Abstract
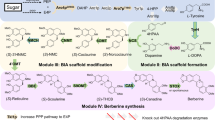
Cannabis sativa L. has been cultivated and used around the globe for its medicinal properties for millennia1. Some cannabinoids, the hallmark constituents of Cannabis, and their analogues have been investigated extensively for their potential medical applications2. Certain cannabinoid formulations have been approved as prescription drugs in several countries for the treatment of a range of human ailments3. However, the study and medicinal use of cannabinoids has been hampered by the legal scheduling of Cannabis, the low in planta abundances of nearly all of the dozens of known cannabinoids4, and their structural complexity, which limits bulk chemical synthesis. Here we report the complete biosynthesis of the major cannabinoids cannabigerolic acid, Δ9-tetrahydrocannabinolic acid, cannabidiolic acid, Δ9-tetrahydrocannabivarinic acid and cannabidivarinic acid in Saccharomyces cerevisiae, from the simple sugar galactose. To accomplish this, we engineered the native mevalonate pathway to provide a high flux of geranyl pyrophosphate and introduced a heterologous, multi-organism-derived hexanoyl-CoA biosynthetic pathway5. We also introduced the Cannabis genes that encode the enzymes involved in the biosynthesis of olivetolic acid6, as well as the gene for a previously undiscovered enzyme with geranylpyrophosphate:olivetolate geranyltransferase activity and the genes for corresponding cannabinoid synthases7,8. Furthermore, we established a biosynthetic approach that harnessed the promiscuity of several pathway genes to produce cannabinoid analogues. Feeding different fatty acids to our engineered strains yielded cannabinoid analogues with modifications in the part of the molecule that is known to alter receptor binding affinity and potency9. We also demonstrated that our biological system could be complemented by simple synthetic chemistry to further expand the accessible chemical space. Our work presents a platform for the production of natural and unnatural cannabinoids that will allow for more rigorous study of these compounds and could be used in the development of treatments for a variety of human health problems.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Nucleotide sequence data of Cannabis candidate prenyltransferases are available in the third-party annotation section of the DDBJ/ENA/GenBank databases (Extended Data Table 4). Strains and plasmids developed for this study (Extended Data Table 1), along with annotated sequences, have been deposited in the Synthetic Biology Engineering Research Center (Synberc) Registry (https://synberc-registry.jbei.org/) and are physically available from the authors upon reasonable request. Custom Python 3.6 scripts for data analysis are available from the authors upon reasonable request. Contractual obligations from commercial partnerships prohibit us from distributing (by ourselves or through a third party) strains described in our manuscript to for-profit commercial entities. However, we provide extensive genotypic descriptions of our strains, fully annotated DNA sequences, and detailed methods that enable others to build upon our work. Strains will be provided to nonprofit, government, or academic laboratories and institutions. Strains producing controlled substances or their direct precursors can only be provided to laboratories and institutions with appropriate approvals and licenses (for example, DEA permits).
Change history
17 March 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Pacher, P., Bátkai, S. & Kunos, G. The endocannabinoid system as an emerging target of pharmacotherapy. Pharmacol. Rev. 58, 389–462 (2006).
Whiting, P. F. et al. Cannabinoids for medical use: a systematic review and meta-analysis. J. Am. Med. Assoc. 313, 2456–2473 (2015).
Hazekamp, A., Ware, M. A., Muller-Vahl, K. R., Abrams, D. & Grotenhermen, F. The medicinal use of cannabis and cannabinoids—an international cross-sectional survey on administration forms. J. Psychoactive Drugs 45, 199–210 (2013).
ElSolhly, M. A. & Gul, W. in Handbook of Cannabis (ed. Pertwee, R. G.) Ch. 1 (Oxford Univ. Press, Oxford, 2014).
Dekishima, Y., Lan, E. I., Shen, C. R., Cho, K. M. & Liao, J. C. Extending carbon chain length of 1-butanol pathway for 1-hexanol synthesis from glucose by engineered Escherichia coli. J. Am. Chem. Soc. 133, 11399–11401 (2011).
Gagne, S. J. et al. Identification of olivetolic acid cyclase from Cannabis sativa reveals a unique catalytic route to plant polyketides. Proc. Natl Acad. Sci. USA 109, 12811–12816 (2012).
Zirpel, B., Stehle, F. & Kayser, O. Production of Δ9-tetrahydrocannabinolic acid from cannabigerolic acid by whole cells of Pichia (Komagataella) pastoris expressing Δ9-tetrahydrocannabinolic acid synthase from Cannabis sativa L. Biotechnol. Lett. 37, 1869–1875 (2015).
