Abstract
The genetic compensation response (GCR) has recently been proposed as a possible explanation for the phenotypic discrepancies between gene-knockout and gene-knockdown1,2; however, the underlying molecular mechanism of the GCR remains uncharacterized. Here, using zebrafish knockdown and knockout models of the capn3a and nid1a genes, we show that mRNA bearing a premature termination codon (PTC) promptly triggers a GCR that involves Upf3a and components of the COMPASS complex. Unlike capn3a-knockdown embryos, which have small livers, and nid1a-knockdown embryos, which have short body lengths2, capn3a-null and nid1a-null mutants appear normal. These phenotypic differences have been attributed to the upregulation of other genes in the same families. By analysing six uniquely designed transgenes, we demonstrate that the GCR is dependent on both the presence of a PTC and the nucleotide sequence of the transgene mRNA, which is homologous to the compensatory endogenous genes. We show that upf3a (a member of the nonsense-mediated mRNA decay pathway) and components of the COMPASS complex including wdr5 function in GCR. Furthermore, we demonstrate that the GCR is accompanied by an enhancement of histone H3 Lys4 trimethylation (H3K4me3) at the transcription start site regions of the compensatory genes. These findings provide a potential mechanistic basis for the GCR, and may help lead to the development of therapeutic strategies that treat missense mutations associated with genetic disorders by either creating a PTC in the mutated gene or introducing a transgene containing a PTC to trigger a GCR.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The authors declare that all data supporting the findings of this study are available within the article, its Extended Data or from the corresponding author upon reasonable request. All zebrafish mutants, transgenic lines and plasmid constructs generated in this study are readily available from the authors.
References
Rossi, A. et al. Genetic compensation induced by deleterious mutations but not gene knockdowns. Nature 524, 230–233 (2015).
Zhu, P. et al. Short body length phenotype is compensated by the upregulation of nidogen family members in a deleterious nid1a mutation of zebrafish. J. Genet. Genomics 44, 553–556 (2017).
Barabási, A. L. & Oltvai, Z. N. Network biology: understanding the cell’s functional organization. Nat. Rev. Genet. 5, 101–113 (2004).
De Souza, A. T. et al. Transcriptional and phenotypic comparisons of Ppara knockout and siRNA knockdown mice. Nucleic Acids Res. 34, 4486–4494 (2006).
Gao, Y. et al. Auxin binding protein 1 (ABP1) is not required for either auxin signaling or Arabidopsis development. Proc. Natl Acad. Sci. USA 112, 2275–2280 (2015).
Stalder, L. & Mühlemann, O. Transcriptional silencing of nonsense codon-containing immunoglobulin micro genes requires translation of its mRNA. J. Biol. Chem. 282, 16079–16085 (2007).
Bühler, M., Mohn, F., Stalder, L. & Mühlemann, O. Transcriptional silencing of nonsense codon-containing immunoglobulin minigenes. Mol. Cell 18, 307–317 (2005).
Schuermann, A., Helker, C. S. & Herzog, W. Metallothionein 2 regulates endothelial cell migration through transcriptional regulation of vegfc expression. Angiogenesis 18, 463–475 (2015).
Chang, Y. F., Imam, J. S. & Wilkinson, M. F. The nonsense-mediated decay RNA surveillance pathway. Annu. Rev. Biochem. 76, 51–74 (2007).
Lykke-Andersen, J., Shu, M. D. & Steitz, J. A. Human Upf proteins target an mRNA for nonsense-mediated decay when bound downstream of a termination codon. Cell 103, 1121–1131 (2000).
Kunz, J. B., Neu-Yilik, G., Hentze, M. W., Kulozik, A. E. & Gehring, N. H. Functions of hUpf3a and hUpf3b in nonsense-mediated mRNA decay and translation. RNA 12, 1015–1022 (2006).
Kim, V. N., Kataoka, N. & Dreyfuss, G. Role of the nonsense-mediated decay factor hUpf3 in the splicing-dependent exon-exon junction complex. Science 293, 1832–1836 (2001).
