Abstract
Serine hydroxymethyltransferase 2 (SHMT2) regulates one-carbon transfer reactions that are essential for amino acid and nucleotide metabolism, and uses pyridoxal-5′-phosphate (PLP) as a cofactor. Apo SHMT2 exists as a dimer with unknown functions, whereas PLP binding stabilizes the active tetrameric state. SHMT2 also promotes inflammatory cytokine signalling by interacting with the deubiquitylating BRCC36 isopeptidase complex (BRISC), although it is unclear whether this function relates to metabolism. Here we present the cryo-electron microscopy structure of the human BRISC–SHMT2 complex at a resolution of 3.8 Å. BRISC is a U-shaped dimer of four subunits, and SHMT2 sterically blocks the BRCC36 active site and inhibits deubiquitylase activity. Only the inactive SHMT2 dimer—and not the active PLP-bound tetramer—binds and inhibits BRISC. Mutations in BRISC that disrupt SHMT2 binding impair type I interferon signalling in response to inflammatory stimuli. Intracellular levels of PLP regulate the interaction between BRISC and SHMT2, as well as inflammatory cytokine responses. These data reveal a mechanism in which metabolites regulate deubiquitylase activity and inflammatory signalling.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
References
Giardina, G. et al. How pyridoxal 5′-phosphate differentially regulates human cytosolic and mitochondrial serine hydroxymethyltransferase oligomeric state. FEBS J. 282, 1225–1241 (2015).
Anderson, D. D., Woeller, C. F., Chiang, E.-P., Shane, B. & Stover, P. J. Serine hydroxymethyltransferase anchors de novo thymidylate synthesis pathway to nuclear lamina for DNA synthesis. J. Biol. Chem. 287, 7051–7062 (2012).
Szebenyi, D. M., Liu, X., Kriksunov, I. A., Stover, P. J. & Thiel, D. J. Structure of a murine cytoplasmic serine hydroxymethyltransferase quinonoid ternary complex: evidence for asymmetric obligate dimers. Biochemistry 39, 13313–13323 (2000).
Patterson-Fortin, J., Shao, G., Bretscher, H., Messick, T. E. & Greenberg, R. A. Differential regulation of JAMM domain deubiquitinating enzyme activity within the RAP80 complex. J. Biol. Chem. 285, 30971–30981 (2010).
Cooper, E. M., Boeke, J. D. & Cohen, R. E. Specificity of the BRISC deubiquitinating enzyme is not due to selective binding to Lys63-linked polyubiquitin. J. Biol. Chem. 285, 10344–10352 (2010).
Feng, L., Wang, J. & Chen, J. The Lys63-specific deubiquitinating enzyme BRCC36 is regulated by two scaffold proteins localizing in different subcellular compartments. J. Biol. Chem. 285, 30982–30988 (2010).
Sobhian, B. et al. RAP80 targets BRCA1 to specific ubiquitin structures at DNA damage sites. Science 316, 1198–1202 (2007).
Wang, B. et al. Abraxas and RAP80 form a BRCA1 protein complex required for the DNA damage response. Science 316, 1194–1198 (2007).
Kim, H., Chen, J. & Yu, X. Ubiquitin-binding protein RAP80 mediates BRCA1-dependent DNA damage response. Science 316, 1202–1205 (2007).
Jiang, Q. et al. MERIT40 cooperates with BRCA2 to resolve DNA interstrand cross-links. Genes Dev. 29, 1955–1968 (2015).
Sowa, M. E., Bennett, E. J., Gygi, S. P. & Harper, J. W. Defining the human deubiquitinating enzyme interaction landscape. Cell 138, 389–403 (2009).
Zheng, H. et al. A BRISC-SHMT complex deubiquitinates IFNAR1 and regulates interferon responses. Cell Reports 5, 180–193 (2013).
Zeqiraj, E. et al. Higher-order assembly of BRCC36–KIAA0157 is required for DUB activity and biological function. Mol. Cell 59, 970–983 (2015).
