Abstract
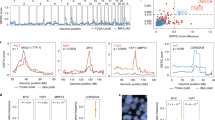
Metastasis is the main cause of death for patients with breast cancer. Many studies have characterized the genomic landscape of breast cancer during its early stages. However, there is evidence that genomic alterations are acquired during the evolution of cancers from their early to late stages, and that the genomic landscape of early cancers is not representative of that of lethal cancers1,2,3,4,5,6,7. Here we investigated the landscape of somatic alterations in 617 metastatic breast cancers. Nine driver genes (TP53, ESR1, GATA3, KMT2C, NCOR1, AKT1, NF1, RIC8A and RB1) were more frequently mutated in metastatic breast cancers that expressed hormone receptors (oestrogen and/or progesterone receptors; HR+) but did not have high levels of HER2 (HER2−; n = 381), when compared to early breast cancers from The Cancer Genome Atlas. In addition, 18 amplicons were more frequently observed in HR+/HER2− metastatic breast cancers. These cancers showed an increase in mutational signatures S2, S3, S10, S13 and S17. Among the gene alterations that were enriched in HR+/HER2− metastatic breast cancers, mutations in TP53, RB1 and NF1, together with S10, S13 and S17, were associated with poor outcome. Metastatic triple-negative breast cancers showed an increase in the frequency of somatic biallelic loss-of-function mutations in genes related to homologous recombination DNA repair, compared to early triple-negative breast cancers (7% versus 2%). Finally, metastatic breast cancers showed an increase in mutational burden and clonal diversity compared to early breast cancers. Thus, the genomic landscape of metastatic breast cancer is enriched in clinically relevant genomic alterations and is more complex than that of early breast cancer. The identification of genomic alterations associated with poor outcome will allow earlier and better selection of patients who require the use of treatments that are still in clinical trials. The genetic complexity observed in advanced breast cancer suggests that such treatments should be introduced as early as possible in the disease course.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Data are available at https://www.ebi.ac.uk/ega/ (EGAS00001003290). In addition, the data and scripts used to generate the figures are accessible at https://github.com/gustaveroussy/mBC_WES_Fabrice_Andre_2019, or on request from the corresponding author.
Change history
16 July 2019
An Amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Navin, N. et al. Tumour evolution inferred by single-cell sequencing. Nature 472, 90–94 (2011).
Wang, Y. et al. Clonal evolution in breast cancer revealed by single nucleus genome sequencing. Nature 512, 155–160 (2014).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Yates, L. R. et al. Genomic evolution of breast cancer metastasis and relapse. Cancer Cell 32, 169–184 (2017).
Ng, C. K. Y. et al. Genetic heterogeneity in therapy-naïve synchronous primary breast cancers and their metastases. Clin. Cancer Res. 23, 4402–4415 (2017).
Yu, H. A. et al. Analysis of tumor specimens at the time of acquired resistance to EGFR-TKI therapy in 155 patients with EGFR-mutant lung cancers. Clin. Cancer Res. 19, 2240–2247 (2013).
Gramza, A. W., Corless, C. L. & Heinrich, M. C. Resistance to tyrosine kinase inhibitors in gastrointestinal stromal tumors. Clin. Cancer Res. 15, 7510–7518 (2009).
Robinson, D. R. et al. Activating ESR1 mutations in hormone-resistant metastatic breast cancer. Nat. Genet. 45, 1446–1451 (2013).
Fumagalli, D. et al. Somatic mutation, copy number and transcriptomic profiles of primary and matched metastatic estrogen receptor-positive breast cancers. Ann. Oncol. 27, 1860–1866 (2016).
Lefebvre, C. et al. Mutational profile of metastatic breast cancers: a retrospective analysis. PLoS Med. 13, e1002201 (2016).
Brastianos, P. K. et al. Genomic characterization of brain metastases reveals branched evolution and potential therapeutic targets. Cancer Discov. 5, 1164–1177 (2015).
Brown, D. et al. Phylogenetic analysis of metastatic progression in breast cancer using somatic mutations and copy number aberrations. Nat. Commun. 8, 14944 (2017).
Savas, P. et al. The subclonal architecture of metastatic breast cancer: results from a prospective community-based rapid autopsy program “CASCADE”. PLoS Med. 13, e1002204 (2016).
Murtaza, M. et al. Multifocal clonal evolution characterized using circulating tumour DNA in a case of metastatic breast cancer. Nat. Commun. 6, 8760 (2015).
