Abstract
Intestinal organoids are complex three-dimensional structures that mimic the cell-type composition and tissue organization of the intestine by recapitulating the self-organizing ability of cell populations derived from a single intestinal stem cell. Crucial in this process is a first symmetry-breaking event, in which only a fraction of identical cells in a symmetrical sphere differentiate into Paneth cells, which generate the stem-cell niche and lead to asymmetric structures such as the crypts and villi. Here we combine single-cell quantitative genomic and imaging approaches to characterize the development of intestinal organoids from single cells. We show that their development follows a regeneration process that is driven by transient activation of the transcriptional regulator YAP1. Cell-to-cell variability in YAP1, emerging in symmetrical spheres, initiates Notch and DLL1 activation, and drives the symmetry-breaking event and formation of the first Paneth cell. Our findings reveal how single cells exposed to a uniform growth-promoting environment have the intrinsic ability to generate emergent, self-organized behaviour that results in the formation of complex multicellular asymmetric structures.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Code availability
Code used for image analysis in this study was developed in the Liberali laboratory in Matlab and Python 3. For Python 3, multiple open source Python libraries for scientific computing and image analysis were used. The code for organoid 2D and 3D segmentation, feature extraction and organoid linkage is available under https://github.com/fmi-basel/glib-nature2018-materials. An R implementation of the Griph algorithm for scRNA-seq analysis is available as an R package at https://github.com/ppapasaikas/Griph.
References
Sasai, Y. Cytosystems dynamics in self-organization of tissue architecture. Nature 493, 318–326 (2013).
Ohnishi, Y. et al. Cell-to-cell expression variability followed by signal reinforcement progressively segregates early mouse lineages. Nat. Cell Biol. 16, 27–37 (2014).
Goolam, M. et al. Heterogeneity in Oct4 and Sox2 targets biases cell fate in 4-cell mouse embryos. Cell 165, 61–74 (2016).
Hanna, J. et al. Direct cell reprogramming is a stochastic process amenable to acceleration. Nature 462, 595–601 (2009).
Eldar, A. & Elowitz, M. B. Functional roles for noise in genetic circuits. Nature 467, 167–173 (2010).
Snijder, B. et al. Population context determines cell-to-cell variability in endocytosis and virus infection. Nature 461, 520–523 (2009).
Battich, N., Stoeger, T. & Pelkmans, L. Control of transcript variability in single mammalian cells. Cell 163, 1596–1610 (2015).
Loewer, A. & Lahav, G. We are all individuals: causes and consequences of non-genetic heterogeneity in mammalian cells. Curr. Opin. Genet. Dev. 21, 753–758 (2011).
Snijder, B. & Pelkmans, L. Origins of regulated cell-to-cell variability. Nat. Rev. Mol. Cell Biol. 12, 119–125 (2011).
Wennekamp, S., Mesecke, S., Nédélec, F. & Hiiragi, T. A self-organization framework for symmetry breaking in the mammalian embryo. Nat. Rev. Mol. Cell Biol. 14, 452–459 (2013).
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
Spence, J. R. et al. Directed differentiation of human pluripotent stem cells into intestinal tissue in vitro. Nature 470, 105–109 (2011).
Sato, T. et al. Paneth cells constitute the niche for Lgr5 stem cells in intestinal crypts. Nature 469, 415–418 (2011).
Shoshkes-Carmel, M. et al. Subepithelial telocytes are an important source of Wnts that supports intestinal crypts. Nature 557, 242–246 (2018).
San Roman, A. K., Jayewickreme, C. D., Murtaugh, L. C. & Shivdasani, R. A. Wnt secretion from epithelial cells and subepithelial myofibroblasts is not required in the mouse intestinal stem cell niche in vivo. Stem Cell Reports 2, 127–134 (2014).
Farin, H. F. et al. Visualization of a short-range Wnt gradient in the intestinal stem-cell niche. Nature 530, 340–343 (2016).
van Es, J. H. et al. Dll1+ secretory progenitor cells revert to stem cells upon crypt damage. Nat. Cell Biol. 14, 1099–1104 (2012).
Tetteh, P. W. et al. Replacement of lost Lgr5-positive stem cells through plasticity of their enterocyte-lineage daughters. Cell Stem Cell 18, 203–213 (2016).
Yan, K.S. et al. intestinal enteroendocrine lineage cells possess homeostatic and injury-inducible stem cell activity. Cell Stem Cell 21, 78–90.76 (2017).
