Abstract
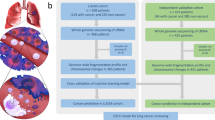
Cell-free DNA in the blood provides a non-invasive diagnostic avenue for patients with cancer1. However, characteristics of the origins and molecular features of cell-free DNA are poorly understood. Here we developed an approach to evaluate fragmentation patterns of cell-free DNA across the genome, and found that profiles of healthy individuals reflected nucleosomal patterns of white blood cells, whereas patients with cancer had altered fragmentation profiles. We used this method to analyse the fragmentation profiles of 236 patients with breast, colorectal, lung, ovarian, pancreatic, gastric or bile duct cancer and 245 healthy individuals. A machine learning model that incorporated genome-wide fragmentation features had sensitivities of detection ranging from 57% to more than 99% among the seven cancer types at 98% specificity, with an overall area under the curve value of 0.94. Fragmentation profiles could be used to identify the tissue of origin of the cancers to a limited number of sites in 75% of cases. Combining our approach with mutation-based cell-free DNA analyses detected 91% of patients with cancer. The results of these analyses highlight important properties of cell-free DNA and provide a proof-of-principle approach for the screening, early detection and monitoring of human cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Sequence data used in this study have been deposited at the database of Genotypes and Phenotypes (dbGaP, study ID 34536).
Code availability
Code for analyses is available at http:github.com/Cancer-Genomics/delfi_scripts.
References
Wan, J. C. M. et al. Liquid biopsies come of age: towards implementation of circulating tumour DNA. Nat. Rev. Cancer 17, 223–238 (2017).
Bray, F. et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68, 394–424 (2018).
World Health Organization. Guide to Cancer Early Diagnosis https://www.who.int/cancer/publications/cancer_early_diagnosis/en/ (WHO, 2017).
National Comprehensive Cancer Network. NCCN Clinical Practice Guidelines in Oncology https://www.nccn.org/professionals/physician_gls/default.aspx (accessed 16 April 2019).
Phallen, J. et al. Direct detection of early-stage cancers using circulating tumor DNA. Sci. Transl. Med. 9, eaan2415 (2017).
Cohen, J. D. et al. Detection and localization of surgically resectable cancers with a multi-analyte blood test. Science 359, 926–930 (2018).
Newman, A. M. et al. An ultrasensitive method for quantitating circulating tumor DNA with broad patient coverage. Nat. Med. 20, 548–554 (2014).
Bettegowda, C. et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci. Transl. Med. 6, 224ra24 (2014).
Leary, R. J. et al. Development of personalized tumor biomarkers using massively parallel sequencing. Sci. Transl. Med. 2, 20ra14 (2010).
Leary, R. J. et al. Detection of chromosomal alterations in the circulation of cancer patients with whole-genome sequencing. Sci. Transl. Med. 4, 162ra154 (2012).
Chan, K. C. et al. Noninvasive detection of cancer-associated genome-wide hypomethylation and copy number aberrations by plasma DNA bisulfite sequencing. Proc. Natl Acad. Sci. USA 110, 18761–18768 (2013).
Jiang, P. et al. Lengthening and shortening of plasma DNA in hepatocellular carcinoma patients. Proc. Natl Acad. Sci. USA 112, E1317–E1325 (2015).
Wang, B. G. et al. Increased plasma DNA integrity in cancer patients. Cancer Res. 63, 3966–3968 (2003).
Umetani, N. et al. Prediction of breast tumor progression by integrity of free circulating DNA in serum. J. Clin. Oncol. 24, 4270–4276 (2006).
Chan, K. C., Leung, S. F., Yeung, S. W., Chan, A. T. & Lo, Y. M. Persistent aberrations in circulating DNA integrity after radiotherapy are associated with poor prognosis in nasopharyngeal carcinoma patients. Clin. Cancer Res. 14, 4141–4145 (2008).
Mouliere, F. et al. High fragmentation characterizes tumour-derived circulating DNA. PLoS ONE 6, e23418 (2011).
Mouliere, F. et al. Enhanced detection of circulating tumor DNA by fragment size analysis. Sci. Transl. Med. 10, eaat4921 (2018).