Schrijver, W. A. M. E. et al. Mutation profiling of key cancer genes in primary breast cancers and their distant metastases. Cancer Res. 78, 3112–3121 (2018).
De Mattos-Arruda, L. et al. Genetic heterogeneity and actionable mutations in HER2-positive primary breast cancers and their brain metastases. Oncotarget 9, 20617–20630 (2018).
Razavi, P. et al. The genomic landscape of endocrine-resistant advanced breast cancers. Cancer Cell 34, 427–438 (2018).
Nayar, U. et al. Acquired HER2 mutations in ER+ metastatic breast cancer confer resistance to estrogen receptor-directed therapies. Nat. Genet. 51, 207–216 (2019).
Li, Z. et al. Loss of the FAT1 tumor suppressor promotes resistance to CDK4/6 inhibitors via the hippo pathway. Cancer Cell 34, 893–905 (2018).
Knudsen, E. S. & Wang, J. Y. J. Targeting the RB-pathway in cancer therapy. Clin. Cancer Res. 16, 1094–1099 (2010).
Lock, R. et al. Cotargeting MNK and MEK kinases induces the regression of NF1-mutant cancers. J. Clin. Invest. 126, 2181–2190 (2016).
Gala, K. et al. KMT2C mediates the estrogen dependence of breast cancer through regulation of ERα enhancer function. Oncogene 37, 4692–4710 (2018).
Chakravarty, D. et al. OncoKB: a precision oncology knowledge base. JCO Precis. Oncol. https://doi.org/10.1200/PO.17.00011 (2017).
Law, E. K. et al. The DNA cytosine deaminase APOBEC3B promotes tamoxifen resistance in ER-positive breast cancer. Sci. Adv. 2, e1601737 (2016).
Popova, T. et al. Ploidy and large-scale genomic instability consistently identify basal-like breast carcinomas with BRCA1/2 inactivation. Cancer Res. 72, 5454–5462 (2012).
Riaz, N. et al. Pan-cancer analysis of bi-allelic alterations in homologous recombination DNA repair genes. Nat. Commun. 8, 857 (2017).
Polak, P. et al. A mutational signature reveals alterations underlying deficient homologous recombination repair in breast cancer. Nat. Genet. 49, 1476–1486 (2017).
Edwards, S. L. et al. Resistance to therapy caused by intragenic deletion in BRCA2. Nature 451, 1111–1115 (2008).
Sakai, W. et al. Secondary mutations as a mechanism of cisplatin resistance in BRCA2-mutated cancers. Nature 451, 1116–1120 (2008).
Lee, J. Y. et al. Lobular carcinomas in situ display intralesion genetic heterogeneity and clonal evolution in the progression to invasive lobular carcinoma. Clin. Cancer Res. 25, 674–686 (2019).
André, F. et al. Comparative genomic hybridisation array and DNA sequencing to direct treatment of metastatic breast cancer: a multicentre, prospective trial (SAFIR01/UNICANCER). Lancet Oncol. 15, 267–274 (2014).
Massard, C. et al. High-throughput genomics and clinical outcome in hard-to-treat advanced cancers: results of the MOSCATO 01 trial. Cancer Discov. 7, 586–595 (2017).
Le Tourneau, C. et al. Molecularly targeted therapy based on tumour molecular profiling versus conventional therapy for advanced cancer (SHIVA): a multicentre, open-label, proof-of-concept, randomised, controlled phase 2 trial. Lancet Oncol. 16, 1324–1334 (2015).
Hortobagyi, G. N. et al. Ribociclib as first-line therapy for HR-positive, advanced breast cancer. N. Engl. J. Med. 375, 1738–1748 (2016).
Tripathy, D. et al. Ribociclib plus endocrine therapy for premenopausal women with hormone-receptor-positive, advanced breast cancer (MONALEESA-7): a randomised phase 3 trial. Lancet Oncol. 19, 904–915 (2018).
Slamon, D. J. et al. Phase III randomized study of ribociclib and fulvestrant in hormone receptor-positive, human epidermal growth factor receptor 2-negative advanced breast cancer: MONALEESA-3. J. Clin. Oncol. 36, 2465–2472 (2018).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Saunders, C. T. et al. Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics 28, 1811–1817 (2012).
Chang, M. T. et al. Accelerating discovery of functional mutant alleles in cancer. Cancer Discov. 8, 174–183 (2018).
