Abstract
The serotonin transporter (SERT) regulates neurotransmitter homeostasis through the sodium- and chloride-dependent recycling of serotonin into presynaptic neurons1,2,3. Major depression and anxiety disorders are treated using selective serotonin reuptake inhibitors—small molecules that competitively block substrate binding and thereby prolong neurotransmitter action2,4. The dopamine and noradrenaline transporters, together with SERT, are members of the neurotransmitter sodium symporter (NSS) family. The transport activities of NSSs can be inhibited or modulated by cocaine and amphetamines2,3, and genetic variants of NSSs are associated with several neuropsychiatric disorders including attention deficit hyperactivity disorder, autism and bipolar disorder2,5. Studies of bacterial NSS homologues—including LeuT—have shown how their transmembrane helices (TMs) undergo conformational changes during the transport cycle, exposing a central binding site to either side of the membrane1,6,7,8,9,10,11,12. However, the conformational changes associated with transport in NSSs remain unknown. To elucidate structure-based mechanisms for transport in SERT we investigated its complexes with ibogaine, a hallucinogenic natural product with psychoactive and anti-addictive properties13,14. Notably, ibogaine is a non-competitive inhibitor of transport but displays competitive binding towards selective serotonin reuptake inhibitors15,16. Here we report cryo-electron microscopy structures of SERT–ibogaine complexes captured in outward-open, occluded and inward-open conformations. Ibogaine binds to the central binding site, and closure of the extracellular gate largely involves movements of TMs 1b and 6a. Opening of the intracellular gate involves a hinge-like movement of TM1a and the partial unwinding of TM5, which together create a permeation pathway that enables substrate and ion diffusion to the cytoplasm. These structures define the structural rearrangements that occur from the outward-open to inward-open conformations, and provide insight into the mechanism of neurotransmitter transport and ibogaine inhibition.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon request. The coordinates for the 15B8 X-ray structure have been deposited in the Protein Data Bank under the accession code 6D9G. The coordinates and associated volumes for the cryo-EM reconstruction of ts2-inactive Fab–scFv paroxetine, ts2-active Fab–scFv ibogaine-outward, ΔN72/C13 Fab ibogaine-occluded, and ΔN72/C13 Fab ibogaine-inward-open datasets have been deposited in the PDB and Electron Microscopy Data Bank (EMDB) under the accession codes 6DZW and 8941, 6DZY and 8942, 6DZV and 8940, and 6DZZ and 8943, respectively. The volume for the cryo-EM reconstruction of the ΔN72/C13 Fab noribogaine-inward-open reconstruction has been deposited in the EMDB under accession code 0437. The half maps and masks used for refinement for each dataset have also been deposited in the EMDB.
References
Singh, S. K. & Pal, A. Biophysical approaches to the study of LeuT, a prokaryotic homolog of neurotransmitter sodium symporters. Methods Enzymol. 557, 167–198 (2015).
Kristensen, A. S. et al. SLC6 neurotransmitter transporters: structure, function, and regulation. Pharmacol. Rev. 63, 585–640 (2011).
Gether, U., Andersen, P. H., Larsson, O. M. & Schousboe, A. Neurotransmitter transporters: molecular function of important drug targets. Trends Pharmacol. Sci. 27, 375–383 (2006).
Cipriani, A. et al. Comparative efficacy and acceptability of 21 antidepressant drugs for the acute treatment of adults with major depressive disorder: a systematic review and network meta-analysis. Lancet 391, 1357–1366 (2018).
Lin, Z. & Madras, B. K. in Neurotransmitter Transporters. Handbook of Experimental Pharmacology Vol. 175 (eds Sitte, H. H. & Freissmuth, M.) 327–371 (Springer, Berlin, Heidelberg, 2006).
Kazmier, K. et al. Conformational dynamics of ligand-dependent alternating access in LeuT. Nat. Struct. Mol. Biol. 21, 472–479 (2014).
Malinauskaite, L. et al. A mechanism for intracellular release of Na+ by neurotransmitter/sodium symporters. Nat. Struct. Mol. Biol. 21, 1006–1012 (2014).
Krishnamurthy, H. & Gouaux, E. X-ray structures of LeuT in substrate-free outward-open and apo inward-open states. Nature 481, 469–474 (2012).
Merkle, P. S. et al. Substrate-modulated unwinding of transmembrane helices in the NSS transporter LeuT. Sci. Adv. 4, eaar6179 (2018).
Grouleff, J., Søndergaard, S., Koldsø, H. & Schiøtt, B. Properties of an inward-facing state of LeuT: conformational stability and substrate release. Biophys. J. 108, 1390–1399 (2015).
