Abstract
Chromatin remodelling complexes evict, slide, insert or replace nucleosomes, which represent an intrinsic barrier for access to DNA. These remodellers function in most aspects of genome utilization including transcription-factor binding, DNA replication and repair1,2. Although they are frequently mutated in cancer3, it remains largely unclear how the four mammalian remodeller families (SWI/SNF, ISWI, CHD and INO80) orchestrate the global organization of nucleosomes. Here we generated viable embryonic stem cells that lack SNF2H, the ATPase of ISWI complexes, enabling study of SNF2H cellular function, and contrast it to BRG1, the ATPase of SWI/SNF. Loss of SNF2H decreases nucleosomal phasing and increases linker lengths, providing in vivo evidence for an ISWI function in ruling nucleosomal spacing in mammals. Systematic analysis of transcription-factor binding reveals that these remodelling activities have specific effects on binding of different transcription factors. One group critically depends on BRG1 and contains the transcriptional repressor REST, whereas a non-overlapping set of transcription factors, including the insulator protein CTCF, relies on SNF2H. This selectivity readily explains why chromosomal folding and insulation of topologically associated domains requires SNF2H, but not BRG1. Collectively, this study shows that mammalian ISWI is critical for nucleosomal periodicity and nuclear organization and that transcription factors rely on specific remodelling pathways for correct genomic binding.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Next-generation sequencing data reported in this study have been deposited at the Gene Expression Omnibus with accession number GSE112136.
References
Becker, P. B. & Hörz, W. ATP-dependent nucleosome remodeling. Annu. Rev. Biochem. 71, 247–273 (2002).
Clapier, C. R. & Cairns, B. R. The biology of chromatin remodeling complexes. Annu. Rev. Biochem. 78, 273–304 (2009).
Kadoch, C. & Crabtree, G. R. Mammalian SWI/SNF chromatin remodeling complexes and cancer: Mechanistic insights gained from human genomics. Sci. Adv. 1, e1500447 (2015).
Ho, L. et al. esBAF facilitates pluripotency by conditioning the genome for LIF/STAT3 signalling and by regulating polycomb function. Nat. Cell Biol. 13, 903–913 (2011).
Ho, L. & Crabtree, G. R. Chromatin remodelling during development. Nature 463, 474–484 (2010).
Miller, E. L. et al. TOP2 synergizes with BAF chromatin remodeling for both resolution and formation of facultative heterochromatin. Nat. Struct. Mol. Biol. 24, 344–352 (2017).
King, H. W. & Klose, R. J. The pioneer factor OCT4 requires the chromatin remodeller BRG1 to support gene regulatory element function in mouse embryonic stem cells. eLife 6, e22631 (2017).
Fry, C. J. & Peterson, C. L. Chromatin remodeling enzymes: who’s on first? Curr. Biol. 11, R185–R197 (2001).
Vignali, M., Hassan, A. H., Neely, K. E. & Workman, J. L. ATP-dependent chromatin-remodeling complexes. Mol. Cell. Biol. 20, 1899–1910 (2000).
Corona, D. F. V. & Tamkun, J. W. Multiple roles for ISWI in transcription, chromosome organization and DNA replication. Biochim. Biophys. Acta 1677, 113–119 (2004).
Deuring, R. et al. The ISWI chromatin-remodeling protein is required for gene expression and the maintenance of higher order chromatin structure in vivo. Mol. Cell 5, 355–365 (2000).
Yen, K., Vinayachandran, V., Batta, K., Koerber, R. T. & Pugh, B. F. Genome-wide nucleosome specificity and directionality of chromatin remodelers. Cell 149, 1461–1473 (2012).
Längst, G. & Becker, P. B. Nucleosome mobilization and positioning by ISWI-containing chromatin-remodeling factors. J. Cell Sci. 114, 2561–2568 (2001).
Yamada, K. et al. Structure and mechanism of the chromatin remodelling factor ISW1a. Nature 472, 448–453 (2011).
Ocampo, J., Chereji, R. V., Eriksson, P. R. & Clark, D. J. The ISW1 and CHD1 ATP-dependent chromatin remodelers compete to set nucleosome spacing in vivo. Nucleic Acids Res. 44, 4625–4635 (2016).
