Abstract
Hepatocellular carcinoma is the third leading cause of deaths from cancer worldwide. Infection with the hepatitis B virus is one of the leading risk factors for developing hepatocellular carcinoma, particularly in East Asia1. Although surgical treatment may be effective in the early stages, the five-year overall rate of survival after developing this cancer is only 50–70%2. Here, using proteomic and phospho-proteomic profiling, we characterize 110 paired tumour and non-tumour tissues of clinical early-stage hepatocellular carcinoma related to hepatitis B virus infection. Our quantitative proteomic data highlight heterogeneity in early-stage hepatocellular carcinoma: we used this to stratify the cohort into the subtypes S-I, S-II and S-III, each of which has a different clinical outcome. S-III, which is characterized by disrupted cholesterol homeostasis, is associated with the lowest overall rate of survival and the greatest risk of a poor prognosis after first-line surgery. The knockdown of sterol O-acyltransferase 1 (SOAT1)—high expression of which is a signature specific to the S-III subtype—alters the distribution of cellular cholesterol, and effectively suppresses the proliferation and migration of hepatocellular carcinoma. Finally, on the basis of a patient-derived tumour xenograft mouse model of hepatocellular carcinoma, we found that treatment with avasimibe, an inhibitor of SOAT1, markedly reduced the size of tumours that had high levels of SOAT1 expression. The proteomic stratification of early-stage hepatocellular carcinoma presented in this study provides insight into the tumour biology of this cancer, and suggests opportunities for personalized therapies that target it.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this study—including clinical information, and proteome, phospho-proteome and gene expression data—are available within the paper and its Supplementary Information, or from CNHPP liver data portal (http://liver.cnhpp.ncpsb.org/). The thermo.raw files of proteome and phospho-proteome datasets can be obtained from PRIDE database (www.ebi.ac.uk/pride/archive, accession numbers PXD006512 and PXD008373)63 or iProX database (www.iprox.org, accession number IPX0000937000)64. Gene expression profiles by RNA-seq can be obtained from Gene Expression Omnibus (accession number GSE124535).
The uploading and sharing of individual genetic data from this project is not permissible, according to a review by the Human Genetic Resources Administration of China on the basis of regulations documented in the Interim Measures for the Administration of Human Genetic Resources. We have summaries of the data that are as detailed as possible, and which are available to other researchers. This includes the exonic and splicing mutations, given in Supplementary Table 3. Researchers who wish to gain access to allele frequency information and other summary statistics data are required to fill in a simple application form (http://liver.cnhpp.ncpsb.org/) and send an email to F.H. for identity verification purposes, to adhere to the Chinese regulations.
The TCGA RNA-seq data are publicly available at the Genomic Data Commons Data Portal (CDC, http://www.ncbi.nlm.nih.gov/geo). Additional publically available microarray data for HCC (Fudan cohort) was analysed from NCBI GEO repository (accession number GSE14520).
References
Forner, A., Reig, M. & Bruix, J. Hepatocellular carcinoma. Lancet 391, 1301–1314 (2018).
European Association for the Study of the Liver. EASL clinical practice guidelines: management of hepatocellular carcinoma. J. Hepatol. 69, 182–236 (2018).
Ho, D. W. H. et al. TSC1/2 mutations define a molecular subset of HCC with aggressive behaviour and treatment implication. Gut 66, 1496–1506 (2017).
Tornesello, M. L. et al. Mutations in TP53, CTNNB1 and PIK3CA genes in hepatocellular carcinoma associated with hepatitis B and hepatitis C virus infections. Genomics 102, 74–83 (2013).
Lachenmayer, A. et al. Wnt-pathway activation in two molecular classes of hepatocellular carcinoma and experimental modulation by sorafenib. Clin. Cancer Res. 18, 4997–5007 (2012).
The Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Roessler, S. et al. A unique metastasis gene signature enables prediction of tumor relapse in early-stage hepatocellular carcinoma patients. Cancer Res. 70, 10202–10212 (2010).
Hoshida, Y. et al. Integrative transcriptome analysis reveals common molecular subclasses of human hepatocellular carcinoma. Cancer Res. 69, 7385–7392 (2009).