Chan, W. K. et al. A UPF3-mediated regulatory switch that maintains RNA surveillance. Nat. Struct. Mol. Biol. 16, 747–753 (2009).
Shum, E. Y. et al. The antagonistic gene paralogs Upf3a and Upf3b govern nonsense-mediated RNA decay. Cell 165, 382–395 (2016).
Abudayyeh, O. O. et al. RNA targeting with CRISPR–Cas13. Nature 550, 280–284 (2017).
Wang, K. C. et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 472, 120–124 (2011).
Gomez, J. A. et al. The NeST long ncRNA controls microbial susceptibility and epigenetic activation of the interferon-γ locus. Cell 152, 743–754 (2013).
Vastenhouw, N. L. et al. Chromatin signature of embryonic pluripotency is established during genome activation. Nature 464, 922–926 (2010).
Haberle, V. et al. Two independent transcription initiation codes overlap on vertebrate core promoters. Nature 507, 381–385 (2014).
Shilatifard, A. The COMPASS family of histone H3K4 methylases: mechanisms of regulation in development and disease pathogenesis. Annu. Rev. Biochem. 81, 65–95 (2012).
Wu, M. et al. Molecular regulation of H3K4 trimethylation by Wdr82, a component of human Set1/COMPASS. Mol. Cell. Biol. 28, 7337–7344 (2008).
Wang, P. et al. Global analysis of H3K4 methylation defines MLL family member targets and points to a role for MLL1-mediated H3K4 methylation in the regulation of transcriptional initiation by RNA polymerase II. Mol. Cell. Biol. 29, 6074–6085 (2009).
Hu, D. et al. The Mll2 branch of the COMPASS family regulates bivalent promoters in mouse embryonic stem cells. Nat. Struct. Mol. Biol. 20, 1093–1097 (2013).
Hu, D. et al. Not all H3K4 methylations are created equal: Mll2/COMPASS dependency in primordial germ cell specification. Mol. Cell 65, 460–475.e466 (2017).
Tao, T. et al. Def defines a conserved nucleolar pathway that leads p53 to proteasome-independent degradation. Cell Res. 23, 620–634 (2013).
Wittkopp, N. et al. Nonsense-mediated mRNA decay effectors are essential for zebrafish embryonic development and survival. Mol. Cell. Biol. 29, 3517–3528 (2009).
Huang, H. T. et al. A network of epigenetic regulators guides developmental haematopoiesis in vivo. Nat. Cell Biol. 15, 1516–1525 (2013).
Chen, J. et al. Loss of function of def selectively up-regulates Δ113p53 expression to arrest expansion growth of digestive organs in zebrafish. Genes Dev. 19, 2900–2911 (2005).
Chang, C. et al. liver-enriched gene 1a and 1b encode novel secretory proteins essential for normal liver development in zebrafish. PLoS One 6, e22910 (2011).
Bogdanović, O., Fernández-Miñán, A., Tena, J. J., de la Calle-Mustienes, E. & Gómez-Skarmeta, J. L. The developmental epigenomics toolbox: ChIP-seq and MethylCap-seq profiling of early zebrafish embryos. Methods 62, 207–215 (2013).
Acknowledgements
We thank X. Feng and W. Shou for reading the manuscript; and C. Wood for language editing. This work was supported by the National Key R&D Program of China (2017YFA0504501, 2015CB942802), the National Natural Science Foundation of China (31571511 and 31871500) and the Fundamental Research Funds for the Central Universities.