Tian, H. et al. A reserve stem cell population in small intestine renders Lgr5-positive cells dispensable. Nature 478, 255–259 (2011).
Gut, G., Herrmann, M. D. & Pelkmans, L. Multiplexed protein maps link subcellular organization to cellular states. Science 361, eaar7042 (2018).
Setty, M. et al. Wishbone identifies bifurcating developmental trajectories from single-cell data. Nat. Biotechnol. 34, 637–645 (2016).
Herring, C.A. et al. Unsupervised trajectory analysis of single-cell RNA-seq and imaging data reveals alternative tuft cell origins in the gut. Cell Syst. 6, 37–51 e39 (2018).
Zanconato, F. et al. Genome-wide association between YAP/TAZ/TEAD and AP-1 at enhancers drives oncogenic growth. Nat. Cell Biol. 17, 1218–1227 (2015).
Stein, C. et al. YAP1 exerts its transcriptional control via tead-mediated activation of enhancers. PLoS Genet. 11, e1005465 (2015).
Dupont, S. et al. Role of YAP/TAZ in mechanotransduction. Nature 474, 179–183 (2011).
Camargo, F. D. et al. YAP1 increases organ size and expands undifferentiated progenitor cells. Curr. Biol. 17, 2054–2060 (2007).
Gregorieff, A., Liu, Y., Inanlou, M. R., Khomchuk, Y. & Wrana, J. L. Yap-dependent reprogramming of Lgr5+ stem cells drives intestinal regeneration and cancer. Nature 526, 715–718 (2015).
Barry, E. R. et al. Restriction of intestinal stem cell expansion and the regenerative response by YAP. Nature 493, 106–110 (2013).
Azzolin, L. et al. YAP/TAZ incorporation in the β-catenin destruction complex orchestrates the Wnt response. Cell 158, 157–170 (2014).
Yui, S. et al. YAP/TAZ-dependent reprogramming of colonic epithelium links ECM remodeling to tissue regeneration. Cell Stem Cell 22, 35–49.e37 (2018).
Haber, A. L. et al. A single-cell survey of the small intestinal epithelium. Nature 551, 333–339 (2017).
Moor, A.E. et al. Spatial reconstruction of single enterocytes uncovers broad zonation along the intestinal villus axis. Cell 175, 1156–1167.e1115 (2018).
Fre, S. et al. Notch signals control the fate of immature progenitor cells in the intestine. Nature 435, 964–968 (2005).
Noah, T. K. & Shroyer, N. F. Notch in the intestine: regulation of homeostasis and pathogenesis. Annu. Rev. Physiol. 75, 263–288 (2013).
Yin, X. et al. Niche-independent high-purity cultures of Lgr5+ intestinal stem cells and their progeny. Nat. Methods 11, 106–112 (2014).
Totaro, A. et al. YAP/TAZ link cell mechanics to Notch signalling to control epidermal stem cell fate. Nat. Commun. 8, 15206 (2017).
Bray, S. J. Notch signalling in context. Nat. Rev. Mol. Cell Biol. 17, 722–735 (2016).
Chen, K. Y. et al. A Notch positive feedback in the intestinal stem cell niche is essential for stem cell self-renewal. Mol. Syst. Biol. 13, 927 (2017).
Carroll, T. D. et al. Interkinetic nuclear migration and basal tethering facilitates post-mitotic daughter separation in intestinal organoids. J. Cell Sci. 130, 3862–3877 (2017).
Gjorevski, N. et al. Designer matrices for intestinal stem cell and organoid culture. Nature 539, 560–564 (2016).
Sjoqvist, M. & Andersson, E. R. Do as I say, Not(ch) as I do: lateral control of cell fate. Dev. Biol. 447, 58–70 (2017).
Zhang, H., Pasolli, H. A. & Fuchs, E. Yes-associated protein (YAP) transcriptional coactivator functions in balancing growth and differentiation in skin. Proc. Natl Acad. Sci. USA 108, 2270–2275 (2011).
Xin, M. et al. Hippo pathway effector Yap promotes cardiac regeneration. Proc. Natl Acad. Sci. USA 110, 13839–13844 (2013).
Yimlamai, D. et al. Hippo pathway activity influences liver cell fate. Cell 157, 1324–1338 (2014).
Cebola, I. et al. TEAD and YAP regulate the enhancer network of human embryonic pancreatic progenitors. Nat. Cell Biol. 17, 615–626 (2015).
Panciera, T., Azzolin, L., Cordenonsi, M. & Piccolo, S. Mechanobiology of YAP and TAZ in physiology and disease. Nat. Rev. Mol. Cell Biol. 18, 758–770 (2017).