Coulouarn, C., Factor, V. M. & Thorgeirsson, S. S. Transforming growth factor-β gene expression signature in mouse hepatocytes predicts clinical outcome in human cancer. Hepatology 47, 2059–2067 (2008).
Lee, J. S. et al. Classification and prediction of survival in hepatocellular carcinoma by gene expression profiling. Hepatology 40, 667–676 (2004).
Chiang, D. Y. et al. Focal gains of VEGFA and molecular classification of hepatocellular carcinoma. Cancer Res. 68, 6779–6788 (2008).
Sohn, B. H. et al. Inactivation of Hippo pathway is significantly associated with poor prognosis in hepatocellular carcinoma. Clin. Cancer Res. 22, 1256–1264 (2016).
Lee, J. S. et al. A novel prognostic subtype of human hepatocellular carcinoma derived from hepatic progenitor cells. Nat. Med. 12, 410–416 (2006).
Sia, D. et al. Identification of an immune-specific class of hepatocellular carcinoma, based on molecular features. Gastroenterology 153, 812–826 (2017).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013).
Zheng, C. et al. Landscape of infiltrating T cells in liver cancer revealed by single-cell sequencing. Cell 169, 1342–1356.e16 (2017).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453–457 (2015).
The Cancer Genome Atlas Research Network. Comprehensive and integrative genomic characterization of hepatocellular carcinoma. Cell 169, 1327–1341.e23 (2017).
Uhlen, M. et al. A pathology atlas of the human cancer transcriptome. Science 357, eaan2507 (2017).
Yue, S. et al. Cholesteryl ester accumulation induced by PTEN loss and PI3K/AKT activation underlies human prostate cancer aggressiveness. Cell Metab. 19, 393–406 (2014).
Li, J. et al. Abrogating cholesterol esterification suppresses growth and metastasis of pancreatic cancer. Oncogene 35, 6378–6388 (2016).
Pande, G. The role of membrane lipids in regulation of integrin functions. Curr. Opin. Cell Biol. 12, 569–574 (2000).
Shimano, H. & Sato, R. SREBP-regulated lipid metabolism: convergent physiology - divergent pathophysiology. Nat. Rev. Endocrinol. 13, 710–730 (2017).
Wiśniewski, J. R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 6, 359–362 (2009).
Ren, L. et al. TiO2 with tandem fractionation (TAFT): An approach for rapid, deep, reproducible, and high-throughput phosphoproteome analysis. J. Proteome Res. 17, 710–721 (2018).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protocols 2, 1896–1906 (2007).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
R Core Team. R: A language and environment for statistical computing www.R-project.org/ (R Foundation for Statistical Computing, Vienna, 2012).
Fraley, C. & Raftery, A. E. Model-based clustering, discriminant analysis, and density estimation. J. Am. Stat. Assoc. 97, 611–631 (2002).
Hartigan, J. A. & Hartigan, P. M. The dip test of unimodality. Ann. Stat. 13, 70–84 (1985).
Bolstad, B. M., Irizarry, R. A., Astrand, M. & Speed, T. P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003).
Zhang, B. et al. Proteogenomic characterization of human colon and rectal cancer. Nature 513, 382–387 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Saunders, C. T. et al. Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics 28, 1811–1817 (2012).
The 1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Hua, X. et al. DrGaP: a powerful tool for identifying driver genes and pathways in cancer sequencing studies. Am. J. Hum. Genet. 93, 439–451 (2013).
Forbes, S. A. et al. COSMIC: exploring the world’s knowledge of somatic mutations in human cancer. Nucleic Acids Res. 43, D805–D811 (2015).
Mayakonda, A., Lin, D. C., Assenov, Y., Plass, C. & Koeffler, H. P. Maftools: efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 28, 1747–1756 (2018).
Li, J.-W. et al. ViralFusionSeq: accurately discover viral integration events and reconstruct fusion transcripts at single-base resolution. Bioinformatics 29, 649–651 (2013).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-seq. Bioinformatics 25, 1105–1111 (2009).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protocols 7, 562–578 (2012).
