Abstract
A widely held—but rarely tested—hypothesis for the origin of animals is that they evolved from a unicellular ancestor, with an apical cilium surrounded by a microvillar collar, that structurally resembled modern sponge choanocytes and choanoflagellates1,2,3,4. Here we test this view of animal origins by comparing the transcriptomes, fates and behaviours of the three primary sponge cell types—choanocytes, pluripotent mesenchymal archaeocytes and epithelial pinacocytes—with choanoflagellates and other unicellular holozoans. Unexpectedly, we find that the transcriptome of sponge choanocytes is the least similar to the transcriptomes of choanoflagellates and is significantly enriched in genes unique to either animals or sponges alone. By contrast, pluripotent archaeocytes upregulate genes that control cell proliferation and gene expression, as in other metazoan stem cells and in the proliferating stages of two unicellular holozoans, including a colonial choanoflagellate. Choanocytes in the sponge Amphimedon queenslandica exist in a transient metastable state and readily transdifferentiate into archaeocytes, which can differentiate into a range of other cell types. These sponge cell-type conversions are similar to the temporal cell-state changes that occur in unicellular holozoans5. Together, these analyses argue against homology of sponge choanocytes and choanoflagellates, and the view that the first multicellular animals were simple balls of cells with limited capacity to differentiate. Instead, our results are consistent with the first animal cell being able to transition between multiple states in a manner similar to modern transdifferentiating and stem cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All cell-type transcriptome data are available in the NCBI SRA database under accession number PRJNA412708. Additional supplementary data are available from the Dryad Digital Respository: https://doi.org/10.5061/dryad.hp2fr73.
References
Cavalier-Smith, T. Origin of animal multicellularity: precursors, causes, consequences-the choanoflagellate/sponge transition, neurogenesis and the Cambrian explosion. Phil. Trans. R. Soc. Lond. B 372, 20150476 (2017).
Brunet, T. & King, N. The origin of animal multicellularity and cell differentiation. Dev. Cell 43, 124–140 (2017).
Arendt, D., Benito-Gutierrez, E., Brunet, T. & Marlow, H. Gastric pouches and the mucociliary sole: setting the stage for nervous system evolution. Phil. Trans. R. Soc. Lond. B 370, 20150286 (2015).
Nielsen, C. Six major steps in animal evolution: are we derived sponge larvae? Evol. Dev. 10, 241–257 (2008).
Sebé-Pedrós, A., Degnan, B. M. & Ruiz-Trillo, I. The origin of Metazoa: a unicellular perspective. Nat. Rev. Genet. 18, 498–512 (2017).
Sebé-Pedrós, A. et al. The dynamic regulatory genome of Capsaspora and the origin of animal multicellularity. Cell 165, 1224–1237 (2016).
Gaiti, F. et al. Landscape of histone modifications in a sponge reveals the origin of animal cis-regulatory complexity. eLife 6, e22194 (2017).
Gaiti, F., Calcino, A. D., Tanurdžić, M. & Degnan, B. M. Origin and evolution of the metazoan non-coding regulatory genome. Dev. Biol. 427, 193–202 (2017).
Babonis, L. S. & Martindale, M. Q. Phylogenetic evidence for the modular evolution of metazoan signalling pathways. Phil. Trans. R. Soc. Lond. B 372, 20150477 (2017).
Fairclough, S. R. et al. Premetazoan genome evolution and the regulation of cell differentiation in the choanoflagellate Salpingoeca rosetta. Genome Biol. 14, R15 (2013).
Sebé-Pedrós, A. et al. Regulated aggregative multicellularity in a close unicellular relative of metazoa. eLife 2, e01287 (2013).
de Mendoza, A., Suga, H., Permanyer, J., Irimia, M. & Ruiz-Trillo, I. Complex transcriptional regulation and independent evolution of fungal-like traits in a relative of animals. eLife 4, e08904 (2015).
Arendt, D. et al. The origin and evolution of cell types. Nat. Rev. Genet. 17, 744–757 (2016).
Maldonado, M. Choanoflagellates, choanocytes, and animal multicellularity. Invertebr. Biol. 123, 1–22 (2004).
Ereskovsky, A. The Comparative Embryology of Sponges (Springer, 2010).
Nakanishi, N., Sogabe, S. & Degnan, B. M. Evolutionary origin of gastrulation: insights from sponge development. BMC Biol. 12, 26 (2014).
