Abstract
ATP-citrate lyase (ACLY) is a central metabolic enzyme and catalyses the ATP-dependent conversion of citrate and coenzyme A (CoA) to oxaloacetate and acetyl-CoA1,2,3,4,5. The acetyl-CoA product is crucial for the metabolism of fatty acids6,7, the biosynthesis of cholesterol8, and the acetylation and prenylation of proteins9,10. There has been considerable interest in ACLY as a target for anti-cancer drugs, because many cancer cells depend on its activity for proliferation2,5,11. ACLY is also a target against dyslipidaemia and hepatic steatosis, with a compound currently in phase 3 clinical trials4,5. Many inhibitors of ACLY have been reported, but most of them have weak activity5. Here we report the development of a series of low nanomolar, small-molecule inhibitors of human ACLY. We have also determined the structure of the full-length human ACLY homo-tetramer in complex with one of these inhibitors (NDI-091143) by cryo-electron microscopy, which reveals an unexpected mechanism of inhibition. The compound is located in an allosteric, mostly hydrophobic cavity next to the citrate-binding site, and requires extensive conformational changes in the enzyme that indirectly disrupt citrate binding. The observed binding mode is supported by and explains the structure–activity relationships of these compounds. This allosteric site greatly enhances the ‘druggability’ of ACLY and represents an attractive target for the development of new ACLY inhibitors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The atomic coordinates and the electron microscopy maps have been deposited at the Protein Data Bank (PDB) with accession code 6O0H.
References
Chypre, M., Zaidi, N. & Smans, K. ATP-citrate lyase: a mini-review. Biochem. Biophys. Res. Commun. 422, 1–4 (2012).
Zaidi, N., Swinnen, J. V. & Smans, K. ATP-citrate lyase: a key player in cancer metabolism. Cancer Res. 72, 3709–3714 (2012).
Fan, F. et al. On the catalytic mechanism of human ATP citrate lyase. Biochemistry 51, 5198–5211 (2012).
Burke, A. C. & Huff, M. W. ATP-citrate lyase: genetics, molecular biology and therapeutic target for dyslipidemia. Curr. Opin. Lipidol. 28, 193–200 (2017).
Granchi, C. ATP citrate lyase (ACLY) inhibitors: an anti-cancer strategy at the crossroads of glucose and lipid metabolism. Eur. J. Med. Chem. 157, 1276–1291 (2018).
Wakil, S. J., Stoops, J. K. & Joshi, V. C. Fatty acid synthesis and its regulation. Annu. Rev. Biochem. 52, 537–579 (1983).
Tong, L. Structure and function of biotin-dependent carboxylases. Cell. Mol. Life Sci. 70, 863–891 (2013).
Goldstein, J. L. & Brown, M. S. Regulation of the mevalonate pathway. Nature 343, 425–430 (1990).
Wellen, K. E. et al. ATP-citrate lyase links cellular metabolism to histone acetylation. Science 324, 1076–1080 (2009).
McTaggart, S. J. Isoprenylated proteins. Cell. Mol. Life Sci. 63, 255–267 (2006).
Hatzivassiliou, G. et al. ATP citrate lyase inhibition can suppress tumor cell growth. Cancer Cell 8, 311–321 (2005).
Warburg, O. On the origin of cancer cells. Science 123, 309–314 (1956).
Sun, T., Hayakawa, K., Bateman, K. S. & Fraser, M. E. Identification of the citrate-binding site of human ATP-citrate lyase using X-ray crystallography. J. Biol. Chem. 285, 27418–27428 (2010).
Sun, T., Hayakawa, K. & Fraser, M. E. ADP-Mg2+ bound to the ATP-grasp domain of ATP-citrate lyase. Acta Crystallogr. F 67, 1168–1172 (2011).
Hu, J., Komakula, A. & Fraser, M. E. Binding of hydroxycitrate to human ATP-citrate lyase. Acta Crystallogr. D 73, 660–671 (2017).
Li, J. J. et al. 2-hydroxy-N-arylbenzenesulfonamides as ATP-citrate lyase inhibitors. Bioorg. Med. Chem. Lett. 17, 3208–3211 (2007).
Dufort, F. J. et al. Glucose-dependent de novo lipogenesis in B lymphocytes: a requirement for ATP-citrate lyase in lipopolysaccharide-induced differentiation. J. Biol. Chem. 289, 7011–7024 (2014).
Karpusas, M., Branchaud, B. & Remington, S. J. Proposed mechanism for the condensation reaction of citrate synthase: 1.9-Å structure of the ternary complex with oxaloacetate and carboxymethyl coenzyme A. Biochemistry 29, 2213–2219 (1990).