Smyth, G. K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, e3 (2004).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Kim, P. M. & Tidor, B. Subsystem identification through dimensionality reduction of large-scale gene expression data. Genome Res. 13, 1706–1718 (2003).
Brunet, J. P., Tamayo, P., Golub, T. R. & Mesirov, J. P. Metagenes and molecular pattern discovery using matrix factorization. Proc. Natl Acad. Sci. USA 101, 4164–4169 (2004).
Levine, D. A. & The Cancer Genome Atlas Research Network. Integrated genomic characterization of endometrial carcinoma. Nature 497, 67–73 (2013).
The Cancer Genome Atlas Research Network. Comprehensive molecular characterization of gastric adenocarcinoma. Nature 513, 202–209 (2014).
Pascual-Montano, A., Carazo, J. M., Kochi, K., Lehmann, D. & Pascual-Marqui, R. D. Nonsmooth nonnegative matrix factorization (nsNMF). IEEE Trans. Pattern Anal. Mach. Intell. 28, 403–415 (2006).
Hoshida, Y. Nearest template prediction: a single-sample-based flexible class prediction with confidence assessment. PLoS ONE 5, e15543 (2010).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7 (2013).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Milacic, M. et al. Annotating cancer variants and anti-cancer therapeutics in reactome. Cancers (Basel) 4, 1180–1211 (2012).
Croft, D. et al. The reactome pathway knowledgebase. Nucleic Acids Res. 42, D472–D477 (2014).
Schaefer, C. F. et al. PID: the Pathway Interaction Database. Nucleic Acids Res. 37, D674–D679 (2009).
Bindea, G. et al. Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity 39, 782–795 (2013).
Ye, Q. H. et al. GOLM1 Modulates EGFR/RTK cell-surface recycling to drive hepatocellular carcinoma metastasis. Cancer Cell 30, 444–458 (2016).
Schulze, K. et al. Exome sequencing of hepatocellular carcinomas identifies new mutational signatures and potential therapeutic targets. Nat. Genet. 47, 505–511 (2015).
Llovet, J. M., Villanueva, A., Lachenmayer, A. & Finn, R. S. Advances in targeted therapies for hepatocellular carcinoma in the genomic era. Nat. Rev. Clin. Oncol. 12, 408–424 (2015).
Vorkas, P. A. et al. Untargeted UPLC-MS profiling pipeline to expand tissue metabolome coverage: application to cardiovascular disease. Anal. Chem. 87, 4184–4193 (2015).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Ma, J. et al. iProX: an integrated proteome resource. Nucleic Acids Res. 47, D1211–D1217 (2019).
Acknowledgements
This work was supported by Chinese Human Proteome Project, and partially supported by Chinese State Key Projects for Basic Research (973 Program) (no. 2014CBA02001), National Key R&D Program of China (no. 2016YFC0902400, 2017YFC0906603, 2018YFA0507502, 2016YFF0101405, 2013ZX10002009 and 2009ZX09503-002), Program of International S&T Cooperation (no. 2014DFB30020,2014DFB30030, 2014DFB30010, 2009DFB33070 and 2010DFA31260), National Natural Science Foundation of China (no. 81770581, 81570526, 81772551, 81802364, 8153077, 81672839, 81572823, 81772578, 8153077 and 81123001), Innovation project (16CXZ027), Beijing Science and Technology Project (Z161100002616036) and Shanghai 111 Project (B14019). We acknowledge the assistance of Z. Jin and P. Zhang for the immune-related data analyses, and C. Chang for the method development and evaluation of the missing-value imputing analyses (National Center for Protein Sciences (Beijing)). We thank L. Shen, W. Xue and Z. Li (Cancer Hospital & Institute, Peking University) for their examination of tumour cellularity and immune-cell infiltration; and B. Zhen, L. Tang and Y. Wang (State Key Laboratory of Proteomics, Beijing Proteome Research Center, National Center for Protein Sciences (Beijing), Beijing Institute of Lifeomics) for scientific management.