Hashimshony, T. et al. CEL-Seq2: sensitive highly-multiplexed single-cell RNA-Seq. Genome Biol. 17, 77 (2016).
Fernandez-Valverde, S. L., Calcino, A. D. & Degnan, B. M. Deep developmental transcriptome sequencing uncovers numerous new genes and enhances gene annotation in the sponge Amphimedon queenslandica. BMC Genomics 16, 387 (2015).
Lê Cao, K. A., Boitard, S. & Besse, P. Sparse PLS discriminant analysis: biologically relevant feature selection and graphical displays for multiclass problems. BMC Bioinformatics 12, 253 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Domazet-Lošo, T. & Tautz, D. A phylogenetically based transcriptome age index mirrors ontogenetic divergence patterns. Nature 468, 815–818 (2010).
Li, L., Stoeckert, C. J., Jr & Roos, D. S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 13, 2178–2189 (2003).
Fagnocchi, L. & Zippo, A. Multiple roles of MYC in integrating regulatory networks of pluripotent stem cells. Front. Cell Dev. Biol. 5, 7 (2017).
Young, S. L. et al. Premetazoan ancestry of the Myc–Max network. Mol. Biol. Evol. 28, 2961–2971 (2011).
Kress, T. R., Sabò, A. & Amati, B. MYC: connecting selective transcriptional control to global RNA production. Nat. Rev. Cancer 15, 593–607 (2015).
Sogabe, S., Nakanishi, N. & Degnan, B. M. The ontogeny of choanocyte chambers during metamorphosis in the demosponge Amphimedon queenslandica. Evodevo 7, 6 (2016).
Mah, J. L., Christensen-Dalsgaard, K. K. & Leys, S. P. Choanoflagellate and choanocyte collar-flagellar systems and the assumption of homology. Evol. Dev. 16, 25–37 (2014).
Pozdnyakov, I., Sokolova, A., Ereskovsky, A. & Karpov, S. Kinetid structure of choanoflagellates and choanocytes of sponges does not support their close relationship. Protistology 11, 248–264 (2017).
Srivastava, M. et al. The Amphimedon queenslandica genome and the evolution of animal complexity. Nature 466, 720–726 (2010).
Levin, M. et al. The mid-developmental transition and the evolution of animal body plans. Nature 531, 637–641 (2016).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2009).
Kolde, R. pheatmap v.1.0.8 https://cran.r-project.org/package=pheatmap (2012).
Neuwirth, E. RColorBrewer v.1.1-2 https://cran.r-project.org/package=RColorBrewer (2011).
Conesa, A. et al. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676 (2005).
Götz, S. et al. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 36, 3420–3435 (2008).
Kanehisa, M., Sato, Y. & Morishima, K. BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J. Mol. Biol. 428, 726–731 (2016).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44 (D1), D457–D462 (2016).
Rohart, F., Gautier, B., Singh, A. & Lê Cao, K.-A. mixOmics: An R package for ‘omics feature selection and multiple data integration. PLoS Comput. Biol. 13, e1005752 (2017).
Aguilera, F., McDougall, C. & Degnan, B. M. Co-Option and de novo gene evolution underlie molluscan shell diversity. Mol. Biol. Evol. 34, 779–792 (2017).
Domazet-Lošo, T., Brajković, J. & Tautz, D. A phylostratigraphy approach to uncover the genomic history of major adaptations in metazoan lineages. Trends Genet. 23, 533–539 (2007).
Shen, L. GeneOverlap: an R package to test and visualize gene overlaps http://shenlab-sinai.github.io/shenlab-sinai/ (2014).
Wattam, A. R. et al. PATRIC, the bacterial bioinformatics database and analysis resource. Nucleic Acids Res. 42, D581–D591 (2014).
Yates, A. et al. Ensembl 2016. Nucleic Acids Res. 44 (D1), D710–D716 (2016).
Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408, 796–815 (2000).
Ruiz-Trillo, I., Lane, C. E., Archibald, J. M. & Roger, A. J. Insights into the evolutionary origin and genome architecture of the unicellular opisthokonts Capsaspora owczarzaki and Sphaeroforma arctica. J. Eukaryot. Microbiol. 53, 379–384 (2006).