Reviewer information
Nature thanks Ferenc Müller, Ulf Ørom and Miles Wilkinson for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
J.C. and Z.M. conceived and designed the research; Z.M. performed most of the experiments and analysed the data; P.Z. generated the nid1ad2/d2 and 5 capn3a (mu2-6) mutants, and provided assistance with the data analysis; H.S. generated the capn3aΔ14/Δ14 mutant and capn3a polyclonal antibody; L.G. and Z.Z. performed the experiments of uncapped and capped capn3a RNA injection and the wdr5 complementation test; Q.Z. and S.C. provided assistance in the protein co-immunoprecipitation assay; Y.C. performed capn8 and capn12 WISH and Cas13a gRNA knockdown experiments; J.C., J.P. and Z.M. wrote the manuscript; J.C. and J.P. supervised the research; J.C. provided oversight of the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Small liver phenotype is compensated by the upregulation of capn family members in zebrafish capn3aΔ14/Δ14 mutant embryos.
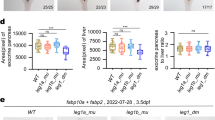
a, WISH of capn3a in wild-type embryos at 1, 1.5, 2, 2.5, 3 and 4 dpf. lv, liver. b, Western blot analysis of Capn3a protein (arrow) in wild-type, wild-type injected with capn3a-MO, and capn3aΔ14/Δ14 (designated capn3a−/−) embryos at 1.5 dpf. Actb1 was used as a loading control. c, Relative mRNA expression of capn3a in wild-type embryos at 1 dpf with different treatments as indicated. The dCas9 mRNA was co-injected with two non-template (NT) gRNAs or a template (T) gRNA (specifically targeting capn3a) into wild-type embryos. Ctr, uninjected embryos. d, Representative images showing WISH of lfabp, a liver-specific marker, to assess the liver sizes in wild-type embryos co-injected with dCas9 mRNA and either template or non-template gRNA at 3.5 dpf. e, Statistical analysis of the average liver sizes in d. Each dot represents the liver size of an individual embryo as described in Fig. 1d. f, Schematic diagram showing the genomic structure and a genetic mutation of zebrafish capn3a gene. ATG denotes the translation start codon; black lines denote introns; blue vertical bar denotes exon. The wild-type genomic DNA sequence of the TAL-spacer is shown. The red line denotes deletion of 14 bp in capn3a gene. Numbers denote the amino acid positions of three domains in zebrafish Capn3a protein and the length of the Capn3a−/− mutant protein. g, Relative mRNA expression of capn3a in wild-type and capn3a−/− embryos at 2.5 dpf. h, i, Relative mRNA expression of capn3a family members in wild-type (+/+), capn3a morphant (MO) and capn3a−/− (−/−) embryos at 1.5 dpf. j, WISH of capn8 and capn12 in wild-type and capn3a−/− embryos at 4 dpf. Black arrow denotes the liver (lv). k, WISH using the lfabp probe was performed to assess the liver sizes in 3.5-dpf wild-type embryos injected with different mRNA as indicated. l, Statistical analysis of the average liver size of the samples as shown in k. m, Relative mRNA expression of capn8 and capn12 in wild-type and capn3a−/− embryos injected with (+) or without (−) capn3a mRNA at 2 dpf. n, Alignment of cDNA sequences between capn3a and its family members. The identity of cDNA sequences and the GeneBank accession number of each gene are provided. o, Relative mRNA expression of capn8 and capn12 in wild-type (+/+), capn3a+/− (+/−) and capn3a−/− (−/−) embryos at 2 dpf. Each experiment was repeated three times with similar results and a representative is shown here. n indicates the number of zebrafish embryos in each group. Data are mean ± s.d. from three biological replicates. *P < 0.05, **P < 0.01, ***P < 0.001, two-tailed t-test.
Extended Data Fig. 2 The GCR is triggered in capn3a genetic mutants carrying a functional PTC.
a, Schematic diagram showing an additional five mutant alleles (mu2–mu6) of the zebrafish capn3a gene. Black line denotes intron; black bar denotes exon; red line denotes deletion or insertion of DNA in different capn3a mutant alleles. b–f, Left, relative mRNA expression of capn3a family members in wild-type and different capn3a mutant alleles at 3 dpf. Right, statistical analysis of the average liver sizes in wild-type and different capn3a mutant alleles at 3.5 dpf. Each experiment was repeated three times with similar results and a representative result is shown. n indicates the number of zebrafish embryos in each group. Data are mean ± s.d. from three biological replicates. *P < 0.05, **P < 0.01, ***P < 0.001, two-tailed t-test.