Reviewer information
Nature thanks Josep M. Llovet and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Consortia
Contributions
F.H., X.Q. J.F. and Y.J. directed and designed research; H.S., B.X., Y.J., W.S., N.C., A.S., X.Z., C.H., M.L. and W.L. coordinated acquisition, distribution and quality evaluation of HCC tumour, and nearby tissue, samples; X.Q., J.Q., W.Y., Yang Zhao, L.R., Manli Zhang, Yan Zhao, M.Y., M.W. and F.T. directed and performed analyses and quality control of mass spectrometry data; X.Y., B.H., Jian Zhou, and J.F. performed the tissue microarray, immunohistochemistry experiments and PDX mouse models experiments; C.L. and Jinan Zhou performed HCC cell-line experiments and the cholesterol staining and quantification experiments; Yang Zhao, G.Q., L.X., Menghuan Zhang, L.Z. and T.S. adapted algorithms and software for data analysis; Y.J., A.S., Yang Zhao and W.S. interpreted data in context of HCC biology; W.Z., Y.H., Y.Q., Y.S., H.X. and Z.W. designed and constructed the database of this study; Y.J., Yang Zhao, A.S., W.S., W.Y., X.Q. and F.H. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Overview of multi-omics landscape of HBV-related early-stage HCC and quality control for the mass spectrometry platform.
a, Schematic of the multi-omics analyses of HBV-related early-stage HCC. The number of proteins, phosphorylation sites, expressed genes and genomic mutations identified in the proteome, phospho-proteome, transcriptome and whole-genome sequencing (WGS) data are displayed on the right of the middle panel. b, Illustration of 110 paired HCC cases used in the individual omics experiments. The proteome, phospho-proteome, genome and transcriptome experiments are coloured red, orange, blue and green, respectively. For each omics dataset, the top panel represents the non-tumour tissues and the bottom panel represents the tumour tissues. c, Longitudinal quality control of mass spectrometry using tryptic digest of HEK293T cells. The bottom-left half of the panel represents the pairwise Spearman’s correlation coefficients of the samples, and the top-right half of the panel depicts the pairwise scatter plots from the same comparison. d, Scatter plots and Spearman’s correlation coefficients for replicate proteome profiling of six HCC samples (three paired tumour and non-tumour samples). The x and y axes represent the log2-transformed protein intensities in each pairwise comparison. Notably, repeat experiments with the same samples have good reproducibility, with a high level of correlation (average, 0.9; range, 0.88–0.92).
Extended Data Fig. 2 Quality assessment of proteomic data and genomic landscape of HBV-related early-stage HCC.
a, Distribution of log2-transformed iBAQ abundance of identified proteins in 199 proteome samples that passed quality control. Green presents non-tumour samples (n = 98), red denotes tumour samples (n = 101). In the box plots, the middle bar represents the median, and the box represents the interquartile range; bars extend to 1.5× the interquartile range. b, Cumulative number of protein identifications. Blue presents non-tumour samples (n = 98), red denotes tumour samples (n = 101). c, Cumulative number of phosphorylation site identifications. Blue presents non-tumour samples (n = 103), red denotes tumour samples (n = 103). d, The genomic landscape and mutational signatures in HBV-related early-stage HCC. Top, individual tumour mutation rates. Middle, AFP and MVI status of the 93 patients with HCC. Bottom, recurrently mutated genes. Mutation types are indicated in the legend at the bottom. The bar plot on the left shows the number of alterations for each gene in the 93 patients.
Extended Data Fig. 3 Protein identification and gene expression in HBV-related early-stage HCC.