Yamashita, A., Singh, S. K., Kawate, T., Jin, Y. & Gouaux, E. Crystal structure of a bacterial homologue of Na+/Cl−-dependent neurotransmitter transporters. Nature 437, 215–223 (2005).
Terry, D. S. et al. A partially-open inward-facing intermediate conformation of LeuT is associated with Na+ release and substrate transport. Nat. Commun. 9, 230 (2018).
Belgers, M. et al. Ibogaine and addiction in the animal model, a systematic review and meta-analysis. Transl. Psychiatry 6, e826 (2016).
Dybowski, J. & Landrin, E. Concerning Iboga, its excitement-producing properties, its composition, and the new alkaloid it contains, ibogaine. C.R. Acad. Sci. 133, 748 (1901).
Bulling, S. et al. The mechanistic basis for noncompetitive ibogaine inhibition of serotonin and dopamine transporters. J. Biol. Chem. 287, 18524–18534 (2012).
Jacobs, M. T., Zhang, Y. W., Campbell, S. D. & Rudnick, G. Ibogaine, a noncompetitive inhibitor of serotonin transport, acts by stabilizing the cytoplasm-facing state of the transporter. J. Biol. Chem. 282, 29441–29447 (2007).
Wu, S. et al. Fabs enable single particle cryoEM studies of small proteins. Structure 20, 582–592 (2012).
Coleman, J. A. & Gouaux, E. Structural basis for recognition of diverse antidepressants by the human serotonin transporter. Nat. Struct. Mol. Biol. 25, 170–175 (2018).
Zhang, Y. W. & Rudnick, G. The cytoplasmic substrate permeation pathway of serotonin transporter. J. Biol. Chem. 281, 36213–36220 (2006).
Burtscher, V., Hotka, M., Li, Y., Freissmuth, M. & Sandtner, W. A label-free approach to detect ligand binding to cell surface proteins in real time. eLife 7, e34944 (2018).
Tavoulari, S., Forrest, L. R. & Rudnick, G. Fluoxetine (Prozac) binding to serotonin transporter is modulated by chloride and conformational changes. J. Neurosci. 29, 9635–9643 (2009).
Ramamoorthy, S., Samuvel, D. J., Buck, E. R., Rudnick, G. & Jayanthi, L. D. Phosphorylation of threonine residue 276 is required for acute regulation of serotonin transporter by cyclic GMP. J. Biol. Chem. 282, 11639–11647 (2007).
Zhang, Y. W., Turk, B. E. & Rudnick, G. Control of serotonin transporter phosphorylation by conformational state. Proc. Natl Acad. Sci. USA 113, E2776–E2783 (2016).
Penmatsa, A., Wang, K. H. & Gouaux, E. X-ray structure of dopamine transporter elucidates antidepressant mechanism. Nature 503, 85–90 (2013).
Wang, K. H., Penmatsa, A. & Gouaux, E. Neurotransmitter and psychostimulant recognition by the dopamine transporter. Nature 521, 322–327 (2015).
Laursen, L. et al. Cholesterol binding to a conserved site modulates the conformation, pharmacology, and transport kinetics of the human serotonin transporter. J. Biol. Chem. 293, 3510–3523 (2018).
Mash, D. C., Staley, J. K., Baumann, M. H., Rothman, R. B. & Hearn, W. L. Identification of a primary metabolite of ibogaine that targets serotonin transporters and elevates serotonin. Life Sci. 57, PL45–PL50 (1995).
Coleman, J. A., Green, E. M. & Gouaux, E. X-ray structures and mechanism of the human serotonin transporter. Nature 532, 334–339 (2016).
Chen, F. et al. Characterization of an allosteric citalopram-binding site at the serotonin transporter. J. Neurochem. 92, 21–28 (2005).
Navratna, V., Tosh, D. K., Jacobson, K. A. & Gouaux, E. Thermostabilization and purification of the human dopamine transporter (hDAT) in an inhibitor and allosteric ligand bound conformation. PLoS ONE 13, e0200085 (2018).
Hasenhuetl, P. S., Freissmuth, M. & Sandtner, W. Electrogenic binding of intracellular cations defines a kinetic decision point in the transport cycle of the human serotonin transporter. J. Biol. Chem. 291, 25864–25876 (2016).
Felts, B. et al. The two Na+ sites in the human serotonin transporter play distinct roles in the ion coupling and electrogenicity of transport. J. Biol. Chem. 289, 1825–1840 (2014).
Blat, Y. Non-competitive inhibition by active site binders. Chem. Biol. Drug Des. 75, 535–540 (2010).
Coleman, J. A., Green, E. M. & Gouaux, E. Thermostabilization, expression, purification, and crystallization of the human serotonin transporter bound to S-citalopram. J. Vis. Exp. 117, e54792 (2016).