Brennecke, P. et al. Accounting for technical noise in single-cell RNA-seq experiments. Nat. Methods 10, 1093–1095 (2013).
Acknowledgements
We thank F. Maurer for organoid preparation, E. Tagliavini for IT support, G. de Medeiros for help in light-sheet imaging, A. Peters for light-sheet, L. Gelman for assistance and training, H. Kohler for sorting, S. Smallwood and S. Thiry for sequencing, R. Cuttat and S. Schuierer for scRNA-seq analysis, L. Pelkmans, J. Betschinger, D. Gilmour, L. Giorgetti, C. Tsiairis and laboratory members for reading the manuscript. Funding: SNSF (POOP3_157531 to P.L.). This work received funding from the ERC under the European Union’s Horizon 2020 research and innovation programme (grant agreement no. 758617).
Reviewer information
Nature thanks Anne Rios and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
P.L. conceived and supervised the study, P.L., D.S., U.M. and A.B. designed the experiments, D.S., U.M., L.C.M. and A.B. performed the time courses, I.L. performed the first YAP1 experiments, U.M. and A.B. designed the image analysis, M.R. and D.V. developed 3D and single-cell imaging, U.M. developed the trajectory inference, A.B. and P.S. built the light-sheet, A.B. and U.M. performed the light-sheet experiments, A.B. performed light-sheet image analysis, D.S. performed all transcriptomics experiments, M.B.S. designed transcription factor motifs analysis, D.S. and M.B.S. performed the analysis of RNA-seq data, M.B.S. and P.P. developed Griph, A.W. and G.R. performed the scRNA-seq analysis, U.M. performed the multiplexing experiments, P.L., D.S., U.M. and A.B. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
A.B. and P.S. are co-founders of Viventis Microscopy Sàrl that commercializes the light-sheet microscope used in this study.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Intestinal development of LGR5+ and LGR5− organoids.
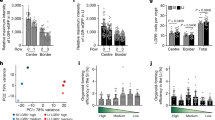
a, LGR5+ and LGR5− single cells sorted from LGR5::DTR-eGFP reporter mouse 3 h after FACS (left), with box plot showing LGR5::DTR-eGFP intensity quantification (right) (n = 602 organoids for LGR5+; n = 576 organoids for LGR5−). Box plot elements show quartiles, and whiskers denote 1.5× the interquartile range. b, FACS histogram of GFP signal of LGR5::DTR-eGFP single cells. Dashed boxes depict gating. c, Representative images of 4i imaging showing the same organoids stained with DAPI, GFP, lysozyme, proliferating cell nuclear antigen (PCNA), aldolaseB and Cell Trace. d, Distribution of eccentricity at 120 h of development for LGR5+ and LGR5− starting populations (LGR5+ n = 463 organoids; LGR5− n = 711 organoids). e, Representative images of LGR5+ and LGR5− cells at 120 h of development. Bottom, enlargements depict budding organoids and enterocysts. f, Representative images of enterocysts stained with DAPI, different cell-type and proliferation markers. g, Left, representative images of organoids grown from LGR5+ and LGR5− single cells in the presence of Wnt signalling inhibition (IWP-2, 2 µM) or activation (CHIR99021, 5 µM). Organoids are fixed at 120 h and stained for nuclear marker (DAPI) and enterocyte marker (aldolaseB). Top row shows complete well overview; bottom row shows magnified examples of single organoid. Right, quantification of enterocysts at 120 h of organoid development after Wnt signalling inhibition (IWP-2, 2 µM) or activation (CHIR99021, 2 µM) (n = 2 replicates). Data are mean values.
Extended Data Fig. 2 Feature selection for organoid development.
a, Pearson correlation heat map of the 14 selected features used for trajectory inference (n = 23,421 organoids). b, Fourteen selected features are grouped based on the underlying information. c, Comparison between eccentricities calculated on projected images and on 3D volumes (n = 16,175 organoids). d, Number of in-focus planes at different time points in cleared and uncleared organoids (n = 2,310 organoids). Planes are classified as in-focus with a logistic regression using the ratio of maximum in-plane intensity to maximum intensity in the whole stack as feature. Shading denotes s.d. e, Diffusion maps for pooled LGR5+ and LGR5− organoids (14 selected features, n = 23,421 organoids). Colour-coding denotes starting population and pseudotime. f, Diffusion maps for LGR5− (first column, 14 selected features, n = 13,623) and LGR5+ (second column, 14 selected features, n = 9,798) organoids, and for the pooled dataset with the full feature set (third column, 66 features, n = 23,421). Colour-coding denotes enterocytes (aldolaseB), PCNA, Paneth cells (lysozyme), organoid area, eccentricity and enterocyst classification (in which class 1 denotes enterocysts).