Snyder, M. W., Kircher, M., Hill, A. J., Daza, R. M. & Shendure, J. Cell-free DNA comprises an in vivo nucleosome footprint that informs its tissues-of-origin. Cell 164, 57–68 (2016).
Underhill, H. R. et al. Fragment length of circulating tumor DNA. PLoS Genet. 12, e1006162 (2016).
Ulz, P. et al. Inferring expressed genes by whole-genome sequencing of plasma DNA. Nat. Genet. 48, 1273–1278 (2016).
Ivanov, M., Baranova, A., Butler, T., Spellman, P. & Mileyko, V. Non-random fragmentation patterns in circulating cell-free DNA reflect epigenetic regulation. BMC Genomics 16 (Suppl. 13), S1 (2015).
Jiang, P. et al. Preferred end coordinates and somatic variants as signatures of circulating tumor DNA associated with hepatocellular carcinoma. Proc. Natl Acad. Sci. USA 115, E10925–E10933 (2018).
Shen, S. Y. et al. Sensitive tumour detection and classification using plasma cell-free DNA methylomes. Nature 563, 579–583 (2018).
Corces, M. R. et al. The chromatin accessibility landscape of primary human cancers. Science 362, eaav1898 (2018).
Polak, P. et al. Cell-of-origin chromatin organization shapes the mutational landscape of cancer. Nature 518, 360–364 (2015).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Fortin, J. P. & Hansen, K. D. Reconstructing A/B compartments as revealed by Hi-C using long-range correlations in epigenetic data. Genome Biol. 16, 180 (2015).
Diehl, F. et al. Circulating mutant DNA to assess tumor dynamics. Nat. Med. 14, 985–990 (2008).
Phallen, J. et al. Early noninvasive detection of response to targeted therapy in non-small cell lung cancer. Cancer Res. 79, 1204–1213 (2019).
Burnham, P. et al. Single-stranded DNA library preparation uncovers the origin and diversity of ultrashort cell-free DNA in plasma. Sci. Rep. 6, 27859 (2016).
Sanchez, C., Snyder, M. W., Tanos, R., Shendure, J. & Thierry, A. R. New insights into structural features and optimal detection of circulating tumor DNA determined by single-strand DNA analysis. NPJ Genom. Med. 3, 31 (2018).
Fisher, S. et al. A scalable, fully automated process for construction of sequence-ready human exome targeted capture libraries. Genome Biol. 12, R1 (2011).
Jones, S. et al. Personalized genomic analyses for cancer mutation discovery and interpretation. Sci. Transl. Med. 7, 283ra53 (2015).
Benjamini, Y. & Speed, T. P. Summarizing and correcting the GC content bias in high-throughput sequencing. Nucleic Acids Res. 40, e72 (2012).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Friedman, J. H. Greedy function approximation: a gradient boosting machine. Ann. Stat. 29, 1189–1232 (2001).
Friedman, J. H. Stochastic gradient boosting. Comput. Stat. Data Anal. 38, 367–378 (2002).
Efron, B. & Tibshirani, R. Improvements on cross-validation: the 632+ bootstrap method. J. Am. Stat. Assoc. 92, 548–560 (1997).
Zurbenko, I. G. The Spectral Analysis of Time Series (Elsevier, 1986).
Robin, X. et al. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinformatics 12, 77 (2011).
Acknowledgements
We thank members of our laboratories for critical review of the manuscript. This work was supported, in part, by the Dr. Miriam and Sheldon G. Adelson Medical Research Foundation, the Stand Up to Cancer–Dutch Cancer Society International Translational Cancer Research Dream Team Grant (SU2C-AACR-DT1415), the Commonwealth Foundation, the Cigarette Restitution Fund, the Burroughs Wellcome Fund and the Maryland Genetics, Epidemiology and Medicine Training Program, the AACR-Janssen Cancer Interception Research Fellowship, the Mark Foundation for Cancer Research, US NIH (grants CA121113, CA006973, and CA180950), the Danish Council for Independent Research (11-105240), the Danish Council for Strategic Research (1309-00006B), the Novo Nordisk Foundation (NNF14OC0012747 and NNF17OC0025052), and the Danish Cancer Society (R133-A8520-00-S41 and R146-A9466-16-S2). Stand Up To Cancer is a program of the Entertainment Industry Foundation administered by the American Association for Cancer Research.