Suga, H. et al. The Capsaspora genome reveals a complex unicellular prehistory of animals. Nat. Commun. 4, 2325 (2013).
King, N. et al. The genome of the choanoflagellate Monosiga brevicollis and the origin of metazoans. Nature 451, 783–788 (2008).
Wilson, D., Charoensawan, V., Kummerfeld, S. K. & Teichmann, S. A. DBD-taxonomically broad transcription factor predictions: new content and functionality. Nucleic Acids Res. 36, D88–D92 (2008).
Srivastava, M. et al. Early evolution of the LIM homeobox gene family. BMC Biol. 8, 4 (2010).
Larroux, C. et al. Genesis and expansion of metazoan transcription factor gene classes. Mol. Biol. Evol. 25, 980–996 (2008).
Larroux, C. et al. Developmental expression of transcription factor genes in a demosponge: insights into the origin of metazoan multicellularity. Evol. Dev. 8, 150–173 (2006).
Shimeld, S. M., Degnan, B. & Luke, G. N. Evolutionary genomics of the Fox genes: origin of gene families and the ancestry of gene clusters. Genomics 95, 256–260 (2010).
Layden, M. J., Meyer, N. P., Pang, K., Seaver, E. C. & Martindale, M. Q. Expression and phylogenetic analysis of the zic gene family in the evolution and development of metazoans. Evodevo 1, 12 (2010).
Presnell, J. S., Schnitzler, C. E. & Browne, W. E. KLF/SP transcription factor family evolution: Expansion, diversification, and innovation in eukaryotes. Genome Biol. Evol. 7, 2289–2309 (2015).
Mukhopadhyay, S. & Jackson, P. K. The tubby family proteins. Genome Biol. 12, 225 (2011).
Larroux, C. et al. The NK homeobox gene cluster predates the origin of Hox genes. Curr. Biol. 17, 706–710 (2007).
Wang, L., Tang, Y., Cole, P. A. & Marmorstein, R. Structure and chemistry of the p300/CBP and Rtt109 histone acetyltransferases: implications for histone acetyltransferase evolution and function. Curr. Opin. Struct. Biol. 18, 741–747 (2008).
Petroni, K. et al. The promiscuous life of plant NUCLEAR FACTOR Y transcription factors. Plant Cell 24, 4777–4792 (2012).
Morrison, A. J. & Shen, X. Chromatin remodelling beyond transcription: the INO80 and SWR1 complexes. Nat. Rev. Mol. Cell Biol. 10, 373–384 (2009).
Jones, M. H., Hamana, N., Nezu, Ji. & Shimane, M. A novel family of bromodomain genes. Genomics 63, 40–45 (2000).
Song, W., Solimeo, H., Rupert, R. A., Yadav, N. S. & Zhu, Q. Functional dissection of a Rice Dr1/DrAp1 transcriptional repression complex. Plant Cell 14, 181–195 (2002).
Matheos, D. P., Kingsbury, T. J., Ahsan, U. S. & Cunningham, K. W. Tcn1p/Crz1p, a calcineurin-dependent transcription factor that differentially regulates gene expression in Saccharomyces cerevisiae. Genes Dev. 11, 3445–3458 (1997).
Rivera, A. S. et al. Gene duplication and the origins of morphological complexity in pancrustacean eyes, a genomic approach. BMC Evol. Biol. 10, 123 (2010).
Romanovskaya, E. V. et al. Transcription factors of the NF1 family: Possible mechanisms of inducible gene expression in the evolutionary lineage of multicellular animals. J. Evol. Biochem. Physiol. 53, 85–92 (2017).
Leys, S. P. et al. Isolation of Amphimedon developmental material. Cold Spring Harb. Protoc. 2008, prot5095 (2008).
Degnan, B. M. et al. Evolutionary Developmental Biology of Invertebrates, vol. 1 (Springer, 2015).
Larroux, C. et al. Whole-mount in situ hybridization in Amphimedon. Cold Spring Harb. Protoc. 2008, prot5096 (2008).
Acknowledgements
This study was supported by funds from the Australian Research Council (B.M.D. and S.M.D.). We thank I. Ruiz-Trillo for primary expression data for Capsaspora and Creolimax and N. Rhodes for assistance with computing and database management.
