Abstract
The sympathetic nervous system drives brown and beige adipocyte thermogenesis through the release of noradrenaline from local axons. However, the molecular basis of higher levels of sympathetic innervation of thermogenic fat, compared to white fat, has remained unknown. Here we show that thermogenic adipocytes express a previously unknown, mammal-specific protein of the endoplasmic reticulum that we term calsyntenin 3β. Genetic loss or gain of expression of calsyntenin 3β in adipocytes reduces or enhances functional sympathetic innervation, respectively, in adipose tissue. Ablation of calsyntenin 3β predisposes mice on a high-fat diet to obesity. Mechanistically, calsyntenin 3β promotes endoplasmic-reticulum localization and secretion of S100b—a protein that lacks a signal peptide—from brown adipocytes. S100b stimulates neurite outgrowth from sympathetic neurons in vitro. A deficiency of S100b phenocopies deficiency of calsyntenin 3β, and forced expression of S100b in brown adipocytes rescues the defective sympathetic innervation that is caused by ablation of calsyntenin 3β. Our data reveal a mammal-specific mechanism of communication between thermogenic adipocytes and sympathetic neurons.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Histone modification marker and transcription factor ChIP–seq datasets generated in this study are available at NIH Sequence Read Archive under the accession code PRJNA526243. Any other relevant data are available from the corresponding author upon reasonable request.
Change history
22 May 2019
In Fig. 6a of this Article, the two dots corresponding to Cidea and S100b were erroneously moved to the top left of the volcano plot; this figure has been corrected online.
An amendment to this paper has been published and can be accessed via a link at the top of the paper
03 February 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41586-020-03161-z
References
Cannon, B. & Nedergaard, J. Brown adipose tissue: function and physiological significance. Physiol. Rev. 84, 277–359 (2004).
Seale, P. & Lazar, M. A. Brown fat in humans: turning up the heat on obesity. Diabetes 58, 1482–1484 (2009).
Morrison, S. F. Central neural control of thermoregulation and brown adipose tissue. Auton. Neurosci. 196, 14–24 (2016).
Zeng, W. et al. Sympathetic neuro-adipose connections mediate leptin-driven lipolysis. Cell 163, 84–94 (2015).
Fedorenko, A., Lishko, P. V. & Kirichok, Y. Mechanism of fatty-acid-dependent UCP1 uncoupling in brown fat mitochondria. Cell 151, 400–413 (2012).
Kazak, L. et al. A creatine-driven substrate cycle enhances energy expenditure and thermogenesis in beige fat. Cell 163, 643–655 (2015).
Daniel, H. & Derry, D. M. Criteria for differentiation of brown and white fat in the rat. Can. J. Physiol. Pharmacol. 47, 941–945 (1969).
Seale, P. et al. Transcriptional control of brown fat determination by PRDM16. Cell Metab. 6, 38–54 (2007).
Cohen, P. et al. Ablation of PRDM16 and beige fat causes metabolic dysfunction and subcutaneous to visceral adipose switch. Cell 156, 304–316 (2013).
Wang, W. & Seale, P. Control of brown and beige fat development. Nat. Rev. Mol. Cell Biol. 17, 691–702 (2016).
Seale, P. et al. Prdm16 determines the thermogenic program of subcutaneous white adipose tissue in mice. J. Clin. Invest. 121, 96–105 (2011).
Chi, J. et al. Three-dimensional adipose tissue imaging reveals regional variation in beige fat biogenesis and PRDM16-dependent sympathetic neurite density. Cell Metab. 27, 226–236 (2018).
Zeng, X. et al. Lysine-specific demethylase 1 promotes brown adipose tissue thermogenesis via repressing glucocorticoid activation. Genes Dev. 30, 1822–1836 (2016).
Pettem, K. L. et al. The specific α-neurexin interactor calsyntenin-3 promotes excitatory and inhibitory synapse development. Neuron 80, 113–128 (2013).
Saunders, A., Johnson, C. A. & Sabatini, B. L. Novel recombinant adeno-associated viruses for Cre activated and inactivated transgene expression in neurons. Front. Neural Circuits 6, 47 (2012).