Extended Data Fig. 3 Cell types and proliferation markers along organoid development trajectories.
Proliferation (PCNA) and cell-type marker (lysozyme and aldolaseB) trends on pseudotime for LGR5+ (n = 9,798) and LGR5− (n = 13,623) organoids.
Extended Data Fig. 4 Dual illumination inverted light-sheet imaging of organoids.
a, Cross-sectional view of objective mounting block showing illumination paths (light blue dashed line) and imaging path (green dashed line). Illumination and imaging objective are solid mounted in the aluminium block and immersed in water contained in the reservoir (black arrow). b, Side view of microscope objectives arrangement: the sample embedded in Matrigel (red arrow) is held on the FEP membrane (black arrow) mounted in a sample holder and positioned between the two illumination objectives. c, Organoids handling procedure: grown organoids are dissociated and single cells are embedded in Matrigel. Several Matrigel drops can be transferred into the imaging chamber allowing imaging parallelization. d, Illustration of image improvement by double illumination. Scattering of single illumination beam by the organoid prevents complete and homogenous illumination (red arrows). This problem is minimized by double illumination. Scale bar, 25 µm. e, Representative images of an organoid expressing H2b–mCherry (red) and membrane-bound GFP (green). Slices every 20 µm across the organoid volume are shown. Scale bar, 25 µm. f, Workflow of light-sheet data analysis. g, Morphological features (major axis, area and volume) derived from light-sheet imaging. Budding organoids n = 6, enterocysts n = 3. h, Dynamic time-warp mapping of light-sheet data onto the trajectory. Budding organoid branch for mean area progression inferred from the trajectory (orange line, n = 23,421) and mean area progression extracted from time-lapse light-sheet imaging (violet line, n = 6) before (left) and after (right) morphing. Red dots indicate positions of real time (h) relative to pseudotime. Shading denotes s.d.
Extended Data Fig. 5 LGR5 dynamics during organoid development.
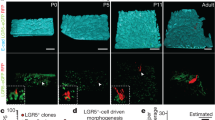
a, Representative images of organoids grown from LGR5+ and LGR5− starting populations at 0, 24 and 120 h of development. b, Cell types and proliferation markers plotted on pseudotime for LGR5+ (top, n = 9,798) and LGR5− (bottom, n = 13,623) starting populations. c, Light-sheet time-lapse imaging of organoid formation starting from a single LGR5+ cell (GFP signal) expressing membrane-bound mCherry (MEM9–mCherry). Green arrows are pointing to cells re-acquiring GFP signal from LGR5 reporter in the organoid crypts. d, Quantification of GFP signal of LGR5 reporter from time lapse shown in c. Cells localized in the organoid crypt are plotted in green, cells localized outside the crypt and on the main body are plotted in grey (n = 91 cells).
Extended Data Fig. 6 Bulk RNA-seq analysis.
a, Principal component analysis (PCA) analysis of time-course (0–120 h) bulk RNA-seq shows no mouse or batch effect (n = 3 biological replicates). b, Unsmoothed gene expression profile mapped on pseudotime trajectory of stem cell marker Olfm4. c, Unsmoothed gene expression profiles mapped on pseudotime trajectory of cell-type-specific genes. Paneth cell: Lyz1; ISC: Lgr5; enterocyte: Apoc3; goblet cell: Muc2; enteroendocrine cell: Sst. d, Left, protein abundance and gene expression profile of ISC marker Lgr5 mapped on pseudotime trajectory. Middle, protein abundance and gene expression profile of Paneth cell marker Lyz1 mapped on pseudotime trajectory. Right, protein abundance of enterocyte marker aldolaseB and gene expression profile of enterocyte marker Apoc3 mapped on pseudotime trajectory. e, Smoothed and unsmoothed gene expression profile of canonical Wnt target gene Axin2, Fzd2 and non-canonical target genes Nfatc2 and Rac1. f, Mean expression profile for each cluster and relative quantiles (0.05, 0.25, 0.50, 0.75 and 0.95). Annotation enrichments are shown on the right. E.S., enrichment score. g, Unsmoothed gene expression profile of Yap1, Tead4 and Ggta1 and top three transcription factors (Tead1, Tead4 and Fosl1) contributing to differential gene expression between 24 h and 0 h. Data in b, c, e, g are mean and s.d. (n = 3 biological replicates).