a, Box plots of the proteins identified in tumour (red, n = 98) and paired non-tumour (green, n = 98) samples. b, Box plots of the protein identifications in AFPlow (grey, n = 51), AFPhigh (red, n = 34), MVI− (grey, n = 51) and MVI+ (red, n = 22) patients from the tumour (red, n = 98) or non-tumour (green, n = 98) samples. To eliminate interaction effects, the patients that are both AFPlow and MVI− are simultaneously considered to be AFPlow or MVI− patients in this analysis. c, Overview of the number of expressed genes in HBV-related early-stage HCC. Similar to the protein identification, the number of genes expressed in the tumour samples is higher than that in the paired non-tumour samples. The MVI+ or AFPhigh patients also have a higher level of gene expression than the MVI− or AFPlow patients in the tumour samples, but there are no statistical differences between the MVI+ or AFPhigh patients and the MVI− or AFPlow patients in the non-tumour samples. The samples are arranged in the same order as in Fig. 1a. Dashed lines denote paired samples from the same cases of HCC. Dashed curves are fitted by lasso regression. The shading that underlies the lasso curves represents the 95% confidence intervals of those lines. d, Box plots of the number of expressed genes in the tumour (red, n = 31) and paired non-tumour (red, n = 31) samples. e, Box plots of the expressed genes in AFPlow (grey, n = 12), AFPhigh (red, n = 14), MVI− (grey, n = 12) and MVI+ (red, n = 9) patients from the tumour (red, n = 31) or non-tumour (green, n = 31) samples. To eliminate interaction effects, patients that are both AFPlow and MVI− are simultaneously considered to be AFPlow or MVI− patients in this analysis. In the box plots, the middle bar represents the median, and the box represents the interquartile range; bars extend to 1.5× the interquartile range. P values are calculated by the two-sided Wilcoxon rank-sum test and are shown on the top of the boxes. NS, not significant, *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Fig. 4 Pathway alterations and the effects of genomic mutations on protein expression or phosphorylation in HBV-related early-stage HCC.
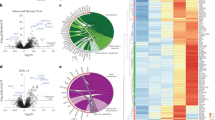
a, Pathway alterations in HBV-related early-stage HCC. Left, alteration pathways in the tumours (n = 101) versus the non-tumour tissues (n = 98). Middle and right, alteration pathways in the tumour samples from the AFPhigh (n = 34) versus AFPlow (n = 67) and MVI+ (n = 22) versus MVI− (n = 79) patients, respectively. P values are calculated by two-sided Fisher’s exact test based on the MSigDB database. Upregulated and downregulated pathways are indicated in red (right) and blue (left) bars. b, The activated pathways in the tumour samples (n = 92) are characterized by phospho-proteome data. The most significant pathways for the phospho-proteome are plotted on the x axis as the −log2 of the P value, compared with the proteome. P values are calculated by two-sided Fisher’s exact test based on the MSigDB database. c, Heat map of hyper-phosphorylated proteins across tumour (n = 92) and non-tumour (n = 92) tissues. The numbers of hyper- phosphorylation sites are shown for each protein and coloured in red, and proteins with no hyper-phosphorylation sites are coloured in white. Biological functions related to these proteins are denoted in the colour panel on the left. The samples are arranged in the same order as in Fig. 1. d, Heat map of dysregulated proteins by TSC1 and TSC2 mutation. The downstream substrate of mTOR, RPS6, was hyper-phosphorylated at the pS236, pS240 and pS244 sites. e, Heat map of dysregulated proteins by CTNNB1 mutation. The liver-related WNT targets GLUL and CYP1A2 were upregulated in tumour samples with CTNNB1 mutations.
Extended Data Fig. 5 NMF consensus-clustering identifies three proteomic subtypes of HBV-related early-stage HCC.
a, Principal component analysis; first two principal components of protein intensities, with samples connected by centroids according to sample types (green, non-tumour; red, tumour). The tumour samples (red, n = 101) exhibit higher heterogeneity than the non-tumour (green, n = 98) samples. The ellipse presents the 0.9 confidence intervals for each type. b, c, Heat map of NMF consensus matrix, cophenetic correlation coefficient and average silhouette-width plots. The NMF input is the quantile-normalized iBAQ intensity matrix of the top 25% most-variant proteins, across 101 tumour samples. On the basis of visual inspection of the hierarchical clustering and the profiles of cophenetic correlation coefficient and average silhouette width for solutions with 2 to 7 clusters, we consider K = 3 to be the preferred solution (indicated by black triangles) and use this to arrange the samples shown in Fig. 2a (yielding the three clusters highlighted in blue, yellow and red). Average cophenetic correlations of each rank of clusters are displayed under the heat map matrix. The green and blue points or lines in the cophenetic correlation and average silhouette-width plots represent the non-smooth NMF (nsNMF) and the Brunet algorithm, respectively.