Green, E. M., Coleman, J. A. & Gouaux, E. Thermostabilization of the human serotonin transporter in an antidepressant-bound conformation. PLoS ONE 10, e0145688 (2015).
Goehring, A. et al. Screening and large-scale expression of membrane proteins in mammalian cells for structural studies. Nat. Protoc. 9, 2574–2585 (2014).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Voss, N. R., Yoshioka, C. K., Radermacher, M., Potter, C. S. & Carragher, B. DoG Picker and TiltPicker: software tools to facilitate particle selection in single particle electron microscopy. J. Struct. Biol. 166, 205–213 (2009).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Grant, T., Rohou, A. & Grigorieff, N. cisTEM, user-friendly software for single-particle image processing. eLife 7, e35383 (2018).
Zhang, C. et al. Analysis of discrete local variability and structural covariance in macromolecular assemblies using Cryo-EM and focused classification. Ultramicroscopy https://doi.org/10.1016/j.ultramic.2018.11.016 (2018).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Wang, R. Y. et al. Automated structure refinement of macromolecular assemblies from cryo-EM maps using Rosetta. eLife 5, e17219 (2016).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
The PyMOL Molecular Graphics System v.2.0 (Schrödinger, 2017).
Chovancova, E. et al. CAVER 3.0: a tool for the analysis of transport pathways in dynamic protein structures. PLOS Comput. Biol. 8, e1002708 (2012).
Afonine, P. V. et al. New tools for the analysis and validation of cryo-EM maps and atomic models. Acta Crystallogr. D 74, 814–840 (2018).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Schreiner, E., Trabuco, L. G., Freddolino, P. L. & Schulten, K. Stereochemical errors and their implications for molecular dynamics simulations. BMC Bioinformatics 12, 190 (2011).
Olsson, M. H., Søndergaard, C. R., Rostkowski, M. & Jensen, J. H. PROPKA3: consistent treatment of internal and surface residues in empirical pK a predictions. J. Chem. Theory Comput. 7, 525–537 (2011).
Lomize, M. A., Pogozheva, I. D., Joo, H., Mosberg, H. I. & Lomize, A. L. OPM database and PPM web server: resources for positioning of proteins in membranes. Nucleic Acids Res. 40, D370–D376 (2012).
Vanommeslaeghe, K. et al. CHARMM general force field: A force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
Vanommeslaeghe, K., Raman, E. P. & MacKerell, A. D., Jr. Automation of the CHARMM General Force Field (CGenFF) II: assignment of bonded parameters and partial atomic charges. J. Chem. Inf. Model. 52, 3155–3168 (2012).
Mayne, C. G., Saam, J., Schulten, K., Tajkhorshid, E. & Gumbart, J. C. Rapid parameterization of small molecules using the force field toolkit. J. Comput. Chem. 34, 2757–2770 (2013).
Vanommeslaeghe, K. & MacKerell, A. D., Jr. Automation of the CHARMM General Force Field (CGenFF) I: bond perception and atom typing. J. Chem. Inf. Model. 52, 3144–3154 (2012).
Gaussian 09 v.A.02 (Gaussian, 2016).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Beauchamp, K. A. et al. MSMBuilder2: modeling conformational dynamics at the picosecond to millisecond scale. J. Chem. Theory Comput. 7, 3412–3419 (2011).
Gumbart, J., Trabuco, L. G., Schreiner, E., Villa, E. & Schulten, K. Regulation of the protein-conducting channel by a bound ribosome. Structure 17, 1453–1464 (2009).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Feller, S. E., Zhang, Y., Pastor, R. W. & Brooks, B. R. Constant pressure molecular dynamics simulation: the Langevin piston method. J. Chem. Phys. 103, 4613–4621 (1995).
Martyna, G. J., Tobias, D. J. & Klein, M. L. Constant pressure molecular dynamics algorithms. J. Chem. Phys. 101, 4177–4189 (1994).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: an N·log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Ryckaert, J.-P., Ciccotti, G. & Berendsen, H. J. C. Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J. Comput. Phys. 23, 327–341 (1977).
Gowers, R. et al. MDAnalysis: a python package for the rapid analysis of molecular dynamics simulations (eds Benthall, S. and Rostrup, S.) Proc. 15th Python Science Conf. 98–105 (2016).
Michaud-Agrawal, N., Denning, E. J., Woolf, T. B. & Beckstein, O. MDAnalysis: a toolkit for the analysis of molecular dynamics simulations. J. Comput. Chem. 32, 2319–2327 (2011).
Trabuco, L. G., Villa, E., Mitra, K., Frank, J. & Schulten, K. Flexible fitting of atomic structures into electron microscopy maps using molecular dynamics. Structure 16, 673–683 (2008).
Bayburt, T. H. & Sligar, S. G. Membrane protein assembly into Nanodiscs. FEBS Lett. 584, 1721–1727 (2010).