Wiechens, N. et al. The chromatin remodelling enzymes SNF2H and SNF2L position nucleosomes adjacent to CTCF and other transcription factors. PLoS Genet. 12, e1005940 (2016).
Stopka, T. & Skoultchi, A. I. The ISWI ATPase Snf2h is required for early mouse development. Proc. Natl Acad. Sci. USA 100, 14097–14102 (2003).
Hakimi, M.-A. et al. A chromatin remodelling complex that loads cohesin onto human chromosomes. Nature 418, 994–998 (2002).
Oppikofer, M. et al. Expansion of the ISWI chromatin remodeler family with new active complexes. EMBO Rep. 18, 1697–1706 (2017).
Gkikopoulos, T. et al. A role for Snf2-related nucleosome-spacing enzymes in genome-wide nucleosome organization. Science 333, 1758–1760 (2011).
Kelly, T. K. et al. Genome-wide mapping of nucleosome positioning and DNA methylation within individual DNA molecules. Genome Res. 22, 2497–2506 (2012).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Ho, L. et al. An embryonic stem cell chromatin remodeling complex, esBAF, is essential for embryonic stem cell self-renewal and pluripotency. Proc. Natl Acad. Sci. USA 106, 5181–5186 (2009).
Valouev, A. et al. Determinants of nucleosome organization in primary human cells. Nature 474, 516–520 (2011).
Teif, V. B. et al. Genome-wide nucleosome positioning during embryonic stem cell development. Nat. Struct. Mol. Biol. 19, 1185–1192 (2012).
Mathelier, A. et al. JASPAR 2016: a major expansion and update of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 44, D110–D115 (2016).
Stadler, M. B. et al. DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature 480, 490–495 (2011).
Lambert, S. A. et al. The human transcription factors. Cell 172, 650–665 (2018).
Dekker, J. & Mirny, L. The 3D genome as moderator of chromosomal communication. Cell 164, 1110–1121 (2016).
Merkenschlager, M. & Nora, E. P. CTCF and cohesin in genome folding and transcriptional gene regulation. Annu. Rev. Genomics Hum. Genet. 17, 17–43 (2016).
Nora, E. P. et al. Targeted degradation of CTCF decouples local insulation of chromosome domains from genomic compartmentalization. Cell 169, 930–944 (2017).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Crane, E. et al. Condensin-driven remodelling of X chromosome topology during dosage compensation. Nature 523, 240–244 (2015).
Mumbach, M. R. et al. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat. Methods 13, 919–922 (2016).
Fyodorov, D. V., Blower, M. D., Karpen, G. H. & Kadonaga, J. T. Acf1 confers unique activities to ACF/CHRAC and promotes the formation rather than disruption of chromatin in vivo. Genes Dev. 18, 170–183 (2004).
Acknowledgements
We thank L. Giorgetti and L. Burger for suggestions and feedback, C. Wirbelauer for technical assistance, S. Smallwood for next-generation sequencing support and members of the Schübeler group for feedback on the project and manuscript. Brg1fl/fl embryonic stem cells were provided by G. Crabtree. Research in the laboratory of D.S. is supported by the Novartis Research Foundation, the European Research Council (ERC) under the European Union’s Horizon research and innovation program (grant agreement no. 667951) and the Swiss National Sciences Foundation. D.B. was funded by a Boehringer Ingelheim Fonds PhD fellowship. M.I. is funded by an EMBO long-term postdoctoral fellowship.
