Abstract
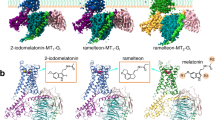
The human MT1 and MT2 melatonin receptors1,2 are G-protein-coupled receptors (GPCRs) that help to regulate circadian rhythm and sleep patterns3. Drug development efforts have targeted both receptors for the treatment of insomnia, circadian rhythm and mood disorders, and cancer3, and MT2 has also been implicated in type 2 diabetes4,5. Here we report X-ray free electron laser (XFEL) structures of the human MT2 receptor in complex with the agonists 2-phenylmelatonin (2-PMT) and ramelteon6 at resolutions of 2.8 Å and 3.3 Å, respectively, along with two structures of function-related mutants: H2085.46A (superscripts represent the Ballesteros–Weinstein residue numbering nomenclature7) and N862.50D, obtained in complex with 2-PMT. Comparison of the structures of MT2 with a published structure8 of MT1 reveals that, despite conservation of the orthosteric ligand-binding site residues, there are notable conformational variations as well as differences in [3H]melatonin dissociation kinetics that provide insights into the selectivity between melatonin receptor subtypes. A membrane-buried lateral ligand entry channel is observed in both MT1 and MT2, but in addition the MT2 structures reveal a narrow opening towards the solvent in the extracellular part of the receptor. We provide functional and kinetic data that support a prominent role for intramembrane ligand entry in both receptors, and suggest that there might also be an extracellular entry path in MT2. Our findings contribute to a molecular understanding of melatonin receptor subtype selectivity and ligand access modes, which are essential for the design of highly selective melatonin tool compounds and therapeutic agents.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Reppert, S. M., Weaver, D.R. & Ebisawa, T. Cloning and characterization of a mammalian melatonin receptor that mediates reproductive and circadian responses. Neuron 13, 1177–1185 (1994).
Reppert, S. M. et al. Molecular characterization of a second melatonin receptor expressed in human retina and brain: the Mel1b melatonin receptor. Proc. Natl Acad. Sci. USA 92, 8734–8738 (1995).
Liu, J. et al. MT1 and MT2 melatonin receptors: a therapeutic perspective. Annu. Rev. Pharmacol. Toxicol. 56, 361–383 (2016).
Bonnefond, A. et al. Rare MTNR1B variants impairing melatonin receptor 1B function contribute to type 2 diabetes. Nat. Genet. 44, 297–301 (2012).
Karamitri, A. et al. Type 2 diabetes-associated variants of the MT2 melatonin receptor affect distinct modes of signaling. Sci. Signal. 11, eaan6622 (2018).
Kato, K. et al. Neurochemical properties of ramelteon (TAK-375), a selective MT1/MT2 receptor agonist. Neuropharmacology 48, 301–310 (2005).
Ballesteros, J. A. & Weinstein, H. Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein-coupled receptors. Meth. Neurosci. 25, 366–428 (1995).
Stauch, B. et al. Structural basis of ligand recognition at the human MT1 melatonin receptor. Nature https://doi.org/10.1038/s41586-019-1141-3 (2019).
White, K. L. et al. Structural connection between activation microswitch and allosteric sodium site in GPCR signaling. Structure 26, 259–269.e5 (2018).
Roth, C. B., Hanson, M. A. & Stevens, R. C. Stabilization of the human β2-adrenergic receptor TM4–TM3–TM5 helix interface by mutagenesis of Glu1223.41, a critical residue in GPCR structure. J. Mol. Biol. 376, 1305–1319 (2008).
Chun, E. et al. Fusion partner toolchest for the stabilization and crystallization of G protein-coupled receptors. Structure 20, 967–976 (2012).
Audet, M. & Bouvier, M. Restructuring G-protein- coupled receptor activation. Cell 151, 14–23 (2012).
Caffrey, M. & Cherezov, V. Crystallizing membrane proteins using lipidic mesophases. Nat. Protocols 4, 706–731 (2009).
Rivara, S., Mor, M., Bedini, A., Spadoni, G. & Tarzia, G. Melatonin receptor agonists: SAR and applications to the treatment of sleep-wake disorders. Curr. Top. Med. Chem. 8, 954–968 (2008).
Bento, A. P. et al. The ChEMBL bioactivity database: an update. Nucleic Acids Res. 42, D1083–D1090 (2014).
Nonno, R. et al. A new melatonin receptor ligand with mt1-agonist and MT2-antagonist properties. J. Pineal Res. 29, 234–240 (2000).