Extended Data Fig. 3 The GCR is differentially activated in nid1a genetic mutants.
a, Alignment of cDNA sequences between nid1a and its family members. The identity of cDNA sequences and the GeneBank accession number of each gene were provided. b, Relative mRNA expression of nid1a in wild-type and nid1ad2/+ embryos at different times as indicated. c, Schematic diagram showing a nid1a mutant allele (nid1a-atgdel) with a deletion in the start codon (the first ATG). Black lines denote introns; black bars denote exons; ATG (surrounded by a black box) denotes the start codon; red horizontal line through the letters denotes deletion of DNA sequence in nid1a-atgdel mutant allele. d, Images of the wild-type and nida-atgdel larvae at 3, 4 and 5 dpf. e, Statistical analysis of the average body lengths in samples as shown in d. f, Relative mRNA expression of nid1a, nid1b, nid2a and nid2b in wild-type and nid1a-atgdel embryos at 1.5 dpf. Each experiment was repeated three times with similar results, and a representative result is shown. n indicates the number of zebrafish embryos in each group. Data are mean ± s.d. from three biological replicates. *P < 0.05, **P < 0.01, ***P < 0.001, two-tailed t-test.
Extended Data Fig. 4 GCR is triggered by a transgene with a PTC and the nucleotide sequence homologous shared by the compensatory endogenous gene.
a, Relative mRNA levels of capn3a, capn8 and capn12 in wild-type and capn3a−/− embryos injected with uncapped egfp RNA, capped capn3a RNA, uncapped capn3a RNA, capped capn3aΔ14 RNA or uncapped capn3aΔ14 RNA at 1.5 dpf. b, Relative transcript levels of the transgene in different transgenic lines injected with or without upf1-MO at 1.5 dpf as indicated. c, Relative transcript levels of the transgene in different transgenic lines treated with or without cycloheximide (CHX) at 3 dpf. d, Relative expression of endogenous nid1a mRNA in different transgenic lines at 3 dpf as indicated. Each experiment was repeated three times with similar results, and a representative result is shown. n indicates the number of zebrafish embryos in each group. Data are mean ± s.d. from three biological replicates. *P < 0.05, **P < 0.01, ***P < 0.001, two-tailed t-test.
Extended Data Fig. 5 Knockdown of upf3a impairs GCR in both capn3a−/− and nid1a−/− embryos.
a, Relative mRNA expression of capn3a, capn8 and capn12 in wild-type and capn3a−/− embryos treated with or without cycloheximide at 2 dpf. Mu, capn3a−/− embryos. b, Top, WISH was performed to assess the liver sizes in wild-type and capn3a−/− embryos injected with different reagents as indicated at 3.5 dpf. Bottom, statistical analysis of the average liver sizes. c, Relative mRNA expression of capn3a, capn8 and capn12 in wild-type and capn3a−/− embryos injected with or without upf3a-MO at 1.5 dpf. d, Images of the wild-type and nidaΔ2/Δ2 mutant (designated nid1a−/−) larvae injected with different reagents as indicated at 3, 4 and 5 dpf. e, Statistical analysis of the average body lengths in samples as shown in d. Each experiment was repeated three times with similar results, and a representative result is shown. n indicates the number of zebrafish embryos in each group. Data are mean ± s.d. from three biological replicates. *P < 0.05, **P < 0.01, ***P < 0.001, two-tailed t-test.