Reviewer information
Nature thanks Daniel De Carvalho, Ellen Heitzer and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
S.C., A.L., J.P., J.F., V. Adleff, R.B.S. and V.E.V. designed and planned the study, and developed and optimized experimental protocols. A.L., J.P., V. Adleff, J.E.M. and D.N.P. performed experiments. S.Ø.J., V. Anagnostou, P.F., J.N., K.M., J.B., B.D.W., H.H., K.L.v.R., M.-B.W.Ø., A.H.M., C.J.H.v.d.V., M.V., A.C., C.J.A.P., G.R.V., N.C.T.v.G., M.K., R.J.A.F., J.S.J., H.J.N., G.A.M. and C.L.A. organized patient enrolment, sample collection, and clinical data curation. S.C., A.L., J.P., J.F., V. Adleff, D.C.B., J.E.M., J.R.W., N.N., G.A.M., C.L.A., R.B.S. and V.E.V. analysed and interpreted data. S.C., A.L., J.P., J.F., R.B.S. and V.E.V. wrote the manuscript and incorporated feedback from all authors. S.C., A.L., J.P. and J.F. contributed equally to this study.
Corresponding authors
Ethics declarations
Competing interests
S.C., A.L., J.P., J.F., V. Adleff, R.B.S. and V.E.V. are inventors on patent applications (62/673,516 and 62/795,900) submitted by Johns Hopkins University related to cell-free DNA for cancer detection. V.E.V. is a founder of Delfi Diagnostics and Personal Genome Diagnostics, a member of their Scientific Advisory Boards and Boards of Directors, and owns Delfi Diagnostics and Personal Genome Diagnostics stock, which are subject to certain restrictions under university policy. Within the last five years, V.E.V. has been an advisor to Daiichi Sankyo, Janssen Diagnostics, Ignyta, and Takeda Pharmaceuticals. The terms of these arrangements are managed by Johns Hopkins University in accordance with its conflict of interest policies.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Simulations of non-invasive cancer detection based on number of alterations analysed and tumour-derived cfDNA fragment distributions.
a, Monte Carlo simulations were performed using different numbers of tumour-specific alterations to evaluate the probability of detecting cancer alterations in cfDNA at the indicated fraction of tumour-derived molecules. The simulations were performed assuming an average of 2,000 genome equivalents of cfDNA and the requirement of five or more observations of any alteration. These analyses indicate that increasing the number of tumour-specific alterations improves the sensitivity of detection of circulating tumour DNA. b, Cumulative density functions of cfDNA fragment lengths of 42 loci containing tumour-specific alterations from 30 patients with breast, colorectal, lung, or ovarian cancer are shown with 95% confidence bands (orange). Lengths of mutant cfDNA fragments were significantly different in size from wild-type cfDNA fragments (blue) at these loci. c, GC content was similar for mutated and non-mutated fragments. d, GC content was not correlated to fragment length.
Extended Data Fig. 2 Germline and haematopoietic cfDNA fragment distributions.
a, Cumulative density functions of fragment lengths at 44 loci containing germline alterations (non-tumour derived) from 38 patients with breast, colorectal, lung or ovarian cancer are shown with 95% confidence bands. Fragments with germline mutations (orange) were comparable in length to wild-type cfDNA fragment lengths (blue). b, Cumulative density functions of fragment lengths at 41 loci containing haematopoietic alterations (non-tumour derived) from 28 patients with breast, colorectal, lung or ovarian cancer are shown with 95% confidence bands. After correction for multiple testing, there were no significant differences (α = 0.05) in the size distributions of mutated haematopoietic cfDNA fragments (orange) and wild-type cfDNA fragments (blue).
