clockwise order the cells were scored around the presynaptic density on the electron micrographs. As the micrograph images are viewed consistently looking towards anterior, so that left and right sides of the image correspond to left and right sides of the worm, this clockwise ordering orients the postsynaptic arrangement in the worm.
Supplementary Information 4
Cell Lists These sheets list all of the cells in the C. elegans adult nervous system along with end organs as they appear in the connectivity matrices.
Supplementary Information 5
Connectome Adjacency Matrices These sheets give the whole-animal connectomes for the two adult sexes of C. elegans. They provide the amount of connectivity between each pair of cells. For chemical connections, rows give the pre-synaptic cell, columns give the post-synaptic cell. Weights in the body of the matrices are the total number of EM serial sections of connectivity, taking into account both the number of synapses and the sizes of synapses. To provide complete coverage of the entire nervous system, the data are assembled from multiple animals and include connections added by extrapolation in gaps where no data were available. The data in these tables is used in the figures of Cook et al.
Supplementary Information 6
Cell Class Lists These are the cell classes and other cell groups that are used in the figures of Cook et al. Shared neurons are the cell classes previously recognized (White et al., 1986). Muscles and the male-specific complement of the male tail are combined into larger groups.
Supplementary Information 7
Cell Class Adjacency Matrices These are whole-animal connectivity matrices based on the cell classes listed in Supplementary Information 6. The data are the same (number of EM serial sections of connectivity) as that in Supplementary Information 5, except given by cell class (or larger combined group) instead of by individual cell. Connectivity between classes is the sum of the connectivity of the class members given in Supplementary Information 5. Data in these tables is used for the Figures in Cook et al.
Supplementary Information 8
Hermaphrodite Left-Right Comparisons These lists give the amount of connectivity (number of EM sections) for the left member and the right member of a homologous pair of neurons onto a class of neurons that itself consists of a left-right pair (that is, connectivity of A/left onto (B/left + B/right) compared to A/right onto (B/left + B/right). Separate sheets for chemical and gap junction connections. sd: standard deviation. Z score: the difference between connection strengths normalized by its estimated variance.
Supplementary Information 9
Ranked Sex Differences The tables give a comparison of connectivity between cell classes. The statistical approach used is described in Cook et al., Methods. Briefly, the approach was to use the variance in connectivty between left/right homologs in the hermaphrodite data (Supplementary Information 8 and Extended Data Fig. 12) to judge how much variance would be expected between the hermaphrodite and male datasets if there were no sex difference. This is a minimum, as it does not take into account possible inter-individual differences not based on sex but on other factors (such as age, experience, and so forth).
Supplementary Information 10
Properties of the EM series used in this study.
Rights and permissions
About this article
Cite this article
Cook, S.J., Jarrell, T.A., Brittin, C.A. et al. Whole-animal connectomes of both Caenorhabditis elegans sexes. Nature 571, 63–71 (2019). https://doi.org/10.1038/s41586-019-1352-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1352-7
This article is cited by
-
A full-body transcription factor expression atlas with completely resolved cell identities in C. elegans
Nature Communications (2024)
-
Comparative connectomics of dauer reveals developmental plasticity
Nature Communications (2024)
-
Molecular and circuit mechanisms underlying avoidance of rapid cooling stimuli in C. elegans
Nature Communications (2024)
-
Distinct neurexin isoforms cooperate to initiate and maintain foraging activity
Translational Psychiatry (2023)
-
Neural engineering with photons as synaptic transmitters
Nature Methods (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.