Extended Data Fig. 6 Knockdown of upf1, upf2 or upf3b has no effects on GCR in both capn3a−/− and nid1a−/− embryos.
a–f, WISH was performed to assess the liver sizes in 3.5-dpf wild-type and capn3a−/− embryos injected with different reagents as indicated (a, c, e). Statistical analysis of the average liver sizes corresponding to a, c and e is shown in b, d and f, respectively. g, Relative mRNA expression of capn3a, capn8 and capn12 in wild-type and capn3a−/− embryos injected with or without upf2-1#MO at 1.5 dpf. h, Left, validation of the second gene-specific morpholino of upf2 (upf2-2#MO). The upf2-2#MO was designed to block the splicing site of exon3–intron3 of upf2. RT–PCR was performed with a pair of primers (pf, forward primer; pv, reverse primer) to distinguish the mRNA produced by the wild-type embryos injected with ST-MO from that injected with upf2-2#MO. DNA fragment of 1,638 bp contains 105 bp of exon 3, 61 bp of exon 4 and the whole intron 3 of upf2. Right, relative mRNA expression of capn3a, capn8 and capn12 in wild-type and capn3a−/− embryos injected with different morpholinos as indicated at 1.5 dpf. i, j, Relative mRNA expression of capn3a, capn8 and capn12 in wild-type embryos, wild-type embryos injected with upf3b-MO (i) or upf1-MO (j), capn3a−/− embryos, or capn3a−/− embryos injected with upf3b-MO or upf1-MO at 1.5 dpf. k–m, Statistical analysis of the average body lengths in 4-dpf wild-type and nid1a−/− larvae injected with or without different morpholinos as indicated. n, Relative mRNA expression of endogenous capn3a in Tg1-1# and Tg3-1# embryos injected with or without upf3a-MO. Each experiment was repeated three times with similar results, and a representative result is shown. n indicates the number of zebrafish embryos in each group. Data are mean ± s.d. from three biological replicates. *P < 0.05, **P < 0.01, ***P < 0.001, two-tailed t-test.
Extended Data Fig. 7 Knockout of upf3a, but not upf1 or upf3b, impairs GCR in capn3a−/− and nid1a−/− embryos.
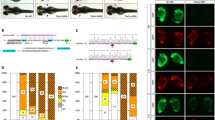
a, Schematic diagram showing the genomic structure and a genetic mutation in zebrafish upf3a gene. ATG denotes the translation start codon; black lines denote introns; blue vertical bars denote exons. Red horizontal line through the letters denotes deletion of 17 bp in the upf3a gene. Numbers denote the amino acid positions of two domains in zebrafish Upf3a protein, and the length of the Upf3a−/− mutant protein. b, Western blot analysis of Upf3a protein in wild-type and upf3a−/− embryos at 12 hpf. Actb1 was used as the protein loading control. MZ, maternal–zygotic mutant embryos. c, Schematic diagram showing the genomic structure and a genetic mutation of zebrafish upf1 gene. Underlined residues denote the additional 4 bp in the upf1 gene. d, Schematic diagram showing the genomic structure and a genetic mutation of the zebrafish upf3b gene. Red horizontal line denotes deletion of 1 bp in the upf3b gene. e, Statistical analysis of the average liver sizes of three genotypes from the in-cross of upf3a+/−;capn3a−/− as indicated at 3.5 dpf. The number of embryos for each genotype is shown. upf3a none-MZ denotes upf3a none-maternal–zygotic mutant embryos. f, WISH of upf3a in the offspring from the in-cross of upf3a+/− at the one-cell stage. g, Images of zebrafish larvae of different genotypes at 3, 4 and 5 dpf as shown. All larvae were from the in-cross of two homozygous parents (either single or double homozygous mutants). h, Statistical analysis of the average body lengths in the samples as shown in g. i, Statistical analysis of the average body lengths of three genotypes from the in-cross of upf3a+/−;nid1a−/− embryos at 4 dpf as indicated. j, Statistical analysis of the average body lengths in the 4-dpf larvae of four genotypes as indicated. The wild-type and upf1−/− embryos were from the in-cross of upf1+/−. The nid1a−/− and upf1−/−;nid1a−/− embryos were from the incross of upf1+/−;nid1a−/−. k, Statistical analysis of the average body lengths in the 4-dpf larvae of the four genotypes as indicated. The wild-type and upf3b−/− embryos were from the in-cross of upf3b+/−. The nid1a−/− and upf3b−/−;nid1a−/− embryos were from the in-cross of upf3b+/−;nid1a−/−. Each experiment was repeated three times with similar results, and a representative result is shown. n indicates the number of zebrafish embryos in each group. Data are mean ± s.d. from three biological replicates. *P < 0.05, **P < 0.01, ***P < 0.001, two-tailed t-test.