Extended Data Fig. 3 cfDNA fragmentation in healthy individuals and patients with lung cancer.
a, cfDNA fragment lengths are shown for healthy individuals (n = 30, grey) and patients with lung cancer (n = 8, blue). b–d, cfDNA fragmentation profiles from healthy individuals (n = 30) had high correlations, whereas patients with lung cancer (n = 8) had lower correlations to median fragmentation profiles of lymphocytes (b), lymphocyte nucleosome distances (c) and healthy cfDNA (d). Pearson correlations are shown with box plots depicting minimum, 25th percentile, median, 75th percentile, and maximum values. e, High coverage (9×) WGS data were subsampled to 2×, 1×, 0.5×, 0.2× and 0.1×-fold coverage. Mean centred genome-wide fragmentation profiles in 5-Mb bins for 30 healthy individuals and 8 patients with lung cancer are depicted for each subsampled fold coverage with median profiles shown in blue. f, Pearson correlation of subsampled profiles to initial profile at 9× coverage for healthy individuals and patients with lung cancer.
Extended Data Fig. 4 cfDNA fragmentation profiles and sequence alterations during therapy.
Detection and monitoring of cancer in serial blood draws from patients with non-small cell lung cancer (n = 19) undergoing treatment with targeted tyrosine kinase inhibitors (black arrows) was performed using targeted sequencing (top) as previously reported29, and genome-wide fragmentation profiles (bottom). For each case, the vertical axis of the bottom panel displays −1 times the Pearson correlation of each sample to the median healthy cfDNA fragmentation profile. Error bars depict confidence intervals from binomial tests for mutant allele fractions, and confidence intervals calculated using Fisher transformation for genome-wide fragmentation profiles. Although the approaches analyse different aspects of cfDNA (whole genome compared with specific alterations), the targeted sequencing and fragmentation profiles were similar for patients responding to therapy as well as those with stable or progressive disease. As fragmentation profiles reflect both genomic and epigenomic alterations (whereas mutant allele fractions only reflect individual mutations), mutant allele fractions alone may not reflect the absolute level of correlation of fragmentation profiles to healthy individuals.
Extended Data Fig. 5 Profiles of cfDNA fragment lengths in copy neutral regions in healthy individuals and one patient with colorectal cancer.
a, The fragmentation profiles in 211 copy neutral windows in chromosomes 1–6 are shown for 25 randomly selected healthy individuals (grey). For a patient with colorectal cancer (CGCRC291) with an estimated mutant allele fraction of 20%, we diluted the cancer fragment length profile to an approximate 10% tumour contribution (blue). a, b, Although the marginal densities of the fragment profiles for the healthy samples and patient with cancer show substantial overlap (a, right), the fragmentation profiles are different as can be seen through visualization of the fragmentation profiles (a, left) and by the separation of the patient with colorectal cancer from the healthy samples (n = 25) in a principal component analysis (b).
Extended Data Fig. 6 Genome-wide GC correction of cfDNA fragments.
To estimate and control for the effects of GC content on sequencing coverage, we calculated coverage in non-overlapping 100-kb genomic windows across the autosomes. For each window, we calculated the average GC of the aligned fragments. a, LOESS smoothing of raw coverage (top row) for two randomly selected healthy subjects (CGPLH189 and CGPLH380) and two patients with cancer (CGPLLU161 and CGPLBR24) with undetectable aneuploidy (PA score < 2.35). After subtracting the average coverage predicted by the LOESS model, the residuals were rescaled to the median autosomal coverage (bottom row). As fragment length may also result in coverage biases, we performed this GC correction procedure separately for short (≤150 bp) and long (>150 bp) fragments. Although the 100-kb bins on chromosome 19 (blue points) consistently have less coverage than predicted by the LOESS model, we did not implement a chromosome-specific correction as such an approach would remove the effects of chromosomal copy number on coverage. b, Overall, we found a limited correlation between short or long fragment coverage and GC content after correction among healthy individuals (n = 211, interquartile range: −0.03–0.03) and patients with cancer (n = 128, interquartile range: −0.06–0.02) with a PA score < 3. Box plots depict 25th percentile, median, and 75th percentile values.