Taura, F. et al. Cannabidiolic-acid synthase, the chemotype-determining enzyme in the fiber-type Cannabis sativa. FEBS Lett. 581, 2929–2934 (2007).
Razdan, R. K. in The Cannabinoid Receptors (ed. Reggio, P. H.) 3–19 (Humana Press, New York, 2009).
Taura, F. et al. Characterization of olivetol synthase, a polyketide synthase putatively involved in cannabinoid biosynthetic pathway. FEBS Lett. 583, 2061–2066 (2009).
Saerens, S. M. G., Delvaux, F. R., Verstrepen, K. J. & Thevelein, J. M. Production and biological function of volatile esters in Saccharomyces cerevisiae. Microb. Biotechnol. 3, 165–177 (2010).
Stout, J. M., Boubakir, Z., Ambrose, S. J., Purves, R. W. & Page, J. E. The hexanoyl-CoA precursor for cannabinoid biosynthesis is formed by an acyl-activating enzyme in Cannabis sativa trichomes. Plant J. 71, 353–365 (2012).
Fellermeier, M. & Zenk, M. H. Prenylation of olivetolate by a hemp transferase yields cannabigerolic acid, the precursor of tetrahydrocannabinol. FEBS Lett. 427, 283–285 (1998).
Page, J. E. & Boubakir, Z. Aromatic prenyltransferase from Cannabis. US patent 2012/0144523 A1 (2012).
Reider Apel, A. et al. A Cas9-based toolkit to program gene expression in Saccharomyces cerevisiae. Nucleic Acids Res. 45, 496–508 (2017).
Ignea, C., Pontini, M., Maffei, M. E., Makris, A. M. & Kampranis, S. C. Engineering monoterpene production in yeast using a synthetic dominant negative geranyl diphosphate synthase. ACS Synth. Biol. 3, 298–306 (2014).
Kuzuyama, T., Noel, J. P. & Richard, S. B. Structural basis for the promiscuous biosynthetic prenylation of aromatic natural products. Nature 435, 983–987 (2005).
Zirpel, B., Degenhardt, F., Martin, C., Kayser, O. & Stehle, F. Engineering yeasts as platform organisms for cannabinoid biosynthesis. J. Biotechnol. 259, 204–212 (2017).
Li, H. et al. A heteromeric membrane-bound prenyltransferase complex from hop catalyzes three sequential aromatic prenylations in the bitter acid pathway. Plant Physiol. 167, 650–659 (2015).
van Bakel, H. et al. The draft genome and transcriptome of Cannabis sativa. Genome Biol. 12, R102 (2011).
Emanuelsson, O., Brunak, S., von Heijne, G. & Nielsen, H. Locating proteins in the cell using TargetP, SignalP and related tools. Nat. Protocols 2, 953–971 (2007).
Emanuelsson, O., Nielsen, H. & von Heijne, G. ChloroP, a neural network-based method for predicting chloroplast transit peptides and their cleavage sites. Protein Sci. 8, 978–984 (1999).
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001).
Taura, F., Morimoto, S., Shoyama, Y. & Mechoulam, R. First direct evidence for the mechanism of Δ1-tetrahydrocannabinolic acid biosynthesis. J. Am. Chem. Soc. 117, 9766–9767 (1995).
Taura, F., Morimoto, S. & Shoyama, Y. Purification and characterization of cannabidiolic-acid synthase from Cannabis sativa L. Biochemical analysis of a novel enzyme that catalyzes the oxidocyclization of cannabigerolic acid to cannabidiolic acid. J. Biol. Chem. 271, 17411–17416 (1996).
Sirikantaramas, S. et al. The gene controlling marijuana psychoactivity: molecular cloning and heterologous expression of Δ1-tetrahydrocannabinolic acid synthase from Cannabis sativa L. J. Biol. Chem. 279, 39767–39774 (2004).
Sirikantaramas, S. et al. Tetrahydrocannabinolic acid synthase, the enzyme controlling marijuana psychoactivity, is secreted into the storage cavity of the glandular trichomes. Plant Cell Physiol. 46, 1578–1582 (2005).
de Meijer, E. P. M. & Hammond, K. M. The inheritance of chemical phenotype in Cannabis sativa L. (V): regulation of the propyl-/pentyl cannabinoid ratio, completion of a genetic model. Euphytica 210, 291–307 (2016).