Walden, M., Masandi, S. K., Pawłowski, K. & Zeqiraj, E. Pseudo-DUBs as allosteric activators and molecular scaffolds of protein complexes. Biochem. Soc. Trans. 46, 453–466 (2018).
Kyrieleis, O. J. P. et al. Three-dimensional architecture of the human BRCA1-A histone deubiquitinase core complex. Cell Reports 17, 3099–3106 (2016).
Zanetti, K. A. & Stover, P. J. Pyridoxal phosphate inhibits dynamic subunit interchange among serine hydroxymethyltransferase tetramers. J. Biol. Chem. 278, 10142–10149 (2003).
Jagath, J. R., Sharma, B., Rao, N. A. & Savithri, H. S. The role of His-134, -147, and -150 residues in subunit assembly, cofactor binding, and catalysis of sheep liver cytosolic serine hydroxymethyltransferase. J. Biol. Chem. 272, 24355–24362 (1997).
Jala, V. R., Appaji Rao, N. & Savithri, H. S. Identification of amino acid residues, essential for maintaining the tetrameric structure of sheep liver cytosolic serine hydroxymethyltransferase, by targeted mutagenesis. Biochem. J. 369, 469–476 (2003).
Krishna Rao, J. V., Jagath, J. R., Sharma, B., Appaji Rao, N. & Savithri, H. S. Asp-89: a critical residue in maintaining the oligomeric structure of sheep liver cytosolic serine hydroxymethyltransferase. Biochem. J. 343, 257–263 (1999).
Xu, D., Jaroszewski, L., Li, Z. & Godzik, A. FFAS-3D: improving fold recognition by including optimized structural features and template re-ranking. Bioinformatics 30, 660–667 (2014).
Zimmermann, L. et al. A completely reimplemented MPI bioinformatics toolkit with a new HHpred server at its core. J. Mol. Biol. 430, 2237–2243 (2018).
Björklund, A. K., Ekman, D. & Elofsson, A. Expansion of protein domain repeats. PLOS Comput. Biol. 2, e114 (2006).
Hu, X. et al. NBA1/MERIT40 and BRE interaction is required for the integrity of two distinct deubiquitinating enzyme BRCC36-containing complexes. J. Biol. Chem. 286, 11734–11745 (2011).
Guettler, S. et al. Structural basis and sequence rules for substrate recognition by tankyrase explain the basis for cherubism disease. Cell 147, 1340–1354 (2011).
Hamilton, G., Colbert, J. D., Schuettelkopf, A. W. & Watts, C. Cystatin F is a cathepsin C-directed protease inhibitor regulated by proteolysis. EMBO J. 27, 499–508 (2008).
Zheng, N. & Shabek, N. Ubiquitin ligases: structure, function, and regulation. Annu. Rev. Biochem. 86, 129–157 (2017).
Yang, X. et al. SHMT2 desuccinylation by SIRT5 drives cancer cell proliferation. Cancer Res. 78, 372–386 (2018).
Cao, J. et al. HDAC11 regulates type I interferon signaling through defatty-acylation of SHMT2. Proc. Natl Acad. Sci. USA 116, 5487–5492 (2019).
Fitzgerald, D. J., et al. Protein complex expression by using multigene baculoviral vectors. Nat. Methods 3, 1021–1032 (2006).
Thompson, R. F., Iadanza, M. G., Hesketh, E. L., Rawson, S. & Ranson, N. A. Collection, pre-processing and on-the-fly analysis of data for high-resolution, single-particle cryo-electron microscopy. Nat. Protocols 14, 100–118 (2019).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Zheng, S. Q., et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 2006 3:12 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. E. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protocols 10, 845–858 (2015).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Pickart, C. M. & Raasi, S. Controlled synthesis of polyubiquitin chains Methods Enzymol. 399, 21–36 (2005).
Wei, Z. et al. Deacetylation of serine hydroxymethyl-transferase 2 by SIRT3 promotes colorectal carcinogenesis. Nat Commun. 9, 4468 (2018).
Byrne, D. P. et al. cAMP-dependent protein kinase (PKA) complexes probed by complementary differential scanning fluorimetry and ion mobility-mass spectrometry. Biochem. J. 473, 3159–3175 (2016).