Extended Data Fig. 7 Machine learning model.
a, We used gradient tree boosting machine learning to examine whether cfDNA can be categorized as having characteristics of a patient with cancer or a healthy individual. The machine learning model included fragmentation size and coverage characteristics in windows throughout the genome, as well as chromosomal arm and mitochondrial DNA copy numbers. We used a tenfold cross-validation approach in which each sample is randomly assigned to a fold, and nine of the folds (90% of the data) are used for training and one fold (10% of the data) is used for testing. The prediction accuracy from a single cross-validation is an average over the ten possible combinations of test and training sets. As this prediction accuracy can reflect bias from the initial randomization of patients, we repeat the entire procedure, including the randomization of patients to folds, ten times. For all cases, feature selection and model estimation were performed on training data and were validated on test data, and the test data were never used for feature selection. Ultimately, we obtained a DELFI score that could be used to classify individuals as likely to be healthy or having cancer. b, Distribution of AUCs across the repeated tenfold cross-validation. The 25th, 50th and 75th percentiles of the 100 AUCs for the cohort of 215 healthy individuals and 208 patients with cancer are indicated by dashed lines.
Extended Data Fig. 8 Whole-genome analyses of chromosomal arm copy number changes and mitochondrial genome representation.
a, Z-scores for each autosome arm are depicted for healthy individuals (n = 215) and patients with cancer (n = 208). The vertical axis depicts normal copy at zero with positive and negative values indicating arm gains and losses, respectively. Z-scores greater than 50 or less than −50 are thresholded at the indicated values. b, The fraction of reads mapping to the mitochondrial genome is depicted for healthy individuals (n = 215) and patients with cancer (n = 208). Box plots depict the minimum, 25th percentile, median, 75th percentile, and maximum values.
Extended Data Fig. 9 DELFI detection of cancer and tissue of origin prediction.
a, Analyses of individual cancer types using DELFI had AUCs ranging from 0.86 to >0.99. b, Receiver operator characteristics for detection of cancer using cfDNA fragmentation profiles and other genome-wide features in a machine learning approach are depicted for a cohort of 215 healthy individuals and each stage of 208 patients with cancer with ≥95% specificity shaded in blue. c, Receiver operator characteristics for DELFI tissue prediction of bile duct, breast, colorectal, gastric, lung, ovarian or pancreatic cancer are depicted. To increase sample sizes within cancer type classes, we included cases detected with a 90% specificity, and the lung cancer cohort was supplemented with the addition of baseline cfDNA data from 18 patients with lung cancer with prior treatment36. d, DELFI tissue of origin prediction.
Extended Data Fig. 10 Detection of cancer using DELFI and mutation-based cfDNA approaches.
DELFI (green) and targeted sequencing10 for mutation identification (blue) were performed independently in a cohort of 126 patients with breast, bile duct, colorectal, gastric, lung or ovarian cancer. The number of individuals detected by each approach and in combination are indicated for DELFI detection with a specificity of 98%, targeted sequencing specificity at >99%, and a combined specificity of 98%. ND, not detected.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables 1-8.
Rights and permissions
About this article
Cite this article
Cristiano, S., Leal, A., Phallen, J. et al. Genome-wide cell-free DNA fragmentation in patients with cancer. Nature 570, 385–389 (2019). https://doi.org/10.1038/s41586-019-1272-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1272-6
This article is cited by
-
Prediction of methylation status using WGS data of plasma cfDNA for multi-cancer early detection (MCED)
Clinical Epigenetics (2024)
-
Multimodal epigenetic sequencing analysis (MESA) of cell-free DNA for non-invasive colorectal cancer detection
Genome Medicine (2024)
-
Terminal modifications independent cell-free RNA sequencing enables sensitive early cancer detection and classification
Nature Communications (2024)
-
An in vitro CRISPR screen of cell-free DNA identifies apoptosis as the primary mediator of cell-free DNA release
Communications Biology (2024)
-
Improving diagnostic accuracy of identifying gastric cancer patients with peritoneal metastases: tumor-guided cell-free DNA analysis of peritoneal fluid
Oncogene (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.