Extended Data Fig. 7 YAP1 is necessary but not sufficient for organoid formation.
a, Fraction of cells per organoid having YAP1 in the ON state (green, nuclear localization) or in the OFF state (red, cytoplasm localization) over time (n = 1,074 organoids). b, Entropy of YAP1 states (that is, variability in YAP1 activation in ON and OFF states) over time (n = 1,074 organoids). Shading denotes s.d. c, Quantification of YAP1 mean intensity after Wnt removal at 48 h or 72 h in organoids fixed at 72 h or 96 h (n = 4 replicates). Violin plot lines show quartiles. d, Images of organoids derived from single cells isolated from organoids at 72 h or at 120 h. e, Organoids treated at different time points with control or with verteporfin. Fixation at 96 h. Left, representative images; right, efficiency quantification (normalization: control, n = 2 replicates). Bar plots depict the mean. f, YAP1 overexpressing organoids with or without Wnt. Fixation at 96 h. Left, representative images; right, efficiency quantification (normalization: control, n = 4 replicates). P = 0.001, two-sided t-test. g, Left, images of organoids with Wnt removal at 48 h, or after double knockout of LATS1 and LATS2 (LATS DKO). Top, fixation at 72 h. Bottom, fixation at 96 h (Wnt removal at 48 h) or 120 h (LATS DKO). Right, quantification of enterocysts as retrieved in the left panel (n = 4 replicates for Wnt removal at 48 h; n = 3 replicates for LATS DKO). P = 0.0001 (Wnt removal); P = 0.002 (LATS DKO), two-sided t-test. h, Representative images of control, verteporfin treatment at 48 h, Wnt removal at 48 h, YAP1 overexpression, LATS DKO or treatment with EREG at 0 h. Top, fixation at 72 h; bottom, fixation at 96 h (Wnt removal at 48 h, YAP1 overexpression, LATS DKO) or 120 h (control, verteporfin 48 h, EREG). i, Scatter plot of Flag mean intensity signal and YAP1 mean intensity signal (left) and of Flag mean intensity and aldolaseB mean intensity (right) in YAP1-overexpressing organoids at 96 h. Data are mean ± s.d.
Extended Data Fig. 8 Griph robustness analysis and comparison to other methods.
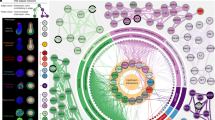
Comparison of Griph lower dimensional embedding (Griph/LargeVis) to different embedding approaches (PCA, PCA combined with distributed stochastic neighbor embedding (t-SNE), diffusion maps) and analysis of method sensitivity for variable gene selection. Five different sets of variable genes have been selected (using Griph to retain 10%, 25% or 50% of genes per bin, by Michaelis–Menten fitting of the gene dropout rates as implemented in M3Drop, or by the previously described mean-variance fitting procedure48) and analysed. a, b, First and second dimension are shown and results are colour-coded for enterocyte marker genes (as in Extended Data Fig. 9b) (a) and YAP1 target genes (as in Fig. 5b) (b). n = 1,863 cells.
Extended Data Fig. 9 scRNA analysis.
a, Experimental workflow for scRNA-seq. b, Griph-based visualization of single-cell degree of expression of marker genes for stem cells, Paneth cells and enterocytes. c, Griph-based visualization of single-cell degree of expression of marker genes for transient-amplifying cells, goblet cells, enteroendocrine cells, tuft cells, immature proximal enterocytes, immature distal enterocytes, transient-amplifying cells G1-phase, transient amplifying cells G2-phase, early progenitor enterocytes, late progenitor enterocytes, mature proximal enterocytes, mature distal enterocytes, enterocytes (villus bottom), enterocytes (villus middle), enterocytes (villus top) and enterocytes. d, Spearman correlation between expression of YAP1 target genes and expression of Dll1, Dll4, Jag1, Jag2 and Atoh1 in single cells at 120 h not expressing Paneth, goblet, enteroendocrine, enterocyte and stem-cell markers (n = 696 cells).