Zlotos, D. P., Jockers, R., Cecon, E., Rivara, S. & Witt-Enderby, P. A. MT1 and MT2 melatonin receptors: ligands, models, oligomers, and therapeutic potential. J. Med. Chem. 57, 3161–3185 (2014).
Teh, M. T. & Sugden, D. Comparison of the structure-activity relationships of melatonin receptor agonists and antagonists: lengthening the N-acyl side-chain has differing effects on potency on Xenopus melanophores. Naunyn Schmiedebergs Arch. Pharmacol. 358, 522–528 (1998).
Valentin-Hansen, L. et al. The arginine of the DRY motif in transmembrane segment III functions as a balancing micro-switch in the activation of the β2-adrenergic receptor. J. Biol. Chem. 287, 31973–31982 (2012).
Spadoni, G. et al. Highly potent and selective MT2 melatonin receptor full agonists from conformational analysis of 1-benzyl-2-acylaminomethyl-tetrahydroquinolines. J. Med. Chem. 58, 7512–7525 (2015).
UniProt Consortium, T. UniProt: the universal protein knowledgebase. Nucleic Acids Res. 46, 2699 (2018).
Alexandrov, A. I., Mileni, M., Chien, E. Y., Hanson, M. A. & Stevens, R. C. Microscale fluorescent thermal stability assay for membrane proteins. Structure 16, 351–359 (2008).
Liu, W. et al. Serial femtosecond crystallography of G protein-coupled receptors. Science 342, 1521–1524 (2013).
Weierstall, U. et al. Lipidic cubic phase injector facilitates membrane protein serial femtosecond crystallography. Nat. Commun. 5, 3309 (2014).
Boutet, S. W. G. J. The Coherent X-ray Imaging (CXI) instrument at the Linac Coherent Light Source (LCLS). New J. Phys. 12, 035024 (2010).
Hart, P. et al. The CSPAD Megapixel X-ray Camera at LCLS. In X-ray Free-electron Lasers: Beam Diagnostics, Beamline Instrumentation and Applications (eds. S. P. Moeller, S. P. et al.) 85040C–85012 (2012).
Barty, A. et al. Cheetah: software for high-throughput reduction and analysis of serial femtosecond X-ray diffraction data. J. Appl. Crystallogr. 47, 1118–1131 (2014).
Battye, T. G., Kontogiannis, L., Johnson, O., Powell, H. R. & Leslie, A. G. iMOSFLM: a new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallogr. D 67, 271–281 (2011).
Duisenberg, A. J. M. Indexing in single-crystal diffractometry with an obstinate list of reflections. J. Appl. Crystallogr. 25, 92–96 (1992).
Kabsch, W. Xds. Acta Crystallogr. D 66, 125–132 (2010).
White, T. A. et al. Recent developments in CrystFEL. J. Appl. Crystallogr. 49, 680–689 (2016).
Karplus, P. A. & Diederichs, K. Linking crystallographic model and data quality. Science 336, 1030–1033 (2012).
Zimmermann, L. et al. A completely reimplemented MPI bioinformatics toolkit with a new HHpred server at its core. J. Mol. Biol. 430, 2237–2243 (2018).
Bunkóczi, G. & Read, R. J. Improvement of molecular-replacement models with Sculptor. Acta Crystallogr. D 67, 303–312 (2011).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Zheng, Y. et al. Structure of CC chemokine receptor 2 with orthosteric and allosteric antagonists. Nature 540, 458–461 (2016).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D 68, 352–367 (2012).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Tan, Q. et al. Structure of the CCR5 chemokine receptor–HIV entry inhibitor maraviroc complex. Science 341, 1387–1390 (2013).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997).
BUSTER v. 2.10.2.
Schüttelkopf, A. W. & van Aalten, D. M. PRODRG: a tool for high-throughput crystallography of protein–ligand complexes. Acta Crystallogr. D 60, 1355–1363 (2004).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
The PyMOL Molecular Graphics System. Version 2.0 Schrödinger, LLC.
Jurcik, A. et al. CAVER Analyst 2.0: analysis and visualization of channels and tunnels in protein structures and molecular dynamics trajectories. Bioinformatics 34, 3586–3588 (2018).
Abagyan, R. A., Totrov, M. M. & Kuznetsov, D. A. ICM: a new method for protein modeling and design: applications to docking and structure prediction from the distorted native conformation. J. Comput. Chem. 15, 488–506 (1994).