Extended Data Fig. 6 Association of proteomic subtypes with clinical features and prognosis signature proteins of HBV-related early stage HCC.
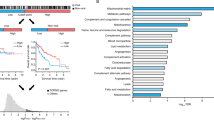
a, b, Association of MVI and serum AFP levels with proteomic subtypes. c, Association of age with proteomic subtypes. S-I (n = 36) HCC developed in older patients (62.0 ± 8.9 years), whereas S-II (n = 32) and S-III (n = 33) HCC generally developed in relatively young patients (54.5 ± 12.0 and 53.4 ± 10.1 years, respectively). The black lines denote the average age of the patients (mean ± s.d.) in the subtypes. d, e, Kaplan–Meier plots of HCC recurrence of subtypes identified in the proteomic (d) and Fudan (e) cohorts. The proteomic subtypes of patients from the Fudan cohort are classified by the nearest template prediction algorithm, with signature genes of proteomic subtypes (Methods). f, Prognosis signature proteins in proteomic subtypes. Heat map shows the relative abundance of each protein. Spearman’s correlation coefficients (cor.) between mRNA and protein expression levels and the log2(hazard ratio) of each protein are displayed on the right. Some of the prognosis signatures identified by proteomic analysis are highlighted by asterisks. For each subtype, the signature proteins are ordered by increasing hazard ratio (HR) values. P values are calculated by one-sided Fisher's exact test (a, b), two tailed Student’s t-test (c) and two-sided log-rank test (d, e).
Extended Data Fig. 7 Pathway alterations in the proteomic subtypes.
a, Heat map of alteration pathways in the proteomic subtypes. Left, alteration pathways identified from the proteome. Right, alteration pathways identified from the phospho-proteome. Colour of each cell represents the average ssGSEA enrichment scores of that subtype; red denotes activation and blue denotes inhibition. Blank cells represent non-enrichment. b, Details of alteration scores of integrin proteins, extracellular matrix proteins, metalloproteinase proteins and glycolysis proteins shown in Fig. 3a. c, Network diagram depicting the interaction of the Rho GTPase signalling pathway. The pink node represents proteins expressed at higher levels in the S-III tumours than in the S-I and S-II tumours; a red node border indicates the hyper-phosphorylation of that protein. d, Identification and quantification of hyper-phosphorylation sites for Rho GTPase activators (FGD3, DOCK2 and ARHGEF2) in S-I (n = 35), S-II (n = 30) and S-III (n = 29) tumours. In the box plots, the middle bar represents the median, and the box represents the interquartile range; bars extend to 1.5× the interquartile range. A grey circle under the box plot indicates that this phosphorylation site is not identified in these samples. P values are calculated by two-sided Wilcoxon rank-sum test.
Extended Data Fig. 8 Signature proteins in S-III tumours and the association of proteomic subtypes with genomic mutations and established transcriptomic signatures of HCC classifications.
a, Global heat map of the relative abundance of the signature proteins in the S-III tumours. Biological functions related to these proteins are denoted by the colour panel on the right, and annotated according to the pathways based on enrichment of MSigDB ontologies. b, Association of the proteomic subtypes with p53 and CTNNB1 mutations, and mRNA-based HCC subclasses. Samples with positive results are coloured in red, and the negative samples are shown in grey. Samples with no statistical significance, or which are not available, are coloured in white. The immunosuppression subclass is labelled by the dashed box. Prediction of mRNA-based HCC subclasses and immune-cell types was performed using nearest template prediction analysis, and ssGSEA and model-based clustering analysis, respectively (Methods). c, Immune infiltration in proteomic subtypes. Samples with high immune infiltration are coloured in red and samples with low immune infiltration are coloured in grey. P values are calculated by one-sided Fisher’s exact test.