Extended Data Fig. 10 YAP1 cell-to-cell variability allows a Notch–DLL1 event and symmetry breaking.
a, Images showing YAP1 variability and DLL1+ cells. Magnified view denotes nuclear localization of HES1 in the same organoid. Arrows: red (DLL1+ cells), yellow (HES1+ cells). Scale bar, 10 µm. b, Images showing overlap of Paneth cells with DLL1+ cells. c, MIP images (top) and magnified single plane images (bottom) of: control, verteporfin addition at 48 h, Wnt removal at 48 h, LATS double-knockout (DKO) and EREG treatment. Organoids are fixed at 72 h. d, MIP images (top) and magnified single plane images (bottom) of: control, verteporfin addition at 48 h, Wnt removal at 48 h, YAP1 overexpression, LATS DKO and EREG treatment. Organoids are fixed at 72 h. e, Fraction of enterocysts (left) and fraction of organoids with Paneth cells (right) for control and organoids treated with Ly411575 or MK-0752 at 0 h. Fixation at 120 h (normalization: control, n = 4 replicates). Ly411575 P = 0.009; MK-0752 P = 0.003, two-sided t-test. Data are mean ± s.d. f, MIP images (top) and magnified single plane images (bottom) of organoids treated with control or DAPT at 120 h and fixed at 144 h and stained for AldoB (left) or LYZ (right) g, Top, annotation enrichment of genes correlated with expression of YAP1 target genes in cells expressing YAP1 target genes and Dll1, Dll4, Jag1, Jag2 and Atoh1 genes. Bottom, annotation enrichment of genes anti-correlated with YAP1 target genes expression in cells expressing YAP1 target genes and Dll1, Dll4, Jag1, Jag2 and Atoh1 genes. E.S., enrichment score. h, MIP images of organoids at 120 h, showing the canonical Wnt signalling response (TCF–GFP) in cells neighbouring Paneth cells.
Supplementary information
Supplementary Information
This file contains Supplementary Methods and additional references.
Supplementary Table 1
Bulk and single cell RNA sequencing analysis. Bulk RNA sequencing time course: Rpkm replicates, normalized mean expression, differential expression 24h-0h (n=3 biological replicates for each time point, statistical test used: Genewise Negative Binomial Generalized Linear Models With Quasi-Likelihood Tests, factors considered: time point, mouse and Dnase treatment) and Beta-coefficient for transcription factor binding motifs analysis. Single cell RNA sequencing: list of Yap1 target genes and intestinal epithelium cell types specific genes used within this study.
Supplementary Table 2
A list of primary and secondary antibodies used within this study.
Video 1: Time lapse Video of a budding organoid expressing H2b-mCherry as nuclear marker
A full stack is acquired every 10 minutes for around 5 days. Left panel –3D volume rendering shows that the whole organoid volume is imaged. Right panel – single plane intersecting the middle of the organoid volume. Experiments were repeated at least three times.
Video 2: Time lapse Video of an enterocyst expressing H2b-mCherry as nuclear marker (red) and membrane bound GFP (mem9-GFP, in green) imaged by light-sheet microscopy.
A full stack is acquired every 10 minutes for around 5 days. Left panel –3D volume rendering shows that the whole organoid volume is imaged. Right panel – single plane intersecting the middle of the organoid volume. Experiments were repeated at least three times.
Video 3: Left panel - 3D volume rendering of segmented light sheet data for one budding organoids and one enterocyst. Right panel – Dynamic plots of budding organoid and enterocyst volume and major axis over time.
Experiments were repeated at least three times.
Video 4: Time lapse Video of a budding organoid expressing Lgr5::GFP reporter (green) and membrane bound mCherry (mem9-mCherry, red) imaged by light-sheet microscopy.
A full stack is acquired every 30 minutes for around 5 days. Left panel –3D volume rendering shows that the whole organoid volume is imaged. Right panel – single plane intersecting the middle of the organoid volume. At the end of time lapse a slide through the all volume is shown where the presence of Lgr5+ cells in the crypt is visible. Experiments were repeated at least twice.
Source data
Rights and permissions
About this article
Cite this article
Serra, D., Mayr, U., Boni, A. et al. Self-organization and symmetry breaking in intestinal organoid development. Nature 569, 66–72 (2019). https://doi.org/10.1038/s41586-019-1146-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1146-y
This article is cited by
-
Hippo pathway in cell–cell communication: emerging roles in development and regeneration
Inflammation and Regeneration (2024)
-
Desulfovibrio vulgaris interacts with novel gut epithelial immune receptor LRRC19 and exacerbates colitis
Microbiome (2024)
-
Bioelectric stimulation controls tissue shape and size
Nature Communications (2024)
-
Open-top multisample dual-view light-sheet microscope for live imaging of large multicellular systems
Nature Methods (2024)
-
A patterned human neural tube model using microfluidic gradients
Nature (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.