Cheng, Y.-C. & Prusoff, W. H. Relationship between the inhibition constant (K I) and the concentration of inhibitor which causes 50 per cent inhibition (I 50) of an enzymatic reaction. Biochem. Pharmacol. 22, 3099–3108 (1973).
Singh, S. K., Piscitelli, C. L., Yamashita, A. & Gouaux, E. A competitive inhibitor traps LeuT in an open-to-out conformation. Science 322, 1655–1661 (2008).
Acknowledgements
We thank the National Institute for Drug Abuse, Drug Supply Program for providing ibogaine and [3H]ibogaine, L. Vaskalis for assistance with figures, H. Owen for help with manuscript preparation, V. Navratna for discussions, and M. Whorton for help with Fab X-ray data collection. Electron microscopy was performed at Oregon Health & Science University (OHSU) at the Multiscale Microscopy Core with technical support from the OHSU-FEI Living Laboratory and OHSU Center for Spatial Systems Biomedicine. We acknowledge the staff of the Northeastern Collaborative Access Team at the Advanced Photon Source. Simulations have been performed using National Science Foundation computing resources allocated through an XSEDE grant (TGMCA06N060) to E.T., and PRAC allocation (grant ACI1713784 to E.T.) at Blue Waters of the National Center for Supercomputing Applications at the University of Illinois. We are grateful to Bernard and Jennifer LaCroute for their support. This work was funded by the National Institutes of Health (5R37MH070039 to E.G.; P41GM104601, U54GM087519 and R01GM123455 to E.T.). E.G. is an investigator of the Howard Hughes Medical Institute.
Reviewer information
Nature thanks Gary Rudnick and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
D.Y. initiated studies on the ibogaine inward-open conformation. J.A.C. initiated cryo-EM studies on SERT–antibody complexes. J.A.C., D.Y. and E.G. designed the project. J.A.C. and D.Y. contributed to all aspects of protein purification, biochemical characterization, electron microscopy data collection and processing, and built atomic models. J.A.C. and C.Y. collected the electron microscopy data on ts2-inactive SERT–paroxetine and ts2-active SERT–ibogaine complexes. J.A.C., C.Y. and D.Y. collected the electron microscopy data on ΔN72/C13 ibogaine occluded, inward-open, and noribogaine inward-open datasets. J.A.C., D.Y. and E.G. wrote the manuscript. Z.Z., P.-C.W. and E.T. performed ibogaine docking and molecular dynamics simulations, and wrote sections related to computational methods. All authors contributed to editing and manuscript preparation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Non-competitive inhibition of transport by ibogaine, ibogaine binding to the outward-open conformation, detection of the inward-open conformation, and the paroxetine ts2-inactive reconstruction.
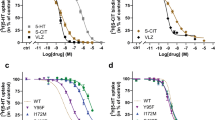
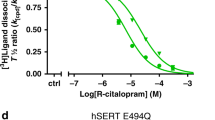
a, Ibogaine inhibition of 5-HT transport for wild-type (blue circles) and ts2-active (red squares) SERT variants using 20 μM [3H]5-HT. Data are mean ± s.e.m. (n = 3 biological replicates). The experiment was performed three times independently with the same results. b, Michaelis–Menten plots of 5-HT uptake for wild-type (blue) transporter in the absence (solid line, circles), or in the presence (dashed line, circles) of 5 μM ibogaine, and for ts2 (red) in the absence (solid line, squares), or in the presence (dashed line, squares) of 5 μM ibogaine. Data are mean ± s.e.m. (n = 3 biological replicates.). The experiment was performed three times independently with the same results. The mean Km and error (s.e.m.) of curve fitting for ΔN72/C13 is 2.2 ± 0.3 µM; and for ts2-active is 4 ± 1 µM. c, Competition binding of ibogaine with [3H]paroxetine for ts2 in the absence (filled squares) or presence (open squares) of 1 μM 15B8 and 8B6 yields a Ki value of 3.2 ± 0.4 µM. Data are mean ± s.e.m. of curve fitting (n = 3 technical replicates). The experiment was performed three times independently with the same results. d, [3H]ibogaine saturation binding experiments of ts2-inactive and ts2-active 15B8 Fab–8B6 scFv complex in 100 mM NaCl, and corresponding mean Kd values derived from the curve fit: ts2-inactive (filled squares, >5 µM), ts2-active 15B8 Fab–8B6 scFv complex (open triangles, >8 µM). Data are mean ± s.e.m. (n = 6 biological replicates). The experiment was performed twice with similar results. e, SDS–PAGE of S277C labelling with MTS-ACMA compared with the C7x construct in nanodiscs in the presence of 1 mM ibogaine and 100 mM NaCl. There is no detectable labelling of the C7x construct. The experiment was performed three times independently with similar results. f, Time-dependent labelling of S277C (background construct: ts2-active, C7x) with MTS-ACMA in the presence of ibogaine (filled circles) and paroxetine (open squares) in 100 mM NaCl. Data are mean ± s.e.m. (n = 3 technical replicates). The experiment was performed three times with similar results. g, Analysis of S277C labelling experiments using MTS-ACMA in the presence of ibogaine or paroxetine, analysed by SDS–PAGE and visualized by in-gel fluorescence. The experiment was performed three times independently with similar results. h, Three-dimensional reconstruction and fit to the density map with the model derived from the paroxetine-bound X-ray structure (PDB code: 6AWN)18. SERT is cyan, 15B8 is purple and 8B6 is green; TM1 and TM6 are orange and red, respectively. i, The fit of paroxetine into the electron-microscopy density map (blue mesh) and interacting residues. j, Left, details of the 15B8–SERT interface with the EL2 region shown as an electrostatic surface potential map and 15B8 shown in ribbon representation. The Fab is coloured dark blue (heavy chain) or light blue (light chain); selected Fab residues within 5 Å of SERT are shown as sticks. Right, a similar view but with the Fab shown as a semi-transparent electrostatic surface potential. EL2 of SERT is shown in ribbon representation and is coloured cyan.