Author information
Authors and Affiliations
Contributions
B.M.D. and S.M.D. conceived and designed the project. S.S., D.S. and K.E.R. identified and isolated the cells and prepared the libraries. W.L.H., S.S. and K.M.K. performed gene expression and annotation and phylostratigraphic analyses with help from T.E.S., S.M.D, S.L.F.-V. and B.M.D. S.S. performed cell-lineage analyses. B.M.D., S.M.D. and S.S. wrote the manuscript with comments and contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
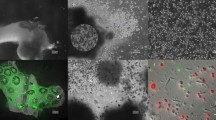
Extended Data Fig. 1 A. queenslandica cell types and sPLS-DA of choanocyte, archaeocyte and pinacocyte transcriptomes.
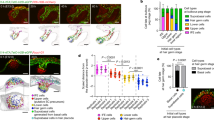
a, Whole-mount internal view of a juvenile A. queenslandica. Cell types are outlined. A, archaeocyte (cluster of four outlined); Cc, choanocyte chamber; S, sclerocyte; Sp, spherulous cell; P, pinacocyte. b, Choanocyte chamber labelled with DiI with an illustration of a single choanocyte below. c, Pinacocyte labelled with DiI with illustration below. d, Archaeocyte labelled with DiI with illustration below. Scale bars, 10 µm (b), 5 µm (c, d). e–i, sPLS-DA identified the gene models that best characterize differences in choanocytes (blue, n = 10), archaeocytes (red, n = 15) and pinacocytes (green, n = 6). e, Sample plot for the optimal number of gene models that discriminate cell types on the first two components; ellipses indicate 95% confidence intervals. f, g, Hierarchically clustered heat maps show the expression of the 110 gene models selected for the first component (f) and the 98 gene models and 2 long non-coding RNAs selected for the second component (g), which accounted for 15% and 5% of explained variance, respectively. h, i, Venn diagrams summarize the significantly differentially expressed genes identified by the DESeq2 analyses for each cell type and the sPLS-DA on the first (h) and the second (i) sPLS-DA component. Percentages are of the total number of differentially expressed genes identified from all analyses.
Extended Data Fig. 2 Percentage of KEGG cellular processes and environmental information processing (that is, cell signalling) genes present in each cell type, corresponding to the number of components that make up each KEGG category identified.
a, Cellular processes genes. b, Environmental information processing (that is, cell signalling) genes.
Extended Data Fig. 3 Evolutionary age of genes expressed in A. queenslandica choanocytes, archaeocytes and pinacocytes using different expression thresholds.
a–e, Phylostratigraphic enrichment of genes expressed in each cell type (Ar, archaeocyte; Ch, choanocyte; Pi, pinacocyte; ArCh, archaeocyte + choanocyte; ArPi, archaeocyte + pinacocyte; ChPi, choanocyte + pinacocyte; ALL, all three cell types combined) at different expression thresholds. Expressed genes are parsed into quartiles based on transcript abundance in each of the cell types. Quartile 1 (Q1) includes the least abundant transcripts and Q4 the most abundant. a, Phylostratigraphy enrichment of all genes expressed in each of the cell types (Q1–Q4). b, Phylostratigraphy enrichment of genes expressed in the top three quartiles (excluding Q1). c, Phylostratigraphy enrichment of genes expressed in the top 50% (Q3 and Q4). d, Phylostratigraphy enrichment of the most highly expressed genes (Q4). e, For comparison, the evolutionary age of differentially expressed genes identified using differential expression analysis, DESeq2. Heat maps indicate enrichment (log-odds ratio based on a two-sided Fisher’s exact test) of phylostrata contained in each gene list in comparison to the A. queenslandica genome (n = 44,719). Asterisks mark significant (P < 0.05; Fisher’s exact test) overlap between gene lists, indicative of phylostrata enrichment. The heat maps on the far right are collapsed versions of the heat maps on the left, in which the premetazoan category contains phylostrata from cellular to holozoan, and the poriferan category contains phylostrata from poriferan to A. queenslandica. To the left of each heat map is a Venn diagram, showing the number of genes in each cell type and set of cell types. Grey boxes on the heat map indicate that there were no genes in that particular gene list characterized by the given phylostrata. See additional supplementary data on Dryad: /ED_Fig. 3_files and /Fig. 3e. f, Pairwise comparison illustrating the number of overlapping genes for each of the quartiles between the three cell types. The numbers in the cells are the number of genes common between two cell types (for example, there are 1,569 expressed genes in common between Q2 in choanocytes and Q3 in archaeocytes). NE, not expressed. g, The percentage of differentially upregulated genes identified in each of the cell types using DESeq2 in the four quartiles.