Nakamura, K. et al. Identification of sympathetic premotor neurons in medullary raphe regions mediating fever and other thermoregulatory functions. J. Neurosci. 24, 5370–5380 (2004).
Cheng, L. et al. Identification of spinal circuits involved in touch-evoked dynamic mechanical pain. Nat. Neurosci. 20, 804–814 (2017).
Reeves, R. H. et al. Astrocytosis and axonal proliferation in the hippocampus of S100b transgenic mice. Proc. Natl Acad. Sci. USA 91, 5359–5363 (1994).
Winningham-Major, F., Staecker, J. L., Barger, S. W., Coats, S. & Van Eldik, L. J. Neurite extension and neuronal survival activities of recombinant S100β proteins that differ in the content and position of cysteine residues. J. Cell Biol. 109, 3063–3071 (1989).
Nishiyama, H., Knöpfel, T., Endo, S. & Itohara, S. Glial protein S100B modulates long-term neuronal synaptic plasticity. Proc. Natl Acad. Sci. USA 99, 4037–4042 (2002).
Steiner, G., Loveland, M. & Schonbaum, E. Effect of denervation on brown adipose tissue metabolism. Am. J. Physiol. 218, 566–570 (1970).
Bachman, E. S. et al. βAR signaling required for diet-induced thermogenesis and obesity resistance. Science 297, 843–845 (2002).
Dulloo, A. G. & Miller, D. S. Energy balance following sympathetic denervation of brown adipose tissue. Can. J. Physiol. Pharmacol. 62, 235–240 (1984).
Cao, Y., Wang, H. & Zeng, W. Whole-tissue 3D imaging reveals intra-adipose sympathetic plasticity regulated by NGF-TrkA signal in cold-induced beiging. Protein Cell 9, 527–539 (2018).
Donato, R. et al. S100B’s double life: intracellular regulator and extracellular signal. Biochim. Biophys. Acta 1793, 1008–1022 (2009).
Nisoli, E., Tonello, C., Benarese, M., Liberini, P. & Carruba, M. O. Expression of nerve growth factor in brown adipose tissue: implications for thermogenesis and obesity. Endocrinology 137, 495–503 (1996).
Sornelli, F., Fiore, M., Chaldakov, G. N. & Aloe, L. Adipose tissue-derived nerve growth factor and brain-derived neurotrophic factor: results from experimental stress and diabetes. Gen. Physiol. Biophys. 28, 179–183 (2009).
Chen, Y. et al. Crosstalk between KCNK3-mediated ion current and adrenergic signaling regulates adipose thermogenesis and obesity. Cell 171, 836–848.e13 (2017).
Long, J. Z. et al. A smooth muscle-like origin for beige adipocytes. Cell Metab. 19, 810–820 (2014).
Lam, S. S. et al. Directed evolution of APEX2 for electron microscopy and proximity labeling. Nat. Methods 12, 51–54 (2015).
Simpson, I. A. et al. Insulin-stimulated translocation of glucose transporters in the isolated rat adipose cells: characterization of subcellular fractions. Biochim. Biophys. Acta 763, 393–407 (1983).
Acknowledgements
We thank Nikon Imaging Center at Harvard Medical School for all imaging studies; RIKEN Institute for sharing the S100b knockout strain; Z. Herbert and the Molecular Biology Core Facilities at Dana Farber Cancer Institute for sequencing studies; the Rodent Histology Core at Harvard Medical School for histology studies; the EM Core at Harvard Medical School for APEX2 imaging studies; the viral core at Children’s Hospital Boston for AAV production; the transgenic core at Beth Israel Deaconess Medical Center for generation of mouse models; Y. Zhu for advice on sequencing data analysis. X.Z. was supported by the American Heart Association postdoctoral fellowship. B.H. is a Cancer Research Institute/Leonard Kahn Foundation Fellow. D.D.G. is an investigator of the Howard Hughes Medical Institute. This study was supported by NIH grant DK31405 to B.M.S.