Scarff, C. A. et al. Examination of ataxin-3 (atx-3) aggregation by structural mass spectrometry techniques: a rationale for expedited aggregation upon polyglutamine (polyQ) expansion. Mol. Cell. Proteomics 14, 1241–1253 (2015).
Gault, J., et al. High-resolution mass spectrometry of small molecules bound to membrane proteins. Nat. Methods 13, 333–336 (2016).
Rose, R. J., Damoc, E., Denisov, E., Makarov, A. & Heck, A. J. R. High-sensitivity Orbitrap mass analysis of intact macromolecular assemblies. Nat. Methods 9, 1084–1086 (2012).
Marty, M. T. et al. Bayesian deconvolution of mass and ion mobility spectra: from binary interactions to polydisperse ensembles. Anal. Chem. 87, 4370–4376 (2015).
Shao, G. et al. MERIT40 controls BRCA1–Rap80 complex integrity and recruitment to DNA double-strand breaks. Genes Dev. 23, 740–754 (2009).
Ochocki, J. D. et al. Arginase 2 suppresses renal carcinoma progression via biosynthetic cofactor pyridoxal phosphate depletion and increased polyamine toxicity. Cell Metab. 27, 1263–1280.e6 (2018).
Acknowledgements
We thank F. Sicheri, J. Fontana, A. Berry, C. Simon, I. Nissim, F. Sobott, J. Cockburn and A. Moloney for useful discussions, and R. Thompson, A. Gordon, M. Iadanza and M. Fuller for technical support. This work was supported by a Sir Henry Dale Fellowship (Wellcome Trust and the Royal Society; 200523/Z/16/Z) to E.Z., MRC MC_PC-16050 grant to E.Z. and F.D.G., NIH R01 CA138835 and a Lupus Research Alliance Target Identification in Lupus grant to R.A.G. and J.M.S., a BBSRC (BB/L021250/1) grant to N.A.R., a BBSRC TDRF grant BB/N021703/1 and North West Cancer Research grants CR1088, CR1097 to P.A.E., and a Polish National Science Centre grant (2014/15/B/NZ1/03359) to K.P. The University of Leeds mass spectrometry facility is supported by BBSRC (BB/E012558/1) and Wellcome Trust (208385/Z/17/Z) grants. The Astbury cryo-EM facility is funded by a University of Leeds ABSL award and Wellcome Trust grant 108466/Z/15/Z.
Reviewer information
Nature thanks Tony Huang and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
M.W. performed electron microscopy experiments, data processing and model building with assistance from E.L.H. and S.K.M. L.T. performed cell-based experiments, immunoprecipitation, challenge assays, metabolic analysis and immunoblots. M.W. and U.M.S. cloned and purified proteins, and performed in vitro interaction studies. R.L.R. and F.D.G. performed qRT-PCR and gene-expression data analysis. D.P.B. and P.A.E. performed differential scanning fluorimetry assays and data analysis. J.C. performed enzyme kinetics. R.G. and J.R.A. performed mass spectrometry analyses. F.E.O. generated reagents. K.P. performed bioinformatics analyses. J.M.S., P.A.E., F.D.G., N.A.R. and R.A.G. contributed to conception and supervision of experiments. E.Z. performed enzyme assays and conceived and supervised the project. E.Z., M.W., L.T. and R.A.G. wrote the manuscript with input from N.A.R., P.A.E. and F.D.G. All authors critically reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
F.E.O. declares competing financial interests as co-founder and shareholder of UbiQ Bio BV.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 SHMT2 exists in a dimer–tetramer equilibrium.
a, Structure of SHMT2ΔN(A285T) dimer (PDB code 6DK3). Unbiased electron density maps are coloured green. The structure was erroneously deposited by the Structural Genomics Consortium (SGC) as wild type with PDB code 3OU5. It was corrected and redeposited with PDB code 6DK3 on 28 May 2018. b, Structure of wild-type SHMT2α tetramer (PDB code 4PVF) bound to PLP. Regions in red show structural rearrangements upon PLP binding. c, Native mass spectrometry spectra (single measurement experiment) of different forms of SHMT2ΔN (top) and measured and calculated masses (bottom). Calculated masses nearest to the measured masses are underlined. d, Size-exclusion chromatography profiles of different forms of SHMT2ΔN (top) and a Coomassie-stained SDS–PAGE gel (bottom). Data are representative of three independent experiments. e, Comparison of SHMT2 protomers with and without PLP.