Reviewer information
Nature thanks David Clark, Jonathan Yuin-Han Loh and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
D.B., M.I., M.B.S. and D.S. conceived and planned the experiments. D.B. and M.I. executed the experiments and contributed to initial data analysis. M.B.S. performed comprehensive computational data analysis. D.S. supervised the project. All authors contributed to interpretation of the results and writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Phenotype description of SNF2H KO mouse ES cells.
a, Generation of SNF2H KO mouse embryonic cell lines. b, Images of wild-type and SNF2H KO cell cultures grown on gelatin-coated plates. c, Cell-cycle-phase composition, measured by fluorescence-activated cell sorting (DNA content and BrdU incorporation). d, Cell-cycle duration estimated by exponential loss of fluorescent dye. Doubling time was estimated from the slope of a linear fit to the data and is indicated (95% confidence intervals in parentheses). e, Images of wild-type and SNF2H KO cells during embryoid body formation. f, Differentially expressed genes between SNF2H KO and wild-type cells. Pluripotency marker genes are unchanged (labelled as 1 to 5). g, Gene ontology (GO) terms enriched in the set of differentially expressed genes between SNF2H KO and wild-type ES cells. The bars are the observed/expected ratios for the 30-most-significant terms from the ‘biological process’ section, with the number of differentially expressed genes associated to a term shown on the right. All reported GO terms have P values of 5.84 × 10−6 or smaller. All genes with an FDR < 0.05 and an absolute fold change >2.0 were used (n = 1,996), and terms with less than 5 or more than 1,000 associated genes were filtered out. h, Correlation-based clustering of RNA-seq profiles, illustrating that the transcriptome of wild-type add-back cells closely resembles the one of wild-type cells, whereas mutant add-back cells are still grouped with SNF2H KO cells.
Extended Data Fig. 2 Local quantification of nucleosome phasing using MNase-seq data.
a, Locus on chromosome 2 showing MNase-seq read densities, amplitudes for different frequencies (periods of 764, 382, 255, 191, 153, 127 and 109 bp) and reconstructed signal using only the nucleosome frequency (191-bp period). b, Bioanalyzer traces of MNase digests used for MNase-seq experiments.
Extended Data Fig. 3 MNase read densities around TSS and DHS.
a, Average MNase read densities around TSS. b, Average MNase read densities around TSS. Genes with high, medium or low RNA levels or for which RNA is not detected in SNF2H KO nor wild-type cells are indicated. c, MNase read densities around TSS as in b. Genes, the RNA levels of which increase, remain unchanged or decrease in SNF2H KO compared to wild-type cells or for which RNA is not detected are indicated. d, MNase-seq alignment densities in a 2-kb window around strong distal DHS. The 5,000 regions with the highest 95th percentile values in wild type were selected from 55,201 regions in total, and aligned at the centre of the DHS (indicated by the arrows at the bottom).
Extended Data Fig. 4 Characterization of genome-wide DNA methylation in SNF2H KO and wild-type cells.
a, Distributions of CpG-read coverage in whole-genome bisulfite sequencing DNA-methylation analysis. b, Distribution of CpA methylation (chromosome 10). c, Two-dimensional densities comparing CpG methylation in wild-type and SNF2H KO cells across different domains of the genome. d, RNA levels and changes relative to wild type of genes involved in DNA methylation.
Extended Data Fig. 5 NRL estimation in BRG1 conditional-knockout cells.
a, Scheme illustrating transposase insertion in chromatin. b, Mono-, di- and tri-nucleosomal fragment lengths estimated from smoothed and de-trended ATAC-seq fragment sizes (here showing wild-type replicate 1 as an example). The three estimates are averaged to obtain a single value per biological replicate. c, Estimated changes in nucleosome-repeat lengths based on ATAC-seq data from previous studies6,7 (as indicated) in BRG1 KO compared to controls. None of the datasets shows a significant increase in values (all P > 0.05, one-sided Welch two-sample t-test). d, e, MNase-based estimation of nucleosome repeat length upon BRG1 deletion using data from a previous study6. e, Counts of distances between same-strand MNase-seq alignments (phase) in Brg1fl/fl and BRG1 KO cells. e, Linear fits to the phase peaks from d and resulting nucleosome-repeat lengths with 95% confidence intervals. f, Genotyping PCR for floxed (2,507-bp product) and deleted (313-bp) BRG1 cells.
Extended Data Fig. 6 NRL estimation in SNF2H KO and wild-type cells using MNase data.
a, Counts of opposite-strand MNase-seq alignment distances illustrating the average fragment-length characteristic of a mono-nucleosome particle. b, Counts of same-strand MNase-seq alignment distances (phases). c, For each replicate MNase-seq dataset, counts of same-strand alignment distances (phases, left) with low- and high-span locally estimated scatterplot smoothing (LOESS) fits (red and blue, respectively), peak detection in residual phase counts (difference between low- and high-span fits, middle), and linear fits to the peak positions with nucleosome phase and 95% CI (right).