Extended Data Fig. 2 Cryo-EM reconstruction of ts2-active SERT–15B8 Fab–8B6 scFv–paroxetine complex.
a, Workflow of cryo-EM data processing of the ts2-inactive SERT–15B8 Fab–8B6 scFv complex with paroxetine in the outward-open conformation. After particle picking, particles were sorted using 2D classification. 3D ab initio reconstructions were performed after 2D classification using cryoSPARC. One out of two predominant classes (boxed) exhibited a subset of homogeneous particles that were used for further processing and global alignment in cryoSPARC. The other class, upon refinement, yielded only a nanometre-resolution map. Local refinement using cisTEM improved the resolution of class 1 (boxed) upon masking of the Fab constant domain and micelle (mask is shown overlaid in blue on top of the reconstruction). The final reconstructed volume was sharpened using PHENIX. b, Representative cryo-EM micrograph. Individual single particles are circled in white. Scale bar, 50 nm. c, 2D class averages after three rounds of classification. d, The angular distribution of particles used in the final reconstruction. e, Cryo-EM density map coloured by local resolution estimation. f, FSC curves for cross-validation, the final map (blue), masked SERT–Fab complex (red), and a mask that isolated SERT (black). The low-resolution limit cut-off for refinement was 7.5 Å. g, FSC curves for model versus half map 1 (working, red), half map 2 (free, black) and model versus final map (blue). h, Cryo-EM density segments of TM1 to TM12. i, A spherical mask placed over SERT was used for focused 3D classification with 3 classes. Comparison of the classes did not reveal any substantial differences. The antibodies were removed for clarity. The number of particles belonging to each class average is: class 1, purple (11.9%, 25,530 particles); class 2, yellow (54.9%, 117,781 particles); class 3, cyan (33.2%, 71,226 particles).
Extended Data Fig. 3 Cryo-EM reconstruction of ts2-active 15B8 Fab–8B6 scFv–ibogaine complex.
a, Workflow of cryo-EM data processing of the ts2-active 15B8 Fab–8B6 scFv complex with ibogaine in the outward-open conformation. After particle picking, particles were sorted using 2D classification. Ab initio reconstructions were performed in cryoSPARC after 2D classification to obtain an initial reconstruction. Particles were used for further processing and global alignment in cryoSPARC followed by recentring in RELION and calculation of the local CTF using Gctf. Local refinement using cisTEM improved the resolution upon masking of the Fab constant domain and micelle (mask is shown overlaid in blue on top of the reconstruction). The final reconstructed volume was sharpened using cisTEM. b, Representative cryo-EM micrograph. Individual single particles are circled in white. Scale bar, 50 nm. c, 2D class averages after three rounds of classification. d, The angular distribution of particles used in the final reconstruction. e, Cryo-EM density map coloured by local resolution estimation. f, FSC curves for cross-validation, the final map (blue), masked SERT–Fab complex (red), and a mask that isolated SERT (black). The low-resolution limit cut-off for refinement was 7.5 Å. g, FSC curves for model versus half map 1 (working, red), half map 2 (free, black) and model versus final map (blue). h, Cryo-EM density segments of TM1 to TM12. i, A spherical mask placed over SERT was used for focused 3D classification with 3 classes. Comparison of the classes did not reveal any substantial differences. The antibodies were removed for clarity. The number of particles belonging to each class average is: class 1, purple (33.6%, 51,739 particles); class 2, yellow (38.8%, 59,747 particles); class 3, cyan (27.6%, 42,500 particles).