Extended Data Fig. 2 PLP regulates SHMT2 dimer-tetramer transition.
a, PLP association with the indicated forms of SHMT2ΔN. SHMT2–PLP internal aldimine linkage was monitored by measuring absorbance at 435 nm after rapid mixing of SHMT2ΔN (5 μM) and PLP (100 μM). Data are mean ± s.e.m. of four independent reactions. b, Enzyme activity of the indicated forms of SHMT2ΔN (0.5 μM) against the synthetic substrate l-threo-phenylserine in the presence of 50 μM PLP. The reaction was monitored by measuring benzyldehyde absorbance at 279 nm. Data are mean ± s.e.m. of three independent experiments carried out in duplicate. c, Differential scanning fluorimetry analysis of different forms of SHMT2ΔN in the presence and absence of PLP. SHMT2ΔN wild-type and mutant proteins (5 μM) were incubated with the indicated concentrations of PLP or 100 μM 2-hydroxy-6-methylpyridine-3-carboxylic acid (HMCA) for 10 min at 20 °C before differential scanning fluorimetry analysis. d, Changes in melting temperatures (ΔTm) for each of the conditions shown in c. ΔTm was calculated by subtracting the Tm of SHMT2 with buffer without ligands from the Tm of SHMT2 with added ligands. Data in c, d represent a single experiment carried out in duplicate. e, Summary of SHMT2ΔN(A285T) dimer–tetramer equilibrium in response to PLP binding.
Extended Data Fig. 3 Reconstitution of BRISC–SHMT2 complex.
a, Summary of limited proteolysis and Edman sequencing results indicating trypsin protease-labile regions for the BRCC45–MERIT40 complex (left) and the BRISC (right). Protease cleavage sites are shown as scissors. Unstructured Abraxas 2 regions are indicated as dashed lines. b, SDS–PAGE of BRISC and BRISCΔNΔC (representative of three independent experiments). c, DUB activity of BRISC (full length) and BRISCΔNΔC (truncated) against a fluorogenic K63-linked diUb substrate. Data are mean ± s.e.m. of three independent experiments carried out in duplicate. d, SHMT2ΔN(A285T) inhibition of BRISC and BRISCΔNΔC DUB activity. Data are mean ± s.e.m. of three independent experiments carried out in duplicate. Ninety-five per cent confidence intervals are shown in square brackets. e, Size-exclusion chromatography of the BRISCΔNΔC–SHMT2ΔN(A285T) complex (top) and Coomassie-stained SDS–PAGE of peak fractions (bottom). f, Native mass spectrometry of the BRISCΔNΔC–SHMT2ΔN(A285T) complex. Data in e, f are representative of three independent experiments.
Extended Data Fig. 4 Electron microscopy analysis of BRISC and BRISC–SHMT2 complex.
a, Negative-stain electron microscopy analysis. Selected 2D-averaged classes of BRISCΔNΔC (left) and BRISCΔNΔC–SHMT2ΔN(A285T) (right). b, Representative cryo-EM micrograph. Green circles (180 Å diameter) indicate auto-picked particles. c, Flow chart of data processing. Final electron microscopy maps, coloured according to local resolution, are provided (scale bar in Å). d, FSC curves for the indicated maps. The final resolution was calculated using the gold-standard FSC cut-off at 0.143 frequency.