Author information
Authors and Affiliations
Contributions
X.Z. conceived the project and designed experiments. X.Z., M.Y. and B.H. performed imaging experiments and data analysis. X.Z. and B.H. performed metabolic assays. J.M.R. performed stereotaxical surgeries, viral injections and post hoc histological analysis for the chemogenetic experiment. M.P.J. performed mass spectrometry analysis. B.B.L. supervised the chemogenetic experiments. D.D.G. supervised analysis of sympathetic innervation. B.M.S. supervised the entire project. X.Z. and B.M.S. wrote the manuscript with discussion and contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Clstn3b encodes an adipocyte-specific protein.
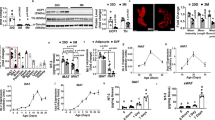
a, Quantitative PCR analysis of Clstn3b expression in wild-type and Lsd1-knockout BAT (n = 3 mice). b, Histone marker and transcription regulator ChIP–seq at the Clstn3 locus from BAT. c, Quantitative PCR analysis of Clstn3b expression in inguinal subcutaneous WAT from mice acclimatized to room temperature or 4 °C (n = 4 mice). d, Mass spectrometry identification of CLSTN3β peptides. e, Conservation of CLSTN3β within the mammalian class. The red cross and green ticks indicates the absence and presence, respectively, of homologues of CLSTN3β in mammalian subclasses. f, Sequence alignment between the unique exon of Clstn3b from human, and a fragment, in an intron upstream of the penultimate exon of Clstn1, in the genome of Chinese softshell turtle. Note how the position of this fragment corresponds to the β-selective exon in Clstn3. All data are mean ± s.e.m. Statistical significance was calculated by unpaired Student’s two-sided t-test.
Extended Data Fig. 2 CLSTN3β localizes to the endoplasmic reticulum.
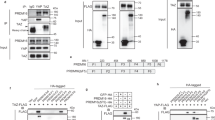
a, b, Electron microscopy analysis of primary brown adipocytes that express CLSTN3β–APEX2. In a, arrows denote the Golgi apparatus. In b, arrows denote peroxisomes. Scale bars, 100 nm. c, Western blot analysis of the fractionation pattern CLSTN3β. Asterisk denotes a nonspecific band. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 3 Ablation of Clstn3b impairs adipose thermogenesis.
a–d, Sanger sequencing (a), western blot (b), quantitative PCR (c) (n = 4 mice) and immunofluorescence (d) confirmation of CRISPR–Cas9 deletion of Clstn3b. Scale bars, 10 μm. e, Quantitative PCR analysis of Clstn3 expression in a panel of wild-type mouse tissues, and wild-type and Clstn3b-knockout brain (n = 2 mice for surveying tissue specificity in wild-type mouse; n = 3 mice for wild type and knockout). The primers target the junction between the third and the penultimate exons. f, g, Body weight curve (f) and body composition (g) of wild-type and Clstn3b-knockout mice on chow diet (n = 8 mice). h, Rates of CO2 production from indirect calorimetry analysis of wild-type and Clstn3b-knockout mice (n = 6 mice). i, j, Movement (i) and daily food intake (j) of wild-type and Clstn3b-knockout mice in metabolic chambers (n = 6 mice). k, Oxygen consumption response to acute β3 agonist injection, of wild-type and Clstn3b-knockout mice (n = 6 mice). All data are mean ± s.e.m. Statistical significance was calculated by unpaired Student’s two-sided t-test.
Extended Data Fig. 4 Transgenic expression of Clstn3b increases adipose thermogenesis.
a, b, Western blot (a) and quantitative PCR (b) confirmation of transgenic overexpression of CLSTN3β in BAT (n = 5 mice). c, d, Body-weight curve (c) and body composition (d) of wild-type and Clstn3b-transgenic mice on chow diet (n = 6 mice). e, Rates of CO2 production from indirect calorimetry analysis of wild-type and Clstn3b-transgenic mice (n = 4 mice). f, g, Movement (f) and daily food intake (g) of wild-type and Clstn3b-transgenic mice in metabolic chambers (n = 4 mice). h, Oxygen consumption response to acute β3 agonist injection of wild-type and Clstn3b-transgenic mice (n = 4 mice). All data are mean ± s.e.m. Statistical significance was calculated by unpaired Student’s two-sided t-test.