Extended Data Fig. 4 Cryo-EM reconstruction of ΔN72/C13 SERT–15B8 Fab–ibogaine complex in NaCl.
a, Workflow of cryo-EM data processing of the ΔN72/C13 SERT–15B8 Fab complex with ibogaine in NaCl in the occluded conformation. After particle picking, particles were sorted using 2D classification. Ab initio reconstructions were performed in cryoSPARC after 2D classification to obtain an initial reconstruction. Particles were used for further processing and global alignment in cryoSPARC followed by recentring in RELION and calculation of the local CTF using Gctf. Local refinement using cisTEM improved the resolution upon masking of the Fab constant domain and micelle (mask is shown overlaid in blue on top of the reconstruction). The final reconstructed volume was sharpened using cisTEM. b, Representative cryo-EM micrograph. Individual single particles are circled in white. Scale bar, 50 nm. c, 2D class averages after three rounds of classification. d, The angular distribution of particles used in the final reconstruction. e, Cryo-EM density map coloured by local resolution estimation. f, FSC curves for cross-validation, the final map (blue), masked SERT–15B8 Fab complex (red), and a mask that isolated SERT (black). The low-resolution limit cut-off for refinement was 7.0 Å. g, FSC curves for model versus half map 1 (working, red), half map 2 (free, black) and model versus final map (blue). h, Cryo-EM density segments of TM1 to TM12. i, A spherical mask placed over SERT was used for focused 3D classification with 3 classes. Comparison of the classes did not reveal any substantial differences. The Fab was removed for clarity. The number of particles belonging to each class average is: class 1, purple (78.9%, 571,547 particles); class 2, yellow (6.9%, 49,983 particles); class 3, cyan (14.2%, 102,863 particles).
Extended Data Fig. 5 Cryo-EM reconstruction of ΔN72/C13 SERT–15B8 Fab–ibogaine complex in KCl.
a, Workflow of cryo-EM data processing of the ΔN72/C13 SERT–15B8 Fab complex with ibogaine in KCl in the inward-open conformation. After particle picking, particles were sorted using 2D classification. Ab initio reconstructions were performed in cryoSPARC after 2D classification to obtain an initial reconstruction. Particles were further sorted in RELION using 3D classification and refined further in cryoSPARC. Local refinement using cisTEM improved the resolution upon masking of the Fab constant domain and micelle (mask is shown overlaid in blue on top of the reconstruction). The final reconstructed volume was sharpened using cisTEM. b, Representative cryo-EM micrograph. Individual single particles are circled in white. Scale bar, 50 nm. c, 2D class averages after three rounds of classification. d, The angular distribution of particles used in the final reconstruction. e, Cryo-EM density map coloured by local resolution estimation. f, FSC curves for cross-validation, the final map (blue), masked SERT–Fab complex (red), and a mask that isolated SERT (black). The low-resolution limit cut-off for refinement was 7.5 Å. g, FSC curves for model versus half map 1 (working, red), half map 2 (free, black) and model versus final map (blue). h, Cryo-EM density segments of TM1 to TM12. i, A spherical mask placed over SERT was used for focused 3D classification with 3 classes. Comparison of the classes did not reveal any substantial differences. The Fab was removed for clarity. The number of particles belonging to each class average is: class 1, purple (32.9%, 121,288 particles); class 2, yellow (33.7%, 124,237 particles); class 3, cyan (33.4%, 123,131 particles).
Extended Data Fig. 6 Cholesteryl hemisuccinate, map features at Thr276 and Ser277, and SERT–noribogaine complex.
a, Interaction between CHS, TM1a and TM5 in the occluded conformation of the ΔN72/C13 SERT–15B8–ibogaine complex in 100 mM NaCl. b, Non-proteinaceous density features (red) near Thr276 and Ser277. c, Noribogaine inhibition of 5-HT transport for ΔN72/C13 SERT. 5-HT transport was measured using 20 μM [3H]5-HT in the presence of the indicated concentrations of noribogaine. The mean IC50 of noribogaine inhibition of serotonin transport was determined from the curve with the error of the fit (s.e.m.): 1.2 ± 0.2 µM. Data are mean ± s.e.m. (n = 3 biological replicates). The experiment was performed twice independently with similar results. d, Michaelis–Menten plots of 5-HT uptake for the ΔN72/C13 transporter in the absence (solid line, circles), or in the presence (dashed line, squares) of 1 µM noribogaine; the mean Km was determined from the curve with the error of the fit (s.e.m.): ΔN72/C13: 2.7 ± 0.6 µM; in the presence of noribogiane: 2.7 ± 0.9 µM. Data are mean ± s.e.m. (n = 3 biological replicates). e, Noribogaine (solid line, circles) and ibogaine (dashed line, squares) competition binding with [3H]paroxetine for ΔN72/C13 SERT. Data are mean ± s.e.m. (n = 3 technical replicates). f, Density map of the ΔN72/C13 SERT–15B8–noribogaine complex, in 100 mM KCl, fit with the model derived from the inward-open ibogaine-bound SERT complex. SERT is cyan and the 15B8 Fab is purple; TM1 and TM6 of SERT are shown in orange and red, respectively. g, Noribogaine density in the central binding pocket. The fit of noribogaine into the electron microscopy density map was derived from ibogaine-bound SERT in the inward-open conformation and is shown in blue mesh, and residues involved in binding (Tyr176, Asp98, Phe341, Phe335, Asn177, Ile172 and Tyr95) are drawn as sticks. h, FSC curve for the noribogaine-bound SERT complex. The low-resolution limit cut-off for refinement was 9.0 Å.