Extended Data Fig. 9 Cholesterol metabolic dysregulation in HBV-related early-stage HCC.
a, Representative immunohistochemical staining of TMA with SOAT1 antibodies in an independent cohort of HCC (n = 254, biologically independent samples). Scale bars, 50 μm. b, Staining intensity of SOAT1 between non-tumour (N) and tumour (T) samples from TMA (n = 254). Data are mean ± s.d. c, Intensities of three cholesteryl esters in paired non-tumour and tumour HCC samples (n = 25) by electrospray ionization mass spectrometry. In the box plots, the middle bar represents the median, and the box represents the interquartile range; bars extend to 1.5× the interquartile range. d, e, Upregulation of the cholesterol-metabolism-related proteins (d) or their mRNAs (e) in paired non-tumour and tumour samples. The black lines in the middle of the plots indicate the mean of protein intensities (± s.e.m.) and mRNA abundance (± s.e.m.). The P values are calculated by two-sided Wilcoxon rank-sum test (b–e).
Extended Data Fig. 10 Downregulation of SOAT1 suppresses the proliferation and migration of HCC cells by reducing plasma-membrane cholesterol content, and then inhibiting the integrins and TGFβ signalling pathway.
a, Western blot of SOAT1 and SOAT2 in PLC/PRF/5 and HepG2 cell lines, with or without treatment with shRNA (n = 3 biologically independent experiments). For full scans, see Supplementary Fig. 1. b, c, SOAT1 protein abundance in PLC/PRF/5 and HepG2 cells based on proteomic data (b, mean ± s.e.m., n = 3 biological repeats) and western blotting analysis (c, n = 3 biologically independent experiments). For full scans, see Supplementary Fig. 1. d, Alteration pathways and proteins in the PLC/PRF/5 cell line after the SOAT1 knockdown or treatment with avasimibe (n = 3 biological repeats). e, Schematic of mechanism that underlies the dysregulation of integrins and TGFβRs by SOAT1 inhibition. f, Body weight of PDX models treated by control (100 μl PBS) or avasimibe (15 mg kg−1 day−1) on the indicated days (n = 36 mice per group). Mean ( ± s.d.) of body weight is plotted. g, Growth curves of PDX models treated by control (100 μl PBS) or avasimibe (15 mg kg−1 day−1) on the indicated days (n = 6 mice per group). Circle and triangle denotes the mean volume of tumours ( ± s.e.m.). The immunohistochemical staining of SOAT1 in PDX models is labelled on the top panel. h, Relative expression of S-III-signature pathway-related proteins among three SOAT1high (SOAT1-H) PDX and three SOAT1low (SOAT1-L) PDX models. Red and blue denote upregulation and downregulation, respectively. Zigzags on the y axis indicate places at which the y axis is truncated. The P values are calculated by two-tailed Student’s t-test. NS, not significant, *P < 0.05, ***P < 0.001.
Supplementary information
Supplementary Information
This file contains Supplementary Notes which shows the full Membership of the Chinese Human Proteome Project (CNHPP) Consortium.
Supplementary Figures
This file contains Supplementary Figure 1.
Supplementary Tables
This file contains Supplementary Tables 1-18.
Source data
Rights and permissions
About this article
Cite this article
Jiang, Y., Sun, A., Zhao, Y. et al. Proteomics identifies new therapeutic targets of early-stage hepatocellular carcinoma. Nature 567, 257–261 (2019). https://doi.org/10.1038/s41586-019-0987-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-0987-8
This article is cited by
-
Signature construction and molecular subtype identification based on liver-specific genes for prediction of prognosis, immune activity, and anti-cancer drug sensitivity in hepatocellular carcinoma
Cancer Cell International (2024)
-
Characterization of the proteome of stable and unstable carotid atherosclerotic plaques using data-independent acquisition mass spectrometry
Journal of Translational Medicine (2024)
-
Metabolic reprogramming in the tumor microenvironment of liver cancer
Journal of Hematology & Oncology (2024)
-
Comprehensive analysis of m6A modification in immune infiltration, metabolism and drug resistance in hepatocellular carcinoma
Cancer Cell International (2024)
-
Long-read transcriptome landscapes of primary and metastatic liver cancers at transcript resolution
Biomarker Research (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.