Acknowledgements
We thank M. Chu, C. Hanson, K. Villers, and J. Velasquez for help with cloning and expression, T. Grant for XFEL data processing, and H. Shaye for technical support. This work was supported by the National Institutes of Health grants R35 GM127086 (V.C.), R21 DA042298 (W.L.), R01 GM124152 (W.L.), U24DK116195 (B.L.R.), R01MH112205 (B.L.R.), the NIMH Psychoactive Drug Screening Program and the Michael Hooker Distinguished Professorship to B.L.R. and F31-NS093917 (R.H.J.O.), the STC Program of the National Science Foundation (NSF) through BioXFEL (No. 1231306) (B.S., U.W., W.L., N.A.Z., V.C.), NSF ABI grant 1565180 (C.L, N.Z., U.W.), HFSP long-term fellowship LT000046/2014-L (L.C.J.), postdoctoral fellowship from the Swedish Research Council (L.C.J.) and EMBO ALTF 677-2014 (B.S.). Parts of this research were carried out at the LCLS, a National User Facility operated by Stanford University on behalf of the US Department of Energy and supported by the US Department of Energy Office of Science, Office of Basic Energy Sciences under Contract No. DE-AC02-76SF00515. This research benefited from the use of credits from the National Institutes of Health (NIH) Cloud Credits Model Pilot, a component of the NIH Big Data to Knowledge (BD2K) program.
Reviewer information
Nature thanks Christian Siebold, Ieva Sutkeviciute, Jean-Pierre Vilardaga and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
L.C.J., B.S., V.K., and V.C. conceived the project, analysed data, and wrote the paper with contributions from all authors. L.C.J. and B.S. designed, optimized, purified and characterized receptor constructs for structural studies and crystallized the receptor, prepared crystal samples and figures, solved and refined the structures, and assisted in generating mutant constructs for binding and functional analyses. J.D.M., X.-P.H., and S.T.S. performed radioligand binding and functional experiments, assisted in making mutant and wild-type constructs, and analysed binding and functional data. L.C.J., B.S., A.B., L.Z., W.L., and V.C. collected XFEL data. S.Z. and U.W. operated the LCP injector during XFEL data collection. S.H., L.Z., and W.L. assisted in XFEL sample preparation. A.B., C.G., C.L., and N.A.Z. processed XFEL data. L.C.J., G.W.H. and A.B. determined the structures. L.C.J. and G.W.H. performed the structure refinement and quality control. N.P., J.M.G., and V.K. designed mutants for radioligand and functional studies, and performed molecular docking and molecular dynamics simulations. R.H.J.O. and A.R.T. assisted with molecular biology and functional experiments. S.Y. synthesized the bitopic compound, analysed data, and edited the paper. R.C.S. contributed to study design and selection of chemical compounds for receptor stabilization and functional characterization, supervised protein expression, and edited the paper. B.L.R. supervised pharmacological experiments and edited the paper. V.K. supervised molecular docking and molecular dynamics calculations. V.C. coordinated and supervised the whole project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Crystallization of MT2: crystals, crystal packing and electron density.
a, b, Bright field (a) and cross-polarized images (b) of representative MT2–2-PMT crystals optimized for synchrotron data collection (representing three independent crystallization setups). c, Cross-polarized image of representative MT2(N86D)–2-PMT crystals used for XFEL data collection (representing three independent crystallization setups). See Extended Data Table 2 for data collection statistics. d, e, Crystal packing (receptor, purple; BRIL, green; rubredoxin, blue). Space for missing rubredoxin in molecule B of the asymmetric unit is indicated with a red circle. Lattice rotated 90° is shown in e. f, Overlay of 2-PMT (purple) and ramelteon (blue) ligands of MT2. g–i, 2mFo – DFc density (grey) contoured at 1σ of ramelteon (g), N862.50D mutation (h), and H2085.46A mutation (i). 2-PMT is shown in purple.
Extended Data Fig. 2 Structural differences between MT1 and MT2.
a, Overlay of MT1–2-PMT (green) and MT2–2-PMT (violet) structures (Cα r.m.s.d = 0.6 Å). b, Comparison of MT1 (green) and MT2 (violet) binding pockets. Overall, the binding pocket in MT2 is about 50 Å3 larger than in MT1. c, Comparison of 2-PMT ligand conformations in MT1 (green) and MT2 (violet). Hydrogen bonds are shown as yellow dashed lines. d, Overlay of MT1 and MT2, showing residues with different conformations in the vicinity of the binding pocket. N4.60 makes a hydrogen bond with Y5.38 in MT2 but not in MT1.