Extended Data Fig. 7 Ibogaine docking and molecular dynamics simulations.
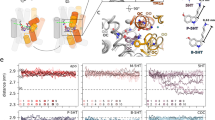
a, Workflow of ligand docking experiments. b, Optimal binding poses of ibogaine in the central binding site of the outward-open, occluded and inward-open conformations. For clarity, only the TM helices surrounding the central binding site (TM1, TM3, TM6 and TM8) are shown. The interaction between ibogaine and Asp98 of SERT, both shown in sticks, is highlighted. c, The simulation system used to study the structural stability and ibogaine binding of different conformations of SERT (two independent 50-ns simulations for each conformation), showing the transporter in cartoon form, with POPC lipids drawn in sticks, bulk water in a transparent surface, and solute ions (100 mM NaCl for the outward-open simulation) in yellow (Na+) and green (Cl−) spheres. d, Structural stability of bound ibogaine measured as the mass-weighted r.m.s.d. (including hydrogen atoms) of the ligand, as well as the Asp98–ibogaine (O–N) distance. The trajectories of outward-open, occluded and inward-open SERT are plotted in red, green and blue, respectively, and are shown for two independent simulations.
Extended Data Fig. 8 Measurement of ibogaine and noribogaine inhibition of mutants, effect of thermostabilizing Y110A mutation, movements of structural elements associated with alternating access mechanism, and alignment of TM5.
a, Inhibition of serotonin uptake by ibogaine or noribogaine for ts2. The mean IC50 of ibogaine and noribogaine inhibition of serotonin transport was determined from the curve with the error of the fit (s.e.m.) (black circles, ibogaine IC50: 7 ± 2 µM; noribogaine IC50: 1.1 ± 0.7 µM), N177L (blue circles, 1 ± 1 µM; 40 ± 10 µM), N177V (green triangles, 0.17 ± 0.04 µM; 24 ± 5 µM), N177A (red squares, 0.6 ± 0.3 µM; 300 ± 200 µM), N177T (cyan diamonds, 1.0 ± 0.2 µM; 8 ± 2 µM), and N177Q (magenta inverted triangles, 1.1 ± 0.7 µM; 1.0 ± 0.5 µM). Data are mean ± s.e.m. (n = 6 and n = 3 biological replicates for ibogaine and noribogaine, respectively). The experiment was performed three times independently with the same results. b, Comparison of EL4 and TM1b in the X-ray structure of the ts3–paroxetine complex (PDB code: 5I6X, purple)28 and the cryo-EM structure of the ts2 active–ibogaine complex in outward-open conformation (grey). Residues Tyr110 (ts2-active) and Ala110 (ts3) are shown in sticks. c, Comparison of the TM helices of the outward-open (grey), occluded (orange), and inward-open (blue) conformations viewed from the extracellular side of the membrane. The positions of TM2, TM4, TM5 and TM12 for each conformation are shown (middle). Right, the helical displacement measured from marker positions in each TM to a position in TM3 (Tyr186); from the outward-open to the occluded conformation (filled circles) and from the occluded to the inward-open conformation (open circles). The TM marker positions are described further in the Methods section ‘Measurements’. Error bars represent the s.d. d, Comparison of the TM helices viewed from the intracellular side of the membrane. The positions of TM5, TM9, TM11 and TM12 for each conformation are shown (middle). Right, the helical displacement measured from marker positions in each TM to a position in TM3 (Gly160); from the outward-open to the occluded conformation (filled circles) and from the occluded to the inward-open conformation (open circles). Error bars represent the s.d. e, Angular changes of TMs associated with transition from the outward-open to the occluded conformation (filled circles) and from the occluded to the inward-open conformation (open circles). Error bars represent the s.d. f, The intracellular region of TM5 ‘unwinds’ in the inward-open conformation. Gly278 and Pro288 in the GX9P motif are shown in sticks. g, Alignment of TM5 of SERT, dopamine transporter (DAT) and noradrenaline transporter (NET) with LeuT and MhsT. The position of the GX9P motif is indicated. h, Comparison of EL3, EL4 and EL6 in the outward-open (grey), the occluded (orange) and the inward-open (blue) conformations.