Wenig, P. & Odermatt, J. OpenChrom: a cross-platform open source software for the mass spectrometric analysis of chromatographic data. BMC Bioinformatics 11, 405 (2010).
McKinney, W. Data structures for statistical computing in Python. In Proc. 9th Python in Science Conference (eds van der Walt, S. & Millman, J.) 51–56 (SciPy, 2010).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Trifinopoulos, J., Nguyen, L.-T., von Haeseler, A. & Minh, B. Q. W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 44, W232–W235 (2016).
Huerta-Cepas, J., Serra, F. & Bork, P. ETE 3: reconstruction, analysis, and visualization of phylogenomic data. Mol. Biol. Evol. 33, 1635–1638 (2016).
Lee, T. S. et al. BglBrick vectors and datasheets: a synthetic biology platform for gene expression. J. Biol. Eng. 5, 12 (2011).
Acknowledgements
We thank G. Wang for the pESC-HlPT1L, pESC-HlPT2 and pESC-HlPT1L-HlPT2 plasmids; T. Laursen for help with microsomal preparations; C. Eiben, T. de Rond, R. Lee, C. Joshua, T. Tian, P. Shih, M. Wehrs and A. Zargar for discussion; L. Davis, T. Lease and B. Sandmann for help on controlled substance regulatory processes; E. Mendez and T. Dunn for strain archiving support; and O. Diaz and E. Coyne for administrative support. This work was supported by a National Science Foundation grant 1330914 to J.D.K; Y.Z. acknowledges support from a Jiangnan University Graduate Scholarship for Oversea Study.
Reviewer information
Nature thanks James Collins, Bernd Lange, Jonathan Mielenz and Jordan Zager for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
X.L., M.A.R., L.d’E., J.W., C.M.D., A.L. and J.D.K. conceived the study. X.L., M.A.R., L.d’E., J.W., A.L., C.M.D., Y.Z., A.T.G., S.H., W.L., H.L., C.Y., J.S. and I.D. constructed the plasmids and yeast strains and performed microbiological manipulations and extractions. X.L., M.A.R. and K.D. performed synthetic organic chemistry. X.L., M.A.R., V.T.B., G.W., E.E.K.B., Y.C. and C.J.P. performed mass spectrometry. M.A.R. performed enzyme kinetic study and in vitro studies. J.W. and L.d’E. identified and functionally expressed CsPT4. X.L., M.A.R., A.L. and J.W. performed bioinformatic analysis. M.A.R. developed data analysis scripts. All authors contributed to the manuscript.
Corresponding author
Ethics declarations
Competing interests
X.L., M.A.R., L.d’E., J.W., C.M.D., E.E.K.B. and C.J.P. have a financial interest in Demetrix. J.D.K. has a financial interest in Amyris, Lygos, Demetrix, Constructive Biology, Napigen, and Maple Bio.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Olivetolic acid production of yCAN03.
All LC–MS chromatograms were selected for the theoretical m/z values of the respective compounds of interest (Extended Data Table 3). a, yCAN03 produced olivetolic acid. An additional peak was observed for the byproduct HTAL (same m/z as olivetolic acid), which has previously been reported. No zymolyase was added during extraction. Chromatography gradient for cannabinoid analogues was used. b, Mass spectrum of HTAL. Mass accuracy for observed m/z at a given retention time (RT) is reported in parts per million (p.p.m.).
Extended Data Fig. 2 CsPT4-T characterization.
a, CsPT4-T localized to the purified microsomal fraction. All LC–MS chromatograms were selected for the theoretical m/z values of the respective compounds of interest (Extended Data Table 3). Chromatography gradient for cannabinoid analogues was used. Signals were compared to genuine standards. We incubated boiled microsomal fraction (mic ΔT), soluble fraction (sol) and microsomal fraction (mic) of yCAN14 in the presence of 500 µM olivetolic acid and 500 µM GPP for 1 h at 30 °C and observed GOT activity only in the microsomal fraction. b, CsPT4-T olivetolic acid kinetics. Using nonlinear regression to fit the Michaelis–Menten kinetic model for varied olivetolic acid (0.25 µM–0.56 mM) and constant GPP (1.67 mM) concentrations revealed a Km(olivetolic acid) = 6.73 ± 0.26 µM (mean ± s.d.) (n = 4 technically independent samples; measurements were plotted individually). c, CsPT4-T GPP kinetics. Eadie–Hofstee linearization of Michaelis–Menten model showed non-Michaelis–Menten type behaviour for CsPT4-T for varied GPP (0.25 µM–1.67 mM) and constant olivetolic acid (1.67 mM) concentrations. Measurements do not fall on a line as would be expected (R2, coefficient of determination; n = 3 technically independent samples; measurements were plotted individually). d, Phylogenetic tree of Cannabis and related prenyltransferases. The numbers indicate the bootstrap value (%) from 1,000 replications. Grey dashed lines depict branch offsets to accommodate labels. The scale bar shows the amino acid substitution ratio. Prenyltransferases cluster by biosynthetic pathway. CsPT4 catalyses CBGA production. The function of the other Cannabis prenyltransferases remains unknown.