Extended Data Fig. 5 CLSTN3β increases sympathetic innervation of thermogenic adipose tissue.
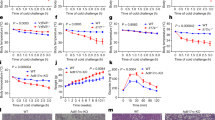
a, Gene expression analysis of wild-type and Clstn3b-knockout BAT upon 5 h of acute cold exposure, following mice being pre-acclimatized to thermoneutrality (n = 4 mice). Blue, wild-type; orange, knockout. b, Indirect calorimetry analysis of Clstn3b-knockout mice with or without Adipoq-cre, receiving AAV-DIO-Clstn3b injection (n = 4 mice). c, Whole-mount tyrosine hydroxylase staining of the inguinal region of the posterior subcutaneous WAT from wild-type and Clstn3b-knockout mice, acclimatized at 4 °C for 1 week. Scale bars, 50 μm. All data are mean ± s.e.m. Statistical significance was calculated by unpaired Student’s two-sided t-test.
Extended Data Fig. 6 CLSTN3β promotes secretion of S100b, an adipocyte-derived neurotrophic factor.
a, b, Quantitative PCR analysis of S100b expression in various fat depots (a) and in inguinal subcutaneous WAT (b), from mice acclimatized to room temperature or 4 °C (n = 4 mice). c, Quantitative PCR analysis of S100b expression in control or Prdm16-transgenic inguinal subcutaneous WAT (n = 4 mice). d, Quantitative PCR analysis of S100b expression in control or Prdm16-knockout inguinal subcutaneous WAT (n = 4 mice). e, PRDM16 ChIP–seq showing binding at the S100b locus. f, Indirect calorimetry analysis of Clstn3b-knockout mice with or without Adipoq-cre, receiving AAV-DIO-S100b injection (n = 4 mice). g, Tyrosine hydroxylase immunostaining of salivary gland from wild-type and S100b-knockout mice. h, Quantitative PCR analysis of S100b expression in wild-type and Clstn3b-knockout BAT from mice housed at room temperature (n = 4 mice). Note that this is a different housing condition from that used for experiments in Extended Data Fig. 5a. i, Western blot analysis of intracellular level of S100b in Clstn3b-knockout brown adipocytes that express S100b alone, or co-expressing S100b with CLSTN3β. j, Western blot analysis of S100b protein level in HEK293T cells transfected with various constructs as indicated. k, Western blot analysis of S100b and complement factor D secretion from HEK293T cells co-transfected with or without CLSTN3β. All data are mean ± s.e.m. Statistical significance was calculated by unpaired Student’s two-sided t-test.
Extended Data Fig. 7 Clstn3b is specifically expressed in human adipose tissue.
RNA sequencing in human tissues that shows adipose-specific expression of Clstn3b. RNA sequencing of 13 human tissue types was analysed for reads that uniquely map to the Clstn3b-specific exon.
Supplementary information
41586_2019_1156_MOESM3_ESM.xlsx
Supplementary Table 1 Proteomics analysis of WT and Clstn3β KO BAT from mice housed at RT (n=4 mice). Statistical significance was calculated by unpaired Student’s two-sided t-test.
Rights and permissions
About this article
Cite this article
Zeng, X., Ye, M., Resch, J.M. et al. Innervation of thermogenic adipose tissue via a calsyntenin 3β–S100b axis. Nature 569, 229–235 (2019). https://doi.org/10.1038/s41586-019-1156-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1156-9
This article is cited by
-
Control of lipolysis by a population of oxytocinergic sympathetic neurons
Nature (2024)
-
Thermogenic adipocyte-derived zinc promotes sympathetic innervation in male mice
Nature Metabolism (2023)
-
Transcriptional repression of beige fat innervation via a YAP/TAZ-S100B axis
Nature Communications (2023)
-
FAM210A is essential for cold-induced mitochondrial remodeling in brown adipocytes
Nature Communications (2023)
-
CLSTN3β enforces adipocyte multilocularity to facilitate lipid utilization
Nature (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.