Russell, R. J., Ferguson, J. M., Hough, D. W., Danson, M. J. & Taylor, G. L. The crystal structure of citrate synthase from the hyperthermophilic archaeon Pyrococcus furiosus at 1.9 Å resolution. Biochemistry 36, 9983–9994 (1997).
Francois, J. A. et al. Structure of a NADH-insensitive hexameric citrate synthase that resists acid inactivation. Biochemistry 45, 13487–13499 (2006).
Chittori, S., Savithri, H. S. & Murthy, M. R. N. Crystal structure of Salmonella typhimurium 2-methylcitrate synthase: insights on domain movement and substrate specificity. J. Struct. Biol. 174, 58–68 (2011).
Holm, L., Kääriäinen, S., Rosenström, P. & Schenkel, A. Searching protein structure databases with DaliLite v.3. Bioinformatics 24, 2780–2781 (2008).
Alter, G. M. et al. Mutation of essential catalytic residues in pig citrate synthase. Biochemistry 29, 7557–7563 (1990).
Evans, C. T., Kurz, L. C., Remington, S. J. & Srere, P. A. Active site mutants of pig citrate synthase: effects of mutations on the enzyme catalytic and structural properties. Biochemistry 35, 10661–10672 (1996).
Kurz, L. C. et al. Effects of changes in three catalytic residues on the relative stabilities of some of the intermediates and transition states in the citrate synthase reaction. Biochemistry 37, 9724–9737 (1998).
Huang, J., Malhi, M., Deneke, J. & Fraser, M. E. Structure of GTP-specific succinyl-CoA synthetase in complex with CoA. Acta Crystallogr. F 71, 1067–1071 (2015).
Fraser, M. E., James, M. N., Bridger, W. A. & Wolodko, W. T. A detailed structural description of Escherichia coli succinyl-CoA synthetase. J. Mol. Biol. 285, 1633–1653 (1999).
Wang, L. et al. Accurate and reliable prediction of relative ligand binding potency in prospective drug discovery by way of a modern free-energy calculation protocol and force field. J. Am. Chem. Soc. 137, 2695–2703 (2015).
Wang, L. et al. Modeling local structural rearrangements using FEP/REST: application to relative binding affinity predictions of CDK2 inhibitors. J. Chem. Theory Comput. 9, 1282–1293 (2013).
Ohi, M., Li, Y., Cheng, Y. & Walz, T. Negative staining and image classification — powerful tools in modern electron microscopy. Biol. Proced. Online 6, 23–34 (2004).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Yang, Z., Fang, J., Chittuluru, J., Asturias, F. J. & Penczek, P. A. Iterative stable alignment and clustering of 2D transmission electron microscope images. Structure 20, 237–247 (2012).
Hohn, M. et al. SPARX, a new environment for Cryo-EM image processing. J. Struct. Biol. 157, 47–55 (2007).
Kimanius, D., Forsberg, B. O., Scheres, S. H. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Bai, X. C., Rajendra, E., Yang, G., Shi, Y. & Scheres, S. H. Sampling the conformational space of the catalytic subunit of human γ-secretase. eLife 4, e11182 (2015).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Harder, E. et al. OPLS3: a force field providing broad coverage of drug-like small molecules and proteins. J. Chem. Theory Comput. 12, 281–296 (2016).
Wang, L., Berne, B. J. & Friesner, R. A. On achieving high accuracy and reliability in the calculation of relative protein-ligand binding affinities. Proc. Natl Acad. Sci. USA 109, 1937–1942 (2012).
Wang, L., Friesner, R. A. & Berne, B. J. Replica exchange with solute scaling: a more efficient version of replica exchange with solute tempering (REST2). J. Phys. Chem. B 115, 9431–9438 (2011).
Acknowledgements
We thank R. Grassucci, D. Bobe, K. Jordan, L. Yen, A. Raczkowski, W. Rice, E. Eng, M. Kopylov, and B. Carragher for access to the electron microscopy facility and help with data collection at the New York Structural Biology Center; and R. Grassucci and Z. Fu for help with data collection at the EM facility at Columbia University. We thank S. Qian and X. Qiang (Pharmaron) for enzyme-activity assay and kinetics studies, M. Geitman (Beactica) for SPR studies, D. Lakshminarasimhan (Xtal BioStructures) for structural studies on the N-terminal segment, proteolysis and early DSF work, Pharmaron for small-molecule synthesis, and H. Blanchette and K. Kreutter (Nimbus) for discussions. The portion of this research that was conducted in the L.T. laboratory was supported by a grant from Nimbus. Some of this work was performed at the Simons Electron Microscopy Center and National Resource for Automated Molecular Microscopy located at the New York Structural Biology Center, supported by grants from the Simons Foundation (349247), NYSTAR, and the NIH National Institute of General Medical Sciences (GM103310).