Extended Data Fig. 3 Molecular dynamics simulations.
a, b, Distance plots for interactions between residues in MT2 (N1754.60, atom ND2; Q194ECL2, atom NE2; N2686.52, atom ND2), and closest oxygen atoms of the ligand methoxy and acetyl groups, respectively, in complexes with melatonin (a) and 2-PMT (b) from three independent simulation runs. c, Distance histograms for interactions of N1754.60 with methoxy of melatonin (yellow) and 2-PMT (violet). d, Distance histograms for interactions of Q194ECL2 with alkylamide tail of melatonin (yellow) and 2-PMT (violet).
Extended Data Fig. 4 Structural and functional differences between MT2–2-PMT and MT2(H208A5.46)–2-PMT.
a, Overlay of the MT2–2-PMT (purple) structure with that of MT2(H2085.46A)–2-PMT (grey) reveals an inward shift of helix V of about 0.9 Å, due to the H2085.46A mutation (black arrow). b, Surface representation of the H2085.46 and H2085.46A residues. Rotation of helix V renders the binding pocket volume about 50 Å3 smaller for the H2085.46A structure (binding site volume for MT2–2-PMT: 766 Å3 compared to 716 Å3 for the MT2(H2085.46A)–2-PMT structure). c, Comparison of the channel profiles (from the outside of the protein towards the ligand) for MT2–2-PMT (purple) and MT2(H2085.46A)–2-PMT (grey) reveals a narrowing of the MT2(H2085.46A)–2-PMT channel at a depth of around 6 Å as a consequence of the mutation and subsequent inward rotation of helix V. d, Functional data for wild-type and H2085.46A mutant MT2 expressed in HEK293T cells by using GloSensor to measure Gi/o-mediated inhibition of cAMP. Data represent mean ± s.e.m. for n independent experiments as indicated in square brackets. %EMAX (maximum effect of a drug) is relative to wild-type receptor (in columns), and %EMAX∗ is relative to melatonin activity (in rows). See Methods for further information and Supplementary Fig. 6 for dose–response curves.
Extended Data Fig. 5 Selectivity analysis of melatonergic compounds.
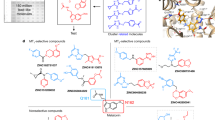
a, Binding affinities of ligands for MT1 (ChEMBL target identifier CHEMBL1945) and MT2 (CHEMBL1946) were retrieved from the ChEMBL database15 (v. 24) of experimental literature values. Of these ligands, 525 have affinities reported for both receptor subtypes. For ligands with multiple reported affinity values for a given receptor, pKi values were averaged. MT1-selective ligands are in the lower right quadrant; MT2-selective ligands are in the upper left quadrant. Data points are coloured by absolute pKi difference between subtypes (that is, selectivity). b, Histogram of observed ligand selectivities. MT2 selective ligands are on the left of the panel, MT1 selective ligands are on the right. c, Plot of the docking score difference of select ligands that were docked between MT2 and MT1 versus their pKi difference (MT2 – MT1). Dashed lines indicate pKi selectivity cutoff criteria (MT1: −1 and MT2: 4). Data points are coloured by molecular mass (Da). See Supplementary Table 1 for details of docked ligands.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-7.
Supplementary Table 1
A table of docked ligands plotted in Extended Data Fig. 5c. MW, molecular weight in Dalton (Da), pKi values in nM. Docking scores are expressed in arbitrary units.
Rights and permissions
About this article
Cite this article
Johansson, L.C., Stauch, B., McCorvy, J.D. et al. XFEL structures of the human MT2 melatonin receptor reveal the basis of subtype selectivity. Nature 569, 289–292 (2019). https://doi.org/10.1038/s41586-019-1144-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1144-0
This article is cited by
-
Protective effects of melatonin on myocardial microvascular endothelial cell injury under hypertensive state by regulating Mst1
BMC Cardiovascular Disorders (2023)
-
Structural basis of hydroxycarboxylic acid receptor signaling mechanisms through ligand binding
Nature Communications (2023)
-
Serial femtosecond crystallography
Nature Reviews Methods Primers (2022)
-
Universal platform for the generation of thermostabilized GPCRs that crystallize in LCP
Nature Protocols (2022)
-
Structural basis of the ligand binding and signaling mechanism of melatonin receptors
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.