Extended Data Fig. 5 Structural analysis of BRISC–SHMT2 structures.
a, Structural overview of the BRISCΔNΔC–SHMT2 structure and cryo-EM maps at a contour level of 0.085 (C2 map), showing secondary structure features. BRCC45 UEV-C and MERIT40 vWFA domains are omitted. CCHB, coiled-coil helical bundle. Asterisks mark the position of the BRCC36 active site. An overreaching loop (SHMT2 residues 296–307) is coloured red. b, Representative electron microscopy map of the BRCC36 active site. c, d, Structure of human (c) and insect (C. floridanus) (d) BRCC36–Abraxas 2 superdimer, highlighting differences and similarities. The connectivity of the MPN- domain of Abraxas 2 and α4 helix by a crossover loop is different in the insect and human complexes, and is coloured orange in the latter structure. e, Superimposition of the BRCC36 MPN+ domain from the insect (grey) and human (blue, green) heterodimers. Abraxas 2 regions with notable differences between the two structures are coloured orange.
Extended Data Fig. 6 Analysis of SHMT1 interaction with BRISC.
a, Elution profile of wild-type SHMT1 from an S75 16/600 size-exclusion chromatography column (data are representative of two independent experiments). b, Elution profile of SHMT1 containing mutated residues designed to break the tetrameric interface (left; data are from a single experiment). Structure of SHMT1 tetramer (PDB code 1RV4), highlighting residues that are important for tetramerization (right). c, BRISC DUB activity against a fluorogenic K63-linked diUb substrate in the presence of the indicated forms of SHMT1 and SHMT2. Data are mean ± s.e.m. of three independent experiments carried out in duplicate. d, Coomassie-stained SDS–PAGE analysis of the indicated SHMT1 and SHMT2 protein preparations (data are from a single experiment). e, Analytical size-exclusion chromatography runs of the indicated protein preparations. SHMT1 proteins were mixed with BRISC for 30 min before injecting on a 2.4-ml Superose 6 column (bottom traces). Traces on top are control runs. A BRISC–SHMT2ΔN run is shown for comparison (see Extended Data Fig. 8e for detailed analysis). f, Coomassie-stained SDS–PAGE analysis of the indicated peak fractions from size-exclusion runs shown in e. Data in e, f are representative of two independent experiments.
Extended Data Fig. 7 Purification and analysis of SHMT2 mutants.
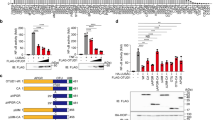
a, Elution profile of the indicated forms of SHMT2ΔN from an S75 10/300 size-exclusion chromatography column (single experiment). b, Coomassie-stained SDS–PAGE analysis of the indicated SHMT2ΔN protein preparations (data are representative of two independent experiments). c, BRISC DUB activity against a fluorogenic K63-linked diUb substrate in the presence of the indicated SHMT2ΔN mutants. Data are mean ± s.e.m. of three independent experiments carried out in duplicate. d, BRISC DUB activity against K63-linked hexaUb chains in the presence of the different forms of SHMT2ΔN, or SHMT1. e, Ubiquitylation levels of IFNAR1 after IFNα stimulation in HEK293T cells that overexpress the indicated forms of Abraxas 2 and SHMT2ΔN. The annotation ‘LL->RR’ denotes SHMT2ΔN(L211R/L215R). IFNAR1 immunoprecipitation (IP) was performed under denaturing conditions and ubiquitin levels were detected using the vu-1 antibody. Mock IP was performed using a generic rabbit IgG antibody. f, Immunoprecipitation (IP) performed using anti-Flag antibody in MEFs that were transiently transfected with Flag–HA epitope-tagged SHMT2ΔN or mutants. Immunoblot was performed for Abraxas 2 and SHMT2, as indicated. UTF, untransfected cells (used as control). g, MEFs that overexpress the indicated SHMT2ΔN or mutants were challenged with LPS, and interferon-receptor-dependent signal transduction response was assessed by immunoblot for STAT1 phosphorylated at Y701. Data shown in d, e are representative of three independent experiments. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 8 SHMT2 blocks the BRISC active site.