Extended Data Fig. 4 Orthologues shared between cell-type-specific gene lists and non-metazoan eukaryotes.
Heat map showing the percentage of A. queenslandica genes with orthogroups shared with select eukaryotes. a, Percentage of genes with orthogroups shared between upregulated and total expressed genes from non-exclusive lists (that is, all genes expressed in each of the three cell types, not excluding genes that overlap between any two cell types). b, Percentage of genes with orthogroups shared between DEG and total expressed genes-exclusive lists (that is, genes uniquely upregulated or expressed in that cell type).
Extended Data Fig. 5 Orthologues found in S. rosetta, C. owczarzaki and C. fragrantissima life-cycle stages, shared with A. queenslandica cell-type transcriptomes and eukaryotic genomes.
a, The percentage and number (in parentheses) of differentially expressed orthogroups found in S. rosetta, C. owczarzaki and C. fragrantissima life-cycle stages that are shared with A. queenslandica cell types. The numbers in parentheses alongside the unicellular holozoan cell states and sponge cell-type names are the total numbers of orthogroups differentially expressed in that specific gene list. b, A heat map showing the percentage of orthogroups shared between genes differentially expressed in S. rosetta, C. owczarzaki and C. fragrantissima life-cycle stages, and genes present in other eukaryotic genomes.
Extended Data Fig. 6 Heat map of transcription factor genes differentially expressed in choanocytes, archaeocytes and pinacocytes.
Ninety-four transcription factor genes that are differentially expressed in A. queenslandica cell types are classified on the basis of phylostratum: premetazoan (light grey), metazoan (dark grey) and poriferan (black). a, Heat map of expression levels in the three cell types combining all analysed CEL-Seq2 data. Depicted values illustrate scaled (z-score) expression levels based on collapsed VST from 10 choanocyte, 15 archaeocyte and 6 pinacocyte transcriptomes. Gene names, families (in parentheses) and phylostrata shading are shown on the right. b, Heat map of uncollapsed expression levels (VST) of all transcriptomes (10 choanocyte, 15 archaeocyte and 6 pinacocyte). Rows in b correspond to the rows and genes in a. c, Venn diagram summary of differentially upregulated transcription factor genes between the three cell types using DESeq2. Percentages are of the total transcription factor genes differentially upregulated in all cell types. d, Bar graph of the number and distribution of transcription factor genes based on evolutionary age in the three cell types.
Extended Data Fig. 7 Analysis of premetazoan transcription factors in A. queenslandica cells and unicellular holozoan cell states.
a, The number and percentage of premetazoan transcription factor orthologues that are present in the genomes of S. rosetta, C. owczarzaki and C. fragrantissima. Percentages are based on the 43 premetazoan genes differentially expressed in the A. queenslandica cell types (Extended Data Fig. 5). The number of transcription factor orthologues in the genome is listed above the bar. The orange bar depicts the percent and number of unicellular holozoan premetazoan transcription factor orthologues that are significantly differentially upregulated in at least one cell state. b, The 15 premetazoan transcription factor orthology groups (listed along the top) that are significantly upregulated in at least one A. queenslandica cell type and one unicellular holozoan cell state. Dots correspond to the cell types and states this occurs. Black dots, orthology group with one gene member; grey dots, orthology group comprising two or more paralogues (see Supplementary File 8 for details).
Extended Data Fig. 8 Choanocyte dedifferentiation into an archaeocyte does not require cell division.
a, b, Four-day-old juveniles 6 h after CM-DiI and EdU labelling. a, CM-DiI-labelled archaeocytes with EdU incorporation (arrows) found near choanocyte chambers. b, Labelled archaeocytes without EdU incorporation (arrowheads), indicating dedifferentiation from choanocytes without cell division. c, d, Choanocyte-derived archaeocytes are capable of generating new choanocyte chambers. c, Four-day-old juvenile 6 h after CM-DiI and EdU labelling. Early choanocyte chamber (dotted line) completely labelled with CM-DiI and EdU, indicating that CM-DiI-labelled archaeocytes with large nuclei are forming this chamber. The absence of cilia and space at the centre of this structure indicates it is not yet a functional choanocyte chamber. d, Four-day-old juvenile 12 h after CM-DiI and 6 h after EdU labelling. Early choanocyte chamber (dotted line) with multiple EdU labelled cells, with both CM-DiI-labelled choanocytes (arrowheads) and non-CM-DiI-labelled choanocytes (arrows) indicate multiple cell lineages contributing to the formation of this chamber. The images presented in a–d represent the consensus of cell behaviours obtained from 10 independent labelling experiments, each comprising a minimum of 24 juveniles. Scale bars, 10 μm.