Reviewer information
Nature thanks Frank M. Raushel and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
J.W. carried out protein expression and purification, grid preparation and optimization, electron microscopy data collection, image processing and reconstruction, atomic model building and refinement, and thermal shift assays. S.L. designed the small-molecule inhibitors. J.K. and H.J.H. designed the enzyme assay and kinetics studies. J.K. designed the SPR studies. E.T. and S.R. performed the free-energy perturbation studies. B.D. identified the conditions for stabilizing the ACLY protein. S.L., J.K., H.J.H., B.D., and L.T. supervised the research and analysed the results. L.T., J.W., J.K., B.D., and E.T. wrote the paper. All authors commented on the paper.
Corresponding author
Ethics declarations
Competing interests
L.T. is a consultant for Nimbus.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Characterization of inhibitor binding to ACLY.
a, The four steps in the reaction catalysed by ACLY. b, Production of ADP is linear over 60 min. Data from duplicate experiments are shown, but the linearity of the progress curve was observed in many additional experiments. c–e, Double reciprocal plots showing that NDI-091143 is competitive versus citrate (Ki = 7.0 ± 0.8 nM) (c); noncompetitive versus ATP (Ki = 218 ± 14 nM) (d); and mixed-type noncompetitive versus CoA (Ki = 5.0 ± 2.1 nM, Ki′ = 156 ± 20 nM) (e). Data from duplicate experiments are shown (n = 2). Although the error on the Ki value versus CoA is relatively high, it does not affect the conclusion regarding the pattern of inhibition versus CoA. The assigned pattern of inhibition has 99.99% probability based on global fitting. The fixed substrate concentrations were 300 μM ATP, 300 μM CoA, and 5 mM citrate. f, Limited proteolysis experiments on full-length human ACLY wild-type (WT), H760A, and H760E mutants in the presence of various substrates and/or NDI-091143. CHY, chymotrypsin. The experiments were repeated at least three times, with similar results.
Extended Data Fig. 2 Negative-stain electron microscopy analysis of ACLY in the absence or presence of inhibitor NDI-091143.
a, A representative negative-stain image of ACLY in the presence of Mg-ATP only. A total of 155 images were examined, with similar results. b, Selected 2D class averages of negatively stained ACLY in the presence of Mg-ATP, generated with the ISAC algorithm. c, A representative negative-stain image of ACLY in the presence of Mg-ATP and NDI-091143. A total of 216 images were examined, with similar results. d, Selected 2D class averages of negatively stained ACLY in the presence of Mg-ATP and inhibitor, generated using the ISAC algorithm. e, The model generated with VIPER and refined with Meridien in SPARX using particles from d.
Extended Data Fig. 3 Processing of single-particle cryo-EM data for ACLY.
Flowchart of data processing showing a representative cryo-EM micrograph, selected reference-free 2D class averages, 3D classification and refinement procedures for the full-length ACLY tetramer as well as two N-terminal segments.
Extended Data Fig. 4 Cryo-EM analysis of the ACLY homo-tetramer and N-terminal segment.
a, Gold-standard FSC curves for masked, unmasked, and high-resolution phase-randomized reconstructions of the ACLY tetramer, as well as its final corrected FSC curve. The correlation is calculated between two independently determined half maps. b, Local-resolution map of the ACLY tetramer. c, Gold-standard FSC curves for the ACLY N-terminal segment, as well as its final corrected FSC curve. The correlation is calculated between two independently determined half maps. d, Local-resolution map of the ACLY N-terminal segment.
Extended Data Fig. 5 Electron microscopy density for selected regions of the ACLY structure.
a, Cryo-EM 3D map of the ACLY tetramer, coloured according to Fig. 1a. b, Electron microscopy density map for a helix in the C-terminal CCLY domain. c, Electron microscopy density of the linker between N-terminal and C-terminal segments (residues 818–830). d, Cryo-EM 3D map of the ACLY N-terminal segment, coloured according to Fig. 1a. e, Electron microscopy density for the β-sheet in the citrate domain. f, Electron microscopy density map for ADP and surrounding residues in the ATP-grasp domain.