Gao, J. et al. 3D clusters of somatic mutations in cancer reveal numerous rare mutations as functional targets. Genome Med. 9, 4 (2017).
Costello, M. et al. Discovery and characterization of artifactual mutations in deep coverage targeted capture sequencing data due to oxidative DNA damage during sample preparation. Nucleic Acids Res. 41, e67 (2013).
Shen, R. & Seshan, V. E. FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing. Nucleic Acids Res. 44, e131 (2016).
Riester, M. et al. PureCN: copy number calling and SNV classification using targeted short read sequencing. Source Code Biol. Med. 11, 13 (2016).
Ye, K., Schulz, M. H., Long, Q., Apweiler, R. & Ning, Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 25, 2865–2871 (2009).
Schröder, J. et al. Socrates: identification of genomic rearrangements in tumour genomes by re-aligning soft clipped reads. Bioinformatics 30, 1064–1072 (2014).
Newman, A. M. et al. Integrated digital error suppression for improved detection of circulating tumor DNA. Nat. Biotechnol. 34, 547–555 (2016).
The Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Rosenthal, R., McGranahan, N., Herrero, J., Taylor, B. S. & Swanton, C. DeconstructSigs: delineating mutational processes in single tumors distinguishes DNA repair deficiencies and patterns of carcinoma evolution. Genome Biol. 17, 31 (2016).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Roth, A. et al. PyClone: statistical inference of clonal population structure in cancer. Nat. Methods 11, 396–398 (2014).
Hundal, J. et al. pVAC-Seq: a genome-guided in silico approach to identifying tumor neoantigens. Genome Med. 8, 11 (2016).
McGranahan, N. et al. Clonal neoantigens elicit T cell immunoreactivity and sensitivity to immune checkpoint blockade. Science 351, 1463–1469 (2016).
Piscuoglio, S. et al. The genomic landscape of male breast cancers. Clin. Cancer Res. 22, 4045–4056 (2016).
Leiserson, M. D., Wu, H. T., Vandin, F. & Raphael, B. J. CoMEt: a statistical approach to identify combinations of mutually exclusive alterations in cancer. Genome Biol. 16, 160 (2015).
Acknowledgements
The SAFIR02 trial is supported by Fondation ARC. Sequencing was supported by French NCI (INCA), Breast Cancer Research Foundation, Operation Parrain Chercheurs, RHU MyProbe (ANR) and Odyssea. S.P. is supported by the Swiss National Science Foundation (Ambizione grant number PZ00P3_168165). F.B. and D.B. are supported by SIRIC, label Ligue EL2016, Ruban Rose and Fondation Groupe EDF. We thank all patients who consented to enter the study, and all investigators and their teams.
Reviewer information
Nature thanks Peter Campbell and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
F.B. designed the study, is the principal investigator (PI) of the PERMED trial and contributed to writing of the paper. C.K.Y.N. and S.P. designed the study, supervised bioinformatics pipelines, generated data from Variant Call Format (VCF) and contributed to writing of the paper. A.P. is the PI of the RUBY trial, designed whole-genome analyses and acquired samples in the SAFIR02 trial. N.D. ran the whole-exome and whole-genome sequencing profiling. N.C. created libraries from PERMED samples. J.C.S. is the PI of the MOSCATO and MATCH-R trials and provided samples from these trials. A.T.D. and Y.A. generated VCF files from raw whole-exome sequencing data and whole-genome sequencing data. M.K. provided DNA from the SHIVA trial. S.G. generated libraries from the PERMED trial. G.M. supervised A.T.D. and Y.A. M.J. is the project manager of the SAFIR01, SAFIR02 and RUBY trials. S.D. and B.V. contributed to writing of the paper. M. Chaffanet and D.B. acquired samples from PERMED and supervised the production of libraries from samples. T.B., M. Campone, C.L., H.B. and F.D. acquired samples for the SAFIR01 and SAFIR02 trials. A.J. is the project manager of the SAFIR02 trial and centralized collected samples and data. M.R.D.F. generated data from VCF files under the supervision of C.K.Y.N. and S.P. N.B. is the translational research head of the MONALEESA trial. T.F. ran all statistical analyses related to outcomes and contributed to the writing. C.L.T. is the PI of the SHIVA trial and helped to design the study. F.A. designed the study, is the PI of the SAFIR01 and SAFIR02 trials, and contributed to the writing. All authors approved the final manuscript and contributed to critical revisions of its intellectual content.