Extended Data Fig. 7 Chromatin accessibility in SNF2H KO, BRG1 KO and control cells.
a, Scatter plots of absolute ATAC signal in promoter and non-promoter peaks (shown in black and red, respectively) of wild-type and SNF2H KO (left), wild-type and mutant add-back (middle), and corresponding changes of ATAC signal (right). R, Pearson’s correlation coefficient. b, Pairwise correlation heat maps for absolute ATAC signal in all (left), promoter (middle) and non-promoter (right) peaks between wild-type, SNF2H KO and add-back samples (two replicates each). c, d, Same as in a and b for wild-type (Brg1fl/fl) and BRG1 KO samples (three replicates each).
Extended Data Fig. 8 Characterization of Snf2h- and Brg1-dependent transcription factors.
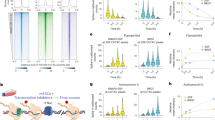
a, b, Enrichments of predicted transcription-factor sites in ATAC-seq peak bins compared to all other bins were calculated using HOMER and shown as Pearson residuals (a, (\({\rm{(Observed}}-{\rm{Expected)}}/\sqrt{{\rm{Expected}}}\)) or significance (b, −log(FDR)) for known motifs (rows) and bins of ATAC-seq peaks (columns). Peaks were binned into unchanged accessibility (change of less than 1.5-fold, grey), and bins of 1,500 peaks, each with increasing or decreasing fold changes (orange and purple). The colour bars on the left indicate the dependency of the transcription factor on chromatin remodellers (SNF2H and BRG1, SNF2H, BRG1 or unknown). c, Significance (−log(FDR)) of motif enrichment in the bin of peaks with the strongest decrease of accessibility, compared between SNF2H KO and BRG1 KO. Dashed lines indicate the significance thresholds used to classify transcription factors that depend on a given chromatin remodeller. d, RNA log2(fold changes) for transcription factor genes with different dependency on chromatin remodellers. The ‘independent’ class refers to transcription factors with motifs that are enriched in the bin with ATAC-seq peaks of unchanged accessibility. Two-sided Wilcoxon rank-sum test with continuity correction. e, f, ATAC profiles around binding sites for transcription factors CTCF (e) and REST (f) with differing dependency on SNF2H or BRG1. g, Western blot showing CTCF protein levels in wild-type and SNF2H KO cells (lamin B is shown as a loading control). For gel source data, see Supplementary Fig. 1. h, i, Average CpG methylation around bound CTCF motifs (h) and REST motifs (i) in wild-type and SNF2H KO cells. j, Profiles of MNase and de-trended methylation data downstream of REST motifs, with indicated nucleosome positions and offsets between wild-type and SNF2H KO. Profiles illustrate high MNase signal at phased nucleosomes, and high methylation values in the respective linker regions. The shift between wild-type and SNF2H KO maximal signals in the third peak is indicated by black lines.
Extended Data Fig. 9 Genome-wide binding of CTCF and REST in SNF2H KO, add-back clones, BRG1 KO and wild-type cells.
a, b, Unsupervised clustering of ChIP enrichments (log2(Immunoprecipitated/Input)) in motif-containing CTCF peaks (a, n = 49,678) and REST peaks (b, n = 2,703) for individual biological replicates from cells with different SNF2H and BRG1 genotypes. The colours between white and green indicate Pearson’s correlation coefficient between pairs of samples. c, CTCF ChIP enrichments (log2(Immunoprecipitated/Input)) for motif-containing CTCF peaks. The panels compare transcription-factor binding in wild-type and SNF2 KO (left), wild-type and mutant add-backs (middle) and Brg1fl/fl to BRG1 KO cells (right). R, Pearson’s correlation coefficient. d, Same as in c for REST. e, f, ChIP–seq alignment density in SNF2H add-back cells ±1 kb around random sets of 2,000 ChIP peaks for CTCF (e) and REST (f). The arrows at the bottom indicate the location of the predicted transcription-factor binding site.