Extended Data Fig. 3 In vitro activity of NphB and HlPT.
All LC–MS chromatograms were selected for the theoretical m/z values of the respective compounds of interest (Extended Data Table 3). Chromatography gradient for cannabinoid analogues was used. Signals were compared to genuine standards. a, Purified NphB catalysed the condensation of GPP and olivetolic acid to CBGA when incubated with 5 mM olivetolic acid and 5 mM GPP at room temperature for 24 h. The enzyme produced at least one other isomer of CBGA, which is consistent with previous reports18. Boiling NphB (NphB ΔT) abolished activity. b, Microsomal fractions were prepared from yCAN21, yCAN22, and yCAN23 that expressed HlPT1L-T (1L-T), HlPT1L-T and HlPT2-T (1L-T/2-T), and HlPT2-T (2-T), respectively. Incubation with 5 mM olivetolic acid and 5 mM GPP for 24 h at 30 °C yielded CBGA as well as several isomers. CBGA production was not observed when incubating HlPT1L-T and HlPT2-T with their native substrates phlorisovalerophenone and dimethylallyl diphosphate (neg).
Extended Data Fig. 4 Divarinolic acid and CBGVA production.
All LC–MS chromatograms were selected for the theoretical m/z values of the respective compounds of interest (Extended Data Table 3). a, Compared with a strain that lacks the olivetolic acid synthesis pathway (yCAN14), yCAN32 produced an additional compound with an m/z value corresponding to divarinolic acid (DA). The retention time of divarinolic acid is reduced relative to olivetolic acid, owing to the reduced hydrophobicity that is conferred by the shorter 5-propyl side chain. No zymolyase was added during extraction. Chromatography gradient for cannabinoid analogues was used. b, Mass spectrum of divarinolic acid. Mass accuracy for observed m/z at a given retention time is reported in parts per million. c, As expected, the prenylated variant of divarinolic acid, CBGVA, was also detected. d, Mass spectrum of CBGVA, with mass accuracy reported in parts per million.
Extended Data Fig. 5 Production of cannabinoid analogues.
a, Pathway assayed for incorporation of different fatty acid precursors into the corresponding olivetolic acid and CBGA analogues. b, Ion-extracted chromatograms of the engineered strain (yCAN31) grown in the presence of butanoic acid (VII), nonanoic acid (VIII), 5-phenylpentanoic acid (IX), 6-phenylhexanoic acid (X) and 7-phenylheptanoic acid (XI); peaks corresponding to their respective olivetolic acid and HTAL analogues were detected. Detected analogue peaks shifted in retention time depending on their change in hydrophobicity relative to olivetolic acid and CBGA standards. Feeding 1 mM hexanoic acid to yCAN30 yielded neither olivetolic acid nor CBGA (neg (I)). Supplementation of the medium with butanoic acid did not increase production of divarinolic acid or CBGVA over baseline levels. Feeding phenyl-substituted acids yielded only either the corresponding HTAL analogues or olivetolic acid analogues. All LC–MS chromatograms were selected for the theoretical m/z values of the respective compounds of interest (Extended Data Table 3).
Extended Data Fig. 6 LC–MS spectra of cannabinoid analogues.
Mass accuracies for observed m/z values at a given retention time are reported in parts per million. a, Mass spectra of cannabinoid analogues produced from fatty acids (I) to (VI). b, Mass spectra of 6hCBGA and 6hTHCA.
Supplementary information
Supplementary Information
This file contains the Supplementary Methods.
Rights and permissions
About this article
Cite this article
Luo, X., Reiter, M.A., d’Espaux, L. et al. Complete biosynthesis of cannabinoids and their unnatural analogues in yeast. Nature 567, 123–126 (2019). https://doi.org/10.1038/s41586-019-0978-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-0978-9
This article is cited by
-
Engineered yeast brews plant hormones
Nature Synