Abstract
Tuberculosis is the leading cause of death by an infectious disease worldwide1. However, the involvement of innate lymphoid cells (ILCs) in immune responses to infection with Mycobacterium tuberculosis (Mtb) is unknown. Here we show that circulating subsets of ILCs are depleted from the blood of participants with pulmonary tuberculosis and restored upon treatment. Tuberculosis increased accumulation of ILC subsets in the human lung, coinciding with a robust transcriptional response to infection, including a role in orchestrating the recruitment of immune subsets. Using mouse models, we show that group 3 ILCs (ILC3s) accumulated rapidly in Mtb-infected lungs and coincided with the accumulation of alveolar macrophages. Notably, mice that lacked ILC3s exhibited a reduction in the accumulation of early alveolar macrophages and decreased Mtb control. We show that the C-X-C motif chemokine receptor 5 (CXCR5)–C-X-C motif chemokine ligand 13 (CXCL13) axis is involved in Mtb control, as infection upregulates CXCR5 on circulating ILC3s and increases plasma levels of its ligand, CXCL13, in humans. Moreover, interleukin-23-dependent expansion of ILC3s in mice and production of interleukin-17 and interleukin-22 were found to be critical inducers of lung CXCL13, early innate immunity and the formation of protective lymphoid follicles within granulomas. Thus, we demonstrate an early protective role for ILC3s in immunity to Mtb infection.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All relevant data are available from the corresponding authors upon reasonable request. RNA-sequencing data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) repository, accession number GSE131031.
Change history
24 July 2019
An Amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
World Health Organization. Global Tuberculosis Report 2018. (2018).
Klose, C. S. & Artis, D. Innate lymphoid cells as regulators of immunity, inflammation and tissue homeostasis. Nat. Immunol. 17, 765–774 (2016).
Diefenbach, A., Colonna, M. & Koyasu, S. Development, differentiation, and diversity of innate lymphoid cells. Immunity 41, 354–365 (2014).
Takatori, H. et al. Lymphoid tissue inducer-like cells are an innate source of IL-17 and IL-22. J. Exp. Med. 206, 35–41 (2009).
Jones, G. W. & Jones, S. A. Ectopic lymphoid follicles: inducible centres for generating antigen-specific immune responses within tissues. Immunology 147, 141–151 (2016).
Lim, A. I. et al. Systemic human ILC precursors provide a substrate for tissue ILC differentiation. Cell 168, 1086–1100 (2017).
Monticelli, L. A. et al. Innate lymphoid cells promote lung-tissue homeostasis after infection with influenza virus. Nat. Immunol. 12, 1045–1054 (2011).
Mchedlidze, T. et al. Interleukin-33-dependent innate lymphoid cells mediate hepatic fibrosis. Immunity 39, 357–371 (2013).
Kløverpris, H. N. et al. Innate lymphoid cells are depleted irreversibly during acute HIV-1 infection in the absence of viral suppression. Immunity 44, 391–405 (2016).
Hedfors, I. A. & Brinchmann, J. E. Long-term proliferation and survival of in vitro-activated T cells is dependent on interleukin-2 receptor signalling but not on the high-affinity IL-2R. Scand. J. Immunol. 58, 522–532 (2003).
Slight, S. R. et al. CXCR5+ T helper cells mediate protective immunity against tuberculosis. J. Clin. Invest. 123, 712–726 (2013).
Constantinides, M. G., McDonald, B. D., Verhoef, P. A. & Bendelac, A. A committed precursor to innate lymphoid cells. Nature 508, 397–401 (2014).
Moro, K. et al. Innate production of TH2 cytokines by adipose tissue-associated c-Kit+Sca-1+ lymphoid cells. Nature 463, 540–544 (2010).
Dorhoi, A. et al. The adaptor molecule CARD9 is essential for tuberculosis control. J. Exp. Med. 207, 777–792 (2010).
Traber, K. E. et al. Induction of STAT3-dependent CXCL5 expression and neutrophil recruitment by oncostatin-M during pneumonia. Am. J. Respir. Cell Mol. Biol. 53, 479–488 (2015).
Nouailles, G. et al. CXCL5-secreting pulmonary epithelial cells drive destructive neutrophilic inflammation in tuberculosis. J. Clin. Invest. 124, 1268–1282 (2014).
Pagán, A. J. et al. Myeloid growth factors promote resistance to mycobacterial infection by curtailing granuloma necrosis through macrophage replenishment. Cell Host Microbe 18, 15–26 (2015).
van de Veerdonk, F. L. et al. Mycobacterium tuberculosis induces IL-17A responses through TLR4 and dectin-1 and is critically dependent on endogenous IL-1. J. Leukoc. Biol. 88, 227–232 (2010).
Yadav, M. & Schorey, J. S. The β-glucan receptor dectin-1 functions together with TLR2 to mediate macrophage activation by mycobacteria. Blood 108, 3168–3175 (2006).
El-Shazly, A. E. et al. Novel association between vasoactive intestinal peptide and CRTH2 receptor in recruiting eosinophils: a possible biochemical mechanism for allergic eosinophilic inflammation of the airways. J. Biol. Chem. 288, 1374–1384 (2013).
Rajaram, M. V. et al. Mycobacterium tuberculosis activates human macrophage peroxisome proliferator-activated receptor γ linking mannose receptor recognition to regulation of immune responses. J. Immunol. 185, 929–942 (2010).
Tientcheu, L. D. et al. Differential transcriptomic and metabolic profiles of M. africanum- and M. tuberculosis-infected patients after, but not before, drug treatment. Genes Immun. 16, 347–355 (2015).
O’Kane, C. M., Elkington, P. T. & Friedland, J. S. Monocyte-dependent oncostatin M and TNF-α synergize to stimulate unopposed matrix metalloproteinase-1/3 secretion from human lung fibroblasts in tuberculosis. Eur. J. Immunol. 38, 1321–1330 (2008).
Khader, S. A. et al. IL-23 and IL-17 in the establishment of protective pulmonary CD4+ T cell responses after vaccination and during Mycobacterium tuberculosis challenge. Nat. Immunol. 8, 369–377 (2007).
Ebihara, T. et al. Runx3 specifies lineage commitment of innate lymphoid cells. Nat. Immunol. 16, 1124–1133 (2015).
Cupedo, T. et al. Human fetal lymphoid tissue-inducer cells are interleukin 17-producing precursors to RORC+ CD127+ natural killer-like cells. Nat. Immunol. 10, 66–74 (2009).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Li, B. D. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Bindea, G. et al. ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25, 1091–1093 (2009).
Upreti, D., Pathak, A. & Kung, S. K. Development of a standardized flow cytometric method to conduct longitudinal analyses of intracellular CD3ζ expression in patients with head and neck cancer. Oncol. Lett. 11, 2199–2206 (2016).
Acknowledgements
This work was supported by Washington University in St Louis, NIH grant HL105427, AI111914-02 and AI123780 to S.A.K. and D.K., AI134236-02 to S.A.K., M.C. and D.K., and NIH/NHLBI T32 HL007317-37 to R.D.-G., the Department of Molecular Microbiology, Washington University in St Louis, and Stephen I. Morse Fellowship to S.D., T32 HL 7317-39 to N.C.H. and T32-AI007172 to M. Dunlap. A.L. was supported by BMGF (OPP1137006) and the Wellcome Trust (210662/Z/18/Z), A. Singh and T.N. were supported by the Sub-Saharan African Network for TB/HIV Research Excellence (SANTHE), a DELTAS Africa Initiative (DEL-15-006). J.R.-M. was supported by funds of the Department of Medicine, University of Rochester, and NIH grant U19 AI91036. A.K.S. was supported, in part, by the Searle Scholars Program, the Beckman Young Investigator Program, a Sloan Fellowship in Chemistry, the NIH (5U24AI118672), the Bill and Melinda Gates Foundation and the Ragon Institute. S.W.K. was supported by an NSF Graduate Student Fellowship Award and the Hugh Hampton Young Memorial Fund Fellowship. We thank Amgen for providing the anti-IL-23 antibody for the study, J. Bando for helping with the flow cytometry and M. Holtzman for gifting Il13−/− mice.
Reviewer information
Nature thanks Gérard Eberl, Tom H. M. Ottenhoff and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
A.A., R.D.-G., S.D., S.W.K., N.C.H., A. Singh, M.A., S.N., J.R.-M., P.O., L.L., D.R., M.d.l.G.-H., T.K.U., M. Darby, E.P., F.K., L.M., R.M., K.J.D., M. Dunlap and N.M.-A. designed, performed and interpreted experiments, T.E., T.N., D.K., A.S.P., J.K.K., A. Steyn, J.Z., W.H., W.M.Y., A.K.S., H.N.K., M.C., A.L. and S.A.K. interpreted experiments, carried out data analysis and/or provided reagents. H.N.K., A.L., M.C. and S.A.K. designed the study, provided funding and wrote the paper, all authors edited and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
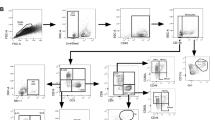
Extended Data Fig. 1 Hierarchical gating strategy used to identify lymphocyte populations in human blood and lung samples.
Single cells from blood (n = 68) or lung samples (n = 42) from human participants were processed for flow cytometry, and all doublets were excluded. Cells were gated as lymphocytes, live, CD45+ and CD3+ T cells or CD3− cells. CD3− cells were gated on CD56 and CD94. CD94+ cells are NK cells and were further subgated as CD16−CD56high NK cells or CD16+CD56low NK cells. ILCs in the CD94− fraction were CD127+ and negative for all lineage markers: CD4, CD11c, CD14, CD19, CD34, FcER1, BDCA2, TCRαβ and TCRγδ. Total ILCs were CD127+CD161+, ILC2s were Lin−CD127+CRTH2+ cells. ILC1s were Lin−CD127+CRTH2−CD56−CD117− cells. ILC3 were Lin−CD127+CRTH2−CD117+ cells with variable CD56 expression.
Extended Data Fig. 2 ILC depletion seen in participants with tuberculosis is not affected by tuberculosis drug resistance or concurrent infection with HIV.
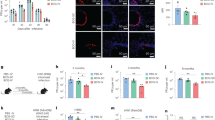
a, The frequencies of the two main circulating NK populations, CD16+CD56low and CD16−CD56high were measured in human participants with tuberculosis (n = 35–45) and healthy controls (n = 19) by flow cytometry. NK cell frequencies in paired samples taken from the same participant with tuberculosis before and after 6 months of standard and successful tuberculosis therapy were also determined by flow cytometry (n = 8–11). b, Percentages of blood ILC1s, ILC2s, ILC3s, CD56low NK cells and CD56high NK cells were measured in control subjects who did not have tuberculosis (TB) or HIV (TB−HIV−, n = 19), participants with tuberculosis and without HIV (TB+HIV−, n = 13–15) and with tuberculosis and HIV coinfection (TB+HIV+, n = 12–15), and participants with multi-drug-resistant tuberculosis without HIV (MDRTB+HIV−, n = 4–8) and with multi-drug-resistant tuberculosis with HIV coinfection (MDRTB+HIV+, n = 7–12). Significance is calculated by a Dunn’s multiple comparison test. *P < 0.05, **P < 0.01. c, Capase-3 expression in circulating lymphocytes from peripheral blood of participants with tuberculosis (n = 14–16) and controls (n = 15–24) was done by flow cytometry. Significance calculated using a Mann–Whitney U-test. Data are mean ± s.d and individual values.
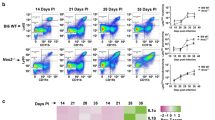
Extended Data Fig. 3 Hierarchical gating strategy used to identify ILC populations in mouse lungs.
a–c, C57BL/6 mice were aerosol infected with approximately 100 CFU Mtb and lungs were collected at different d.p.i. and flow cytometry analysis was carried out on single-cell suspensions. a, b, Flow gating strategies for ILC1s (CD45+CD127+Lin−NKp46+NK1.1+), ILC2s (CD45+CD127+Lin−NK1.1−Sca1+) and ILC3 (CD45+CD127+Lin−NK1.1−Rorγt+) and NKp46-expressing (CD45+CD127+Lin−NK1.1−Rorγt+NKp46+) ILC3s are shown. c, Rorγt–GFP mice were aerosol-infected with approximately 100 CFU Mtb and lungs were collected at 14 d.p.i. ILC3 (CD45+CD127+Lin−NK1.1−GFP+) populations were quantified using flow cytometry.
Extended Data Fig. 4 Pulmonary ILCs are tissue-resident cells and express markers of migration.
a, CD69, CD103, CD62L and NK-p44 expression on the circulating ILCs in human peripheral blood and lung tissue were measured by flow cytometry. Percentage of total human ILCs expressing these markers in paired samples of participants with tuberculosis shown (paired samples n = 5; NK-p44 expression unpaired samples, lung n = 26; blood n = 34). Significance of CD69, CD103 and CD62L expression was calculated using a one-way Wilcoxon matched-pairs test. Significance of NK-p44 expression was analysed using an unpaired Mann–Whitney U-test. b, c, NK-p44 and CD56 expression were measured in tuberculosis-infected lung tissues in comparison to control samples. Significance is analysed using a unpaired Mann–Whitney U-test (b) and a Kruskal–Wallis test with adjustments for multiple comparisons (c). d, Percentages of ILC1, ILC2, ILC3, CD56low NK cells, and CD56high NK cells in human lung tissue were measured by flow cytometry TB−HIV− control subjects (n = 4–6), participants with tuberculosis without HIV (TB+HIV−, n = 7–13) and with tuberculosis with HIV coinfection (TB+HIV+, n = 7–22). e, CXCL13 protein levels were measured in the plasma of participants with tuberculosis without (n = 18–20) and with HIV coinfection(n = 9). Significance is calculated by Mann–Whitney U-test (no significance after Bonferonni correction). f, Frequencies of CD103+ ILCs were measured by flow cytometry in the blood of TB−HIV− control subjects (n = 24), participants with tuberculosis without (n = 10) and with HIV coinfection (n = 5). Significance is analysed using a Mann–Whitney U-test with Bonferroni corrections (only values that were significant after correction are shown). g, Representative FACS plots showing two distinct subpopulations of CD103- and CXCR5-expressing ILCs measured in lung tissues from three subjects with tuberculosis; in these populations most CXCR5-expressing cells are CD117+ ILC3s and CD103+ lung ILCs are a combination of CD117+ ILC3s, CRTH2+ ILCs and CD117−CRTH2− cells. Green, CD117+; red, CRTH2+. h, C57BL/6 mice were aerosol-infected with approximately 100 CFU Mtb and lungs were collected at 14 d.p.i. Lung ILCs were sorted from single-cell suspensions (ILCs: CD45+CD127+Lin–NK1.1−).The ability of sorted ILCs to migrate towards medium alone or a gradient of mouse CXCL13 was quantified in a transwell migration assay. n = 3–5 biological replicates. Significance is calculated by one-way ANOVA, *P < 0.05, **P < 0.01. i, Human ILC3s sorted from lungs migrated in response to recombinant human CXCL13 in transwell migration assays. Significance is calculated by one-tailed Wilcoxon signed-rank test. Data are paired values (a, left three graphs and i, right) or mean ± s.d. and individual values (b–f, h).
Extended Data Fig. 5 ILC3s localize within lymphoid follicles during tuberculosis.
a, IHC staining of nuclear RORγt, CD3 and PAX5 on FFPE lung tissues from human participants with pulmonary tuberculosis or influenza (left) or macaques with latent tuberculosis infections and pulmonary tuberculosis (right). b, Representative fluorescent immunohistology scans of tuberculosis-infected human lung tissues, with CD20 (FITC), CD3 (PE–Texas Red) and CD127 (PE–Cy5). CD3−CD127+ ILCs are present adjacent to follicles (top) and granuloma-like structures (bottom). c, Cumulative total ILCs per mm2 of lung tissue (left), in histological sections from 3 different patients with tuberculosis, are increased in structures of tuberculosis histopathlogy (combined lesional tissue) in comparison to the remainder of unaffected tissue (non-lesional). However, percentages of ILCs per total cell number (DAPI+ cells) (right) are not different between regions of interest (lesional tissue, n = 31 lesions) and unaffected tissue (non-lesional, n = 3 patients). d, e, ILC3 immunofluorescence analysis of nuclear RORγt, B220 and CD3 in FFPE lung tissues from human participants with pulmonary tuberculosis (d) and macaques (e) with pulmonary tuberculosis (PTB) and latent tuberculosis infections (LTBI).
Extended Data Fig. 6 Sort purity of human ILC3s and CD4+ T cells.
ILC3s and CD4+ T cells were sorted from human PBMCs and reflowed back into FACSAria Fusion to confirm purity. Purity of ILC3s was confirmed to be 100% and the CD4+ T cell sort was 97% pure.
Extended Data Fig. 7 VIP and OSM are expressed within human lung tissue infected with tuberculosis.
a, b, VIP (a) and OSM (b) protein expression was confirmed in situ in human lung tissues infected with tuberculosis using multiplexed fluorescent immunohistology. c, The network of upstream drivers enriched in the differentially expressed genes in sorted ILC3s between tuberculosis-infected and uninfected samples are shown. Inset, GO network generated over the genes identified as downstream of OSM by IPA (n = 64, see Methods). Each node represents a specific GO, KEGG or Reactome term (Supplementary Table 4). Broad categories of pathways are annotated. Line width and darkness correspond to number of shared genes between nodes. Node size, **P < 0.01; ***P < 0.001. d, Select predicted downstream pathways enriched in the differentially expressed genes in ILC3s between tuberculosis-infected and uninfected samples are shown.
Extended Data Fig. 8 Sorting purity of mouse ILCs.
a, C57BL/6 mice were aerosol-infected with approximately 100 CFU Mtb and lungs were collected at 5 d.p.i. The lung CD45 population was enriched by using CD45 microbeads. CD45-enriched cells were stained and lung ILCs (CD45+CD127+Lin−NK1.1−) were purified by using FACSJazz. Sort purity is shown here. b, mRNA expression of Ccr6, Rorγt and Ahr relative to Gapdh in the purified ILCs (CD45+CD127+Lin−NK1.1−) and non-ILC population (CD45+CD127−Lin−NK1.1−) were quantified by RT–PCR. Significance is calculated by two-way ANOVA, ****P < 0.0001. Data are mean ± s.d. and individual values.
Extended Data Fig. 9 ILC1s and ILC2s are dispensable, while ILC3s are required for early protection against tuberculosis.
a–d, C57BL/6 (n = 5–10), Ifng−/− (n = 3–7), Il13−/− (n = 5–15) and Rorγt−/− (n = 4–5) mice were aerosol-infected with approximately 100 CFU Mtb. a, c, At 14 d.p.i., the bacterial burden was measured in the lungs by plating on 7H11 agar plates for 14 days. b, d, Numbers of lung ILC1s, ILC2s and ILC3s were quantified by flow cytometry. Significance is calculated by one-way ANOVA (a, b) or Student’s t-test (c, d). e, Cbfbf/fNkp46cre and Cbfbf/f mice were aerosol-infected with approximately 100 CFU Mtb and at 14 and 30 d.p.i. the bacterial burden was determined in the lungs by plating (n = 5–11 Cbfbf/f mouse; n = 3–11 Cbfbf/fNkp46cre mouse). f, Numbers of lung ILC1s, ILC2s, ILC3s and alveolar macrophages were determined by flow cytometry. g, FFPE lung sections from 30 d.p.i. Mtb-infected mice were stained with antibodies against B220 and CD3, and the average size of B cell follicles was quantified. h, i, Tissues from uninfected Cbfbf/fNkp46cre and Cbfbf/f mice were collected, and lung and lymph nodes were analysed for the different myeloid (h; alveolar macrophages, myeloid dendritic cells (mDCs), neutrophils, monocytes and recruited macrophages) and T cell (i; CD3+CD4+, CD3+CD8+, CD3+TCRα+, CD3+γδ+) populations by flow cytometry. Significance is calculated by Student’s t-test. Data are mean ± s.d. and individual values. All experiments were replicated at least twice.
Extended Data Fig. 10 IL-17 and IL-22 are produced by lung ILCs following Mtb infection and mediate protection through the CXCR5 axis.
C57BL/6 mice were aerosol-infected with approximately 100 CFU Mtb and treated with isotype (n = 5 mice) or anti-NK1.1 (n = 5 mice, PK126, 100 μg) antibodies every 3 days. a, Lung NK cells were determined following treatment with isotype or anti-NK1.1 antibodies at 30 d.p.i. by flow cytometry. b, Lung bacterial burden was assessed at 30 d.p.i. a, b, Significance calculated by Student’s t-test. c, d, Tissues from uninfected Ahrf/f and Ahrf/fRorγtcre mice were collected, and lung and lymph nodes were analysed for the different myeloid (c; alveolar macrophages, myeloid dendritic cells, neutrophils, monocytes and recruited macrophages) and T cell (d; CD3+CD4+, CD3+CD8+, CD3+TCRα+, CD3+γδ+) populations by flow cytometry (n = 9 mice per group). e, Lung cells from C57BL/6 mice were infected in vitro with Mtb (MOI of 0.1) and IL-23 protein levels were measured in supernatants on 5 d.p.i. and compared to uninfected (UI) cells. n = 3 uninfected dishes, n = 4 Mtb-infected dishes. f, Left, lung cells from C57BL/6 mice were infected in vitro with Mtb (MOI of 0.1) as in e and stimulated with recombinant (r)IL-23, rIL-1β, and the protein levels of IL-22 and IL-17 were measured in supernatants and compared with levels in uninfected cells. Right, numbers of IL-17- and IL-22-producing ILCs were measured by flow cytometry. g, C57BL/6 and Cxcr5−/− mice were aerosol-infected with around 100 CFU Mtb and at 30 d.p.i. the bacterial burden was determined in the lungs by plating (n = 5 mice per group). h, ILC3 quantification in FFPE lung sections was carried out by staining with antibodies against CD3, B220 and Rorγt, and the number of Rorγt+CD3− ILC3s were counted and are shown. i, FFPE lung sections from 30 d.p.i. Mtb-infected mice were stained with antibodies against B220 and CD3, and the average size of B cell follicles was quantified. Significance is calculated by Student’s t-test (c–i). Data are mean ± s.d. and individual values. All experiments were replicated at least twice.
Supplementary information
Supplementary Table 1
Differentially expressed genes of ILC2s sorted from TB-infected and control lungs. ILC2 were sorted from lung tissue and differential gene expression between TB infected (n=5) and uninfected control tissue (n=2) were determined by RNA sequencing. p-values corrected using Benjamini-Hochberg with a significance cut-off of FDR q< 0.01.
Supplementary Table 2
Differentially expressed genes of ILC2s sorted from TB-infected and control lungs. ILC2 were sorted from lung tissue and differential gene expression between TB infected (n=5) and uninfected control tissue (n=2) were determined by RNA sequencing. p-values corrected using Benjamini-Hochberg with a significance cut-off of FDR q< 0.01.
Supplementary Table 3
Predicted upstream drivers of DE response in ILC2s. Upstream drivers of differentially expressed genes in ILC2s were predicted using Ingenuity Pathway Analysis (IPA). p-values calculated by hypergeometric test between genes in our data and known interactions in the literature for each driver.
Supplementary Table 4
Predicted upstream drivers of DE response in ILC3s. Upstream drivers of differentially expressed genes in ILC3s were predicted using Ingenuity Pathway Analysis (IPA). p-values calculated by hypergeometric test between genes in our data and known interactions in the literature for each driver.
Supplementary Table 5
Downstream driver network for genes known to interact with OSM. Network created using the Mechanistic Networks generated by IPA, using a p-value< 0.01 for overlap significance.
Supplementary Table 6
Select predicted downstream pathways enriched in the differentially expressed genes in ILC3s between TB infected and uninfected samples. Generated in IPA.
Rights and permissions
About this article
Cite this article
Ardain, A., Domingo-Gonzalez, R., Das, S. et al. Group 3 innate lymphoid cells mediate early protective immunity against tuberculosis. Nature 570, 528–532 (2019). https://doi.org/10.1038/s41586-019-1276-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1276-2
This article is cited by
-
Treatment of Psoriasis Patients with Latent Tuberculosis Using IL-17 and IL-23 Inhibitors: A Retrospective, Multinational, Multicentre Study
American Journal of Clinical Dermatology (2024)
-
A candidate subunit vaccine induces protective immunity against Mycobacterium avium subspecies paratuberculosis in mice
npj Vaccines (2023)
-
Innate immune memory in inflammatory arthritis
Nature Reviews Rheumatology (2023)
-
Induction of killing of Mycobacterium avium subsp. hominissuis in macrophages by cytokine stimulated innate-like lymphoid cells is negatively affected by the pathogen
International Microbiology (2023)
-
Targeted isolation of antitubercular cycloheptapeptides and an unusual pyrroloindoline-containing new analog, asperpyrroindotide A, using LC–MS/MS-based molecular networking
Marine Life Science & Technology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.