Extended Data Fig. 10 Comparison of CTCF and REST binding sites with variable dependencies on specific chromatin remodellers.
a, Wild-type CTCF binding (x axis) and change of binding between SNF2H KO and wild-type (y axis) at predicted CTCF motifs. Strongly bound motifs can be subdivided into sites with strong and weak dependence on SNF2H, on the basis of their loss of CTCF binding upon SNF2H KO. b, Comparison of strongly and weakly dependent sites in terms of wild-type CTCF enrichment, accessibility in the neighbourhood measured by DNase I, frequency of sites that are part of cohesin chromatin loops34, CTCF motif score, distance to nearest TSS, accessibility in the close neighbourhood measured by ATAC and isolation score (fraction of DNase I or ATAC signal in central 200 bp versus 2 kb). c, d, Same as in a, b for REST. e, f, Two exemplary loci with CTCF (e) and REST (f) ChIP tracks in cells of different SNF2H and BRG1 genotypes (reads per kilobase of transcript per million mapped reads in 5-bp bins). Weakly and strongly dependent binding sites are indicated.
Extended Data Fig. 11 Measurements of chromatin conformation by Hi-C in individual replicates.
a, b, Normalized Hi-C contact maps at 20-kb resolution for individual replicates for a representative 4-Mb region on chromosome 16 (25–29 Mb), from cells with different SNF2H (a) or BRG1 (b) genotypes. Insulation scores for 200-kb squares are shown at the bottom as a heat map. c, Insulation scores (200-kb squares) for TAD boundaries with CTCF binding sites detected in individual replicates of SNF2H and BRG1-deleted cells with corresponding wild-type controls. d, e, log interaction frequency as a function of log distance for individual Hi-C replicates in Snf2h wild-type and KO cells (d) and Brg1fl/fl and BRG1 KO cells (e). The decay exponents have been estimated for short (50–200 kb) and long (0.5–2 Mb in c and 0.3–1 Mb in d) genomic distances, omitting the nonlinear part at the bottom.
Extended Data Fig. 12 CTCF-cohesin loops and A/B compartments in individual replicates.
a, Interaction frequencies at SMC1A HiChip loops, aggregated over loops separated by 280 to 380 kb (n = 701). b, Interaction Z scores at SMC1A HiChip loops, aggregated over loops separated by 60 to 500 kb (n = 4,375) and averaged over two biological replicates. c, Same as in b for individual replicates. d, Changes of interaction Z scores at SMC1A HiChip loops from c in individual replicate pairs. e, Changes of interaction Z scores at SMC1A HiChip loops, separated by CTCF motif orientation (+−, convergent; ++/−−/−+, other) and dependency on SNF2H (strong or weak). f, Normalized Hi-C contact maps at 150-kb resolution for entire chromosome 2 showing individual replicates. A/B compartments are indicated below the plots in green and red, respectively. g, cis-Eigenvector 1 values (multiplied by 100) across entire chromosome 2.
Supplementary information
Supplementary Information
This file contains Supplementary Methods with references and Supplementary Table 1, with NGS sample information.
Supplementary Figure
This file contains the uncropped western blots.
Rights and permissions
About this article
Cite this article
Barisic, D., Stadler, M.B., Iurlaro, M. et al. Mammalian ISWI and SWI/SNF selectively mediate binding of distinct transcription factors. Nature 569, 136–140 (2019). https://doi.org/10.1038/s41586-019-1115-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1115-5
This article is cited by
-
MYH1G-AS is a chromatin-associated lncRNA that regulates skeletal muscle development in chicken
Cellular & Molecular Biology Letters (2024)
-
The impact of DNA methylation on CTCF-mediated 3D genome organization
Nature Structural & Molecular Biology (2024)
-
Differential requirements for Smarca5 expression during hematopoietic stem cell commitment
Communications Biology (2024)
-
Context-specific functions of chromatin remodellers in development and disease
Nature Reviews Genetics (2024)
-
In vitro reconstitution of chromatin domains shows a role for nucleosome positioning in 3D genome organization
Nature Genetics (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.