Extended Data Fig. 8 Depletion of PTC-bearing mRNA impairs GCR in capn3a−/− embryos.
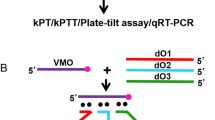
a, b, WISH was performed to assess the liver sizes in 3.5-dpf capn3a−/− embryos injected with different reagents as indicated. c, Relative expression of capn8 and capn12 in wild-type embryos injected with different reagents. d–f, qRT–PCR (d), western blot analysis (e) or WISH (f) in wild-type embryos injected with different reagents. Actb1 was used as a loading control. g, Top, a capn3a mutant mRNA (capn3a3pm), which is resistant to the degradation mediated by Cas13a gRNA, was designed. Red letters denote nucleotide mutations in the target sites; red stars denote the target sites of three gRNAs. Bottom, statistical analysis of the average liver sizes in wild-type embryos injected with or without Cas13a and non-specific gRNA (NS), Cas13a and gRNA (G), or Cas13a and gRNA plus capn3a3pm mRNA at 3.5 dpf. h, Statistical analysis of the average liver sizes in capn3a−/− embryos injected with or without Cas13a and non-specific gRNA (NS) or Cas13a and gRNA (G) at 3.5 dpf. Each experiment was repeated three times with similar results, and a representative result is shown. n indicates the number of zebrafish embryos in each group. Data are mean ± s.d. from three biological replicates. *P < 0.05, **P < 0.01, ***P < 0.001, two-tailed t-test.
Extended Data Fig. 9 ChIP analysis of the occupancies of H3K4me3 and RNA Pol II at different genomic regions in capn3a−/− and wild-type embryos.
a, Western blot analysis of total levels of H3K4me3 in wild-type and capn3a−/− embryos at 1, 2 and 3 dpf. Total histone 3 (H3) protein was used as a loading control. b, A genome browser screenshot of public H3K4me3 data around the TSS regions of capn8 and capn12 in wild-type zebrafish embryos at the stage of 30% epiboly, cited from the UCSC Genome Browser on Zebrafish Jul, 2010 (Zv9/danRer7) Assembly. c, d, ChIP–qPCR analysing the occupancies of H3K4me3 (c) and RNA Pol II (d) at different regions of capn8 and capn12 in wild-type and capn3a−/− embryos. The experiments were performed as described in Fig. 3c, d, e. Four pairs of primers (A, B, C and D) were designed to amplify the corresponding regions of capn8 or capn12 genes (light blue arrows). The occupancy was presented as a percentage of input. The TSS of gapdh was used as a positive control for the H3K4me3 and Pol II ChIP assays. Exon 11 of gapdh and an intergenic region were used as negative controls for the H3K4me3 and Pol II ChIP assays, respectively. The data from the B pair primers for H3K4me3 and the B and C pair primers for Pol II are presented in Fig. 3d, e. e, ChIP–qPCR analysing H3K4me3 occupancy at the TSS of capn8 or capn12 in wild-type and capn3a−/− embryos injected with or without upf3a-MO. The experiment was performed as described in Fig. 3f. The promoter region (A1) of capn2b and an intergenic region were used as two negative controls. The data from the B-pair of primers were presented in Fig. 3f. Each experiment was repeated three times with similar results, and a representative result is shown. n indicates the number of zebrafish embryos in each group. Data are mean ± s.d. from three biological replicates. *P < 0.05, **P < 0.01, ***P < 0.001, two-tailed t-test.