Extended Data Fig. 6 Structural comparison between the CCLY domain of ACLY and CS.
a, Overlay of the CCLY tetramer (labelled 1, 2, 3, and 4) with a dimer of chicken CS (light blue and grey). b, Overlay of the CCLY tetramer with a dimer of CS, viewed after a 90° rotation around the horizontal axis from a. Regions where the N-terminal segment of CS overlaps with protomer 3 of CCLY are indicated by red arrows. The second protomer (B) of the CS dimer does not overlay well with CCLY. c, Overlay of the CCLY core (green) with a C-terminal segment (residues 167–366) of 2-methyl CS (grey). d, Overlay of the CCLY tetramer with a dimer of 2-methyl CS. Regions where the N-terminal segment of 2-methyl CS overlaps with protomer 3 of CCLY are indicated by red arrows. The second protomer (B) of the 2-methyl CS dimer has good overlay with protomer 2 of CCLY, suggesting a CCLY dimer (protomers 1 and 2) has similar organization to the 2-methyl CS dimer, although 2-methyl CS dimers can also assemble into decamers.
Extended Data Fig. 7 Catalytic mechanisms of CS and CCLY.
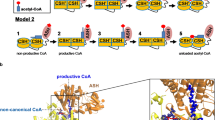
a, Catalytic mechanism of CS. The two steps of the reaction to produce citryl-CoA are labelled. An enol CoA intermediate is produced after the first step. The three arginine residues that recognize OAA are shown. The prime on the label of Arg421 indicates that it is from a different protomer of the enzyme. b, Possible catalytic mechanism of the CCLY domain of ACLY.
Extended Data Fig. 8 Substrate-binding sites in ACLY.
a, Possible binding mode of CoA in the CoA domain. Lys1018 from the CCLY domain of protomer 3 of the tetramer (orange) may contribute to CoA binding. b, Binding sites of NDI-091143 (black) and ADP (brown) as observed in this structure, as well as the possible binding sites of citrate (grey), CoA in the N-terminal CoA domain (CoA1, grey), CoA in the CCLY domain of protomer 3 of the tetramer (CoA2, light blue), and OAA as a mimic of the citryl group (light blue). The translocation of CoA between the two binding sites is indicated by the red arrow. Therefore, step 1 of the ACLY reaction occurs in the ATP-grasp active site, steps 2 and 3 occur at the interface between the citrate and CoA domains, and step 4 occurs in the CCLY domain. The pHis domain provides the His760 residue for both steps 1 and 2.
Extended Data Fig. 9 The inhibitor NDI-091143 is almost completely shielded from the solvent in the binding site.
a, An alternative model for the inhibitor does not fit the density as well and clashes with the protein. The bound conformation of the inhibitor has pseudo two-fold symmetry, around the sulfonamide group. In the alternative model (thin sticks in grey), the distal phenyl group (blue arrow) has weaker density, and an oxygen atom (red arrow) on the sulfonamide has steric clash with the protein (main-chain amide of Gly342). This alternative model is not consistent with the structure–activity relationship of the compounds. The view is related to Fig. 4a by a 40° rotation around the horizontal axis. b, The inhibitor is located in a cavity (grey transparent surface) in the hydrophobic core of the citrate domain. c, The cavity does not exist in the structure of the citrate complex (note the lack of any surfaces near the compound-binding site), and the inhibitor clashes with the enzyme, especially the side chain of Arg379 (red arrow). d, e, The inhibitor is only accessible to the solvent through two small portals. The protein is shown as a molecular surface (cyan).
Supplementary information
Supplementary Information
This file contains Supplementary Text regarding the synthesis of NDI-091143, Scheme 1, and Supplementary Information Figs. 1–3.
Rights and permissions
About this article
Cite this article
Wei, J., Leit, S., Kuai, J. et al. An allosteric mechanism for potent inhibition of human ATP-citrate lyase. Nature 568, 566–570 (2019). https://doi.org/10.1038/s41586-019-1094-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1094-6
This article is cited by
-
Biomimetic chiral hydrogen-bonded organic-inorganic frameworks
Nature Communications (2024)
-
Tumor biomarkers for diagnosis, prognosis and targeted therapy
Signal Transduction and Targeted Therapy (2024)
-
RBM25 is required to restrain inflammation via ACLY RNA splicing-dependent metabolism rewiring
Cellular & Molecular Immunology (2024)
-
Disease-associated astrocyte epigenetic memory promotes CNS pathology
Nature (2024)
-
Metabolic reprogramming in colorectal cancer: regulatory networks and therapy
Cell & Bioscience (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.