Extended Data Fig. 9 Comparison of SERT with bacterial transporters.
a, Superposition of the ibogaine-bound outward-open conformation (light grey) with the LeuT outward-open conformation (PDB code: 3F3A, dark grey)79. The graphs depict the r.m.s.d. and angular differences between the outward-open conformations of SERT and LeuT (3F3A, open triangles), LeuT outward-occluded (PDB code: 2A65, open squares)11, and LeuT inward-open (PDB code: 3TT3, filled circles)8. b, Superposition of the ibogaine-bound occluded conformation (orange) with the LeuT outward-occluded conformation (PDB code: 2A65, dark grey). The graphs compare the occluded conformation of SERT with LeuT conformations as described in a. c, Superposition of the ibogaine-bound inward-open conformation (blue) with the LeuT inward-open conformation (PDB code: 3TT3, dark grey). The graphs compare the occluded conformation of SERT to LeuT conformations as described in a. d, Comparison of the extracellular loops of LeuT in the outward-open (grey), occluded (orange) and inward-open (blue) conformations. e, Comparison of outward-open (light grey), occluded (orange) and inward-open (blue) conformations of SERT with the inward-occluded conformation of MhsT (PDB code: 4US3, dark grey)7. The graphs compare each conformation of SERT with MhsT as described in a.
Extended Data Fig. 10 Sodium and chloride ion-binding sites and putative substrate and ion-release pathways.
a, Comparison of the Na1 site (green boxes) and the Na2 site (purple boxes), and the Cl− site (yellow boxes) with the outward-open (S)-citalopram-bound and paroxetine-bound X-ray structures of SERT (PDB codes: 5I71 and 5I6X, grey)28. Left, outward-open ibogaine-bound conformation; middle, occluded conformation; right, inward-open conformation. The positions of sodium ions found in the X-ray structure are shown in grey. b, Solvent-accessible pathways in the inward-open conformation. Pathway 1 leads from the Na2 site to an opening formed between TM1a and TM5. Pathway 2 leads from the central binding site to an opening between TM1a, TM6b and TM5. c, The minimum radius of the ‘tunnels’ from the central binding site to the intracellular space was plotted as a function of the length of each pathway. The radius of the bicyclic amine moiety of ibogaine is approximately 2.5 Å.
Supplementary information
Video 1
Video depicts the overall map quality of the inward-open ΔN72/C13 SERT–15B8 Fab–ibogaine complex. TM1 and TM6 of SERT are coloured in orange and red with the rest of the transporter blue. The Fab is coloured purple.
Video 2
Animated workflow of ibogaine docking procedure for the outward-open SERT. TM1, TM3, TM6, and TM8 of SERT are shown in cartoon. Ibogaine is drawn using sticks with hydrogen atoms hidden for clarity.
Video 3
Ibogaine optimal poses from the systematic search protocol are shown for outward-open, occluded, and inward-open conformations. Morphing is used to show the transition from the outward-open to occluded, and then to the inward-open state with optimal binding poses of ibogaine. TM1, TM3, TM6, and TM8 of SERT are shown in cartoon, while ibogaine, Asp98, and Phe335 are drawn using sticks.
Video 4
Depiction of the conformational changes from outward-open, occluded, to the inward-open conformation. TM1 and TM6 are coloured in orange and red and TM5 in green. TM2, 7, 8, 10 and EL3, 4, and 6 are coloured in blue while the rest of the transporter is coloured in grey.
Rights and permissions
About this article
Cite this article
Coleman, J.A., Yang, D., Zhao, Z. et al. Serotonin transporter–ibogaine complexes illuminate mechanisms of inhibition and transport. Nature 569, 141–145 (2019). https://doi.org/10.1038/s41586-019-1135-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1135-1
This article is cited by
-
The role of neurotrophic factors in novel, rapid psychiatric treatments
Neuropsychopharmacology (2024)
-
Ligand coupling mechanism of the human serotonin transporter differentiates substrates from inhibitors
Nature Communications (2024)
-
A transcriptomic analysis in mice following a single dose of ibogaine identifies new potential therapeutic targets
Translational Psychiatry (2024)
-
Effects of congeners of amphetamine on the human heart
Naunyn-Schmiedeberg's Archives of Pharmacology (2024)
-
Membrane transporters in drug development and as determinants of precision medicine
Nature Reviews Drug Discovery (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.