Extended Data Fig. 10 Depletion of the members of COMPASS complex impairs GCR in capn3a−/− embryos.
a, Statistical analysis of the average liver sizes in wild-type, capn3a−/−, wild-type embryos injected with ST-MO, capn3a-MO or wdr5-MO, and capn3a−/− embryos injected with ST-MO or wdr5-MO at 3.5 dpf. b, qRT–PCR analysis of capn8 and capn12 mRNA expression in wild-type and capn3a−/− embryos injected with ST-MO or wdr5-MO as shown. The injection of ST-MO was used as a negative control and given a value of one. c, WISH of capn8 in the wild-type and capn3a−/− embryos at around 1 dpf. Left, wild-type and capn3a−/− embryos without injection. Right, wild-type and capn3a−/− embryos with wdr5-MO injection. Red arrow denotes the region with upregulated capn8 expression; green arrow denotes the region with downregulated capn8 expression after wdr5-MO injection. d, Relative mRNA expression of capn8 in wild-type embryos injected with or without wdr5-MO, the wdr5-overexpression plasmid (wdr5-p), or wdr5-MO together with wdr5 plasmid, and in capn3a−/− embryos injected with or without wdr5-MO, the wdr5-overexpression plasmid (wdr5-p), or wdr5-MO together with wdr5 plasmid. e, Schematic diagram showing the genomic structure and a genetic mutation of the zebrafish wdr5 gene. ATG denotes the translation start codon; black lines denote introns; vertical bars denote exons (blue for UTR regions, black for the coding region). The red underlined sequence denotes the additional DNA (10 bp) in the wdr5 mutant; the numbers in the fifth and sixth panels denote the amino acid positions of one domain in zebrafish Wdr5 protein, and the length of predicted Wdr5 mutant protein (Wdr5i10). f, WISH was performed to assess the liver sizes in 3.5-dpf-old wild-type and capn3a−/− embryos injected with different reagents as indicated. Ctr, uninjected control. g, Statistical analysis of the average liver sizes in f under the different treatments as indicated. h, Relative mRNA expression of capn8 and capn12 in 1.5-dpf-old wild-type and capn3a−/− embryos with different treatments as indicated. The injection of ST-MO was used as a negative control and given a value of one. i–k, qRT–PCR analysis of capn8 and capn12 mRNA expression in wild-type and capn3a−/− embryos injected with ST-MO or setd1a-2#MO (i), rbbp5-2#MO (j), or ash2l-2#MO (k). The injection of ST-MO was used as a negative control and given a value of one. Each experiment was repeated three times with similar results, and a representative result is shown. n indicates the number of zebrafish embryos in each group. Data are mean ± s.d. from three biological replicates. *P < 0.05, **P < 0.01, ***P < 0.001, two-tailed t-test.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures which show source gel data scans and a Supplementary Table. A detailed guide of the figures and table is provided in a separate supplementary file.
Supplementary Information
This file contains a detailed guide for Supplementary Figures 1 and 2, and Supplementary Table 1.
Rights and permissions
About this article
Cite this article
Ma, Z., Zhu, P., Shi, H. et al. PTC-bearing mRNA elicits a genetic compensation response via Upf3a and COMPASS components. Nature 568, 259–263 (2019). https://doi.org/10.1038/s41586-019-1057-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1057-y
This article is cited by
-
The Mediator Subunit OsMED16 Interacts with the WRKY Transcription Factor OsWRKY45 to Enhance Rice Resistance Against Magnaporthe oryzae
Rice (2024)
-
miR-430 regulates zygotic mRNA during zebrafish embryogenesis
Genome Biology (2024)
-
HiHo-AID2: boosting homozygous knock-in efficiency enables robust generation of human auxin-inducible degron cells
Genome Biology (2024)
-
Evolutionary origin of germline pathogenic variants in human DNA mismatch repair genes
Human Genomics (2024)
-
dCas13-mediated translational repression for accurate gene silencing in mammalian cells
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.