Supplementary information
Supplementary Table
Supplementary Table 1: The counting report of the cell type specific CEL-Seq2 samples.
Supplementary Table
Supplementary Table 2: Table summarising the details and the statistics of the demultiplexing and mapping steps for the cell type specific CEL-Seq2 samples.
Supplementary Table
Supplementary Table 3: Table of differentially expressed gene lists from DESeq2 with BLAST2GO annotations and phylostrata ID.
Supplementary Table
Supplementary Table 4: Table of differentially expressed gene lists from sPLS-DA with BLAST2GO and KEGG annotations and phylostrata ID.
Supplementary Table
Supplementary Table 5: Table of KEGG enrichment analysis results on differentially expressed gene lists from DESeq2. This spreadsheet contains the output of KEGG enrichment analyses performed on each cell type DEG list. The first sheet contains percentage values of genes/components identified in the DEG lists relative to the A. queenslandica genome.
Supplementary Table
Supplementary Table 6: Table of the phylostrata enrichment of the differentially expressed gene lists from DESeq2.
Supplementary Table
Supplementary Table 7: Table of cell-type gene lists and transcription factor lists from the quartile expression analyses.
Supplementary Table
Supplementary Table 8: Table of transcription factor genes expressed in the three cell types and in the differentially expressed gene lists from DESeq2.
Supplementary Video 1
: Time-lapse video of a 4 day old juvenile Amphimedon queenslandica. This 10 second video captures 20 minutes of cell behavior on the outer edge of the juvenile. Annotated are (i) a choanocyte chamber (cc) comprising of multiple tethered choanocytes, (ii) three migrating archaeocytes (ar) – there are multiple other archaeocytes in this video, and (iii) a pinacocyte (pi), which comes in and out of focus and is characterised by a thin, transparent cytoplasm with small refractive vesicles. Scale bar, 10 µm. n=10 biologically independent samples.
Supplementary Video 2
: Capture and washing of a dissociated archaeocyte. All cells and choanocyte chambers used in this study were fixed or frozen in less than 15 minutes after dissociation from the intact sponge. n=174 biologically independent samples.
Supplementary Video 3
: Time-lapse video of choanocytes transdifferentiating and evacuating chambers in 4 day old juvenile Amphimedon queenslandica. Matching 8-second videos captures 120 min of CM-DiI labelled choanocytes (left, red; right, white), which are initially located in distinct chambers (arrows on four chambers), undergoing transdifferentiation and migrating from the chambers. At the end of the video, none of the CM-DiI labelled chambers are recognisable. Note that many cells vacating the choanocyte chambers are larger, consistent with choanocytes dedifferentiating into larger archaeocytes. Scale bar, 40 µm. n=5 biologically independent samples.
Rights and permissions
About this article
Cite this article
Sogabe, S., Hatleberg, W.L., Kocot, K.M. et al. Pluripotency and the origin of animal multicellularity. Nature 570, 519–522 (2019). https://doi.org/10.1038/s41586-019-1290-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1290-4
This article is cited by
-
Germline-related molecular phenotype in Metazoa: conservation and innovation highlighted by comparative transcriptomics
EvoDevo (2023)
-
Sex-specific expression of pheromones and other signals in gravid starfish
BMC Biology (2022)
-
Neural is Fundamental: Neural Stemness as the Ground State of Cell Tumorigenicity and Differentiation Potential
Stem Cell Reviews and Reports (2022)
-
Evolution of mechanisms controlling epithelial morphogenesis across animals: new insights from dissociation-reaggregation experiments in the sponge Oscarella lobularis
BMC Ecology and Evolution (2021)
-
Insights into the origin of metazoan multicellularity from predatory unicellular relatives of animals
BMC Biology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.