Corresponding author
Ethics declarations
Competing interests
N.B. is an employee of Novartis. F.A. and T.B. received grants from Novartis. F.A. attended the advisory board and gave talks at events sponsored by Novartis. Gustave Roussy Hospital was compensated for these. M. Campone, F.D. and T.B. received honoraria from Novartis. The RUBY trial is funded by Clovis. The MONALEESA trials are sponsored by Novartis. J.C.S. is an employee and holds stock in AstraZeneca since September 2017. Over the past five years he has received consultancy fees from: AstraZeneca, Astex, Clovis, GSK, GamaMabs, Lilly, MSD, Mission Therapeutics, Merus, Pfizer, PharmaMar, Pierre Fabre, Roche-Genentech, Sanofi, Servier, Symphogen and Takeda. He is also a shareholder in Gritstone.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
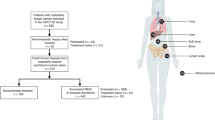
Extended Data Fig. 1 Study workflow.
The main dataset included tumour samples from 617 patients with metastatic breast cancers who were included in six precision medicine trials. The aim was to generate the genomic landscape of metastatic breast cancers. We then explored the clinical relevance of gene alterations that were enriched in mBCs. To address these questions, we used the main dataset to test the prognostic values of these gene alterations; we used genomic data from the MONALEESA2, 3 and 7 trials to test the efficacy of CDK4 inhibitors in RB1 mutant tumours; we used whole-genome sequencing data from four outlier responders to PARP inhibitors (RUBY trial); and we used data from SAFIR02 to explore the efficacy of MEK inhibitors in patients with NF1 mutations.
Extended Data Fig. 2 Outcome of patients with HR+/HER2− mBCs according to the presence of mutational signatures S10, S13 and S17.
Kaplan–Meier plots of overall survival in patients with HR+/HER2− mBCs, stratified by mutational signatures S10 (S10 = 0, n = 171; S10 > 0, n = 194), S13 (S13 = 0, n = 140; S13 > 0, n = 225) and S17 (S17 = 0, n = 245; S17 > 0, n = 120). HR, hazard ratio according to Cox multivariate analysis (log-rank test, two-sided); 95% CI shown in square brackets.
Extended Data Fig. 3 Detection of LST, signature 3 and indels with microhomology according to the presence of biallelic LOF mutations on genes located in the HR pathway.
Scatter plots of LST and contributions of signature 3 in the HR+/HER2−, HER2+ and TNBC cohorts. Each dot represents one tumour and the sizes of the dots are proportion to the number of indels larger than 3 bp with microhomology, a feature of HRD. Tumours associated with biallelic LOF mutations (frameshift, start/stop codon, nonsense and splice sites) in HR-related genes in which one of the alleles lost was a germline variant are shown in purple, and tumours associated with biallelic LOF mutations in which both alleles were lost somatically are shown in red. All cases in the darker pink shaded areas were considered to have HRD, and cases in the lighter pink shaded areas were considered to have HRD if they were LST-high25.
Extended Data Fig. 4 Prediction of tumour neoantigens according to mutational processes.
Scatter plots of the fraction of mutations classified as neoantigens against the number of neoantigens for neoantigens with predicted binding affinity <100 nM (left) and neoantigens with predicted binding affinity <500 nM (right). Above and to the right of the scatter plots are the kernel density plots of the number and fraction of neoantigens, respectively. Each dot represents a sample and is colour-coded according to mutational signature and/or clinical subgroup (see Methods).
Supplementary information
Rights and permissions
About this article
Cite this article
Bertucci, F., Ng, C.K.Y., Patsouris, A. et al. Genomic characterization of metastatic breast cancers. Nature 569, 560–564 (2019). https://doi.org/10.1038/s41586-019-1056-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1056-z
This article is cited by
-
Mutational landscape of inflammatory breast cancer
Journal of Translational Medicine (2024)
-
MYC induces CDK4/6 inhibitors resistance by promoting pRB1 degradation
Nature Communications (2024)
-
CDK4/6i-treated HR+/HER2- breast cancer tumors show higher ESR1 mutation prevalence and more altered genomic landscape
npj Breast Cancer (2024)
-
p4EBP1 staining predicts outcome in ER-positive endocrine-resistant metastatic breast cancer patients treated with everolimus and exemestane
British Journal of Cancer (2024)
-
Navigating Breast Cancer Oligometastasis and Oligoprogression: Current Landscape and Future Directions
Current Oncology Reports (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.