a, Analytical size-exclusion chromatography of different forms of SHMT2ΔN, preincubated with BRISC (left) and SDS–PAGE analysis of peak fractions (right). Data are representative of two independent experiments. b, K63-linked diUb (grey) modelled on the MPN+ domain of BRCC36 using the AMSH-LP–diUb structure (PDB code 2ZNV) as a guide. The SHMT2 obligate dimer sterically clashes with the modelled proximal ubiquitin. c, Michaelis–Menten (top) and Lineweaver–Burk (bottom) plots for BRISC DUB activity against a K63-linked diUb fluorogenic substrate with addition of SHMT2ΔN(A285T). Technical duplicates are shown and data are representative of two independent experiments. d, Superimposition of SHMT2 dimer and tetramer forms. The SHMT2 obligate dimer from the PLP-bound tetramer structure (PDB code 4PVF, coloured grey) was overlaid on the SHMT2ΔN(A285T) dimer bound to BRISC. The second obligate dimer from the SHMT2–PLP holoenzyme sterically clashes with the BRCC36–Abraxas 2 superdimer. Movement of α6 and α7 helices is indicated. e, Analytical size-exclusion chromatography of SHMT2ΔN dimer and tetramer (+PLP) forms with BRISC (left) and SDS–PAGE analysis of peak fractions (right). Data are representative of two independent experiments.
Extended Data Fig. 9 Regulation of BRISC–SHMT2 function in cells.
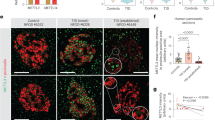
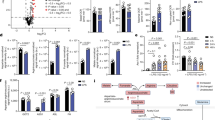
a, Schematic of pairwise comparison of Abraxas2–/– MEFs treated with LPS and expressing the indicated forms of Abraxas 2 (top). Bar chart illustrates the IFN-type I-related genes that were increased by more than twofold in Abraxas 2 wild-type + LPS (black) and Abraxas 2(E144R) + LPS (blue), relative to Abraxas2-/- + LPS. Data are presented as mean ± s.e.m. from three independent experiments (biological replicates). b, Native gel electrophoresis of whole MEF cell lysates cultured in B6-vitamer-free medium, with and without pyridoxal, and expressing the indicated forms of Flag–HA–SHMT2ΔN. c, d, Measurement of phosphorylated STAT1 levels (phosphorylated at Y701) in MEFs (c) or MEFs that stably express SHMT2 (d), cultured in B6-vitamer-free medium. Where indicated, cells were treated with 20 μM pyridoxal and/or 500 units per millilitre of IFNβ for 48 h. e, Immunoprecipitation performed using anti-Flag antibody in MEFs transiently transfected with Flag–HA epitope-tagged SHMT2 or mutants (cultured in DMEM). Immunoblot was performed for Abraxas 2 and SHMT2 as indicated. f, Phosphorylated STAT1 levels (phosphorylated at Y701) measured in MEFs cultured in DMEM containing vitamin B6 and the indicated forms of SHMT, after challenge with HSV. Data shown in b–f are representative of three independent experiments. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 10 Proposed model of BRISC–SHMT2 regulation of cytokine signalling.
K63-linked poly-ubiquitylated interferon receptors (IFNAR1 and IFNAR2) are internalized and degraded through the lysosomal degradation pathway. The BRISC–SHMT2 complex is required for deubiquitylation of IFNAR1 and IFNAR2, and for receptor stabilization. BRISC deubiquitylation of IFNAR1 and IFNAR2 receptors proceeds after SHMT2 displacement, which leads to sustained interferon signalling. The SHMT2 dimer–tetramer transition is regulated by PLP binding. Only an inactive SHMT2 dimer can interact with BRISC, whereas the active SHMT2 tetramer will sterically clash with the BRCC36–Abraxas 2 superdimer. Modulation of SHMT2 dimer–tetramer transition by PLP analogues or other binders could represent ways to inhibit BRISC recruitment to IFNAR1 and IFNAR2, and to reduce interferon signalling.
Supplementary information
Supplementary Information
This file contains uncropped gels and blots for data shown in main-text and extended data figures.
Rights and permissions
About this article
Cite this article
Walden, M., Tian, L., Ross, R. et al. Metabolic control of BRISC–SHMT2 assembly regulates immune signalling. Nature 570, 194–199 (2019). https://doi.org/10.1038/s41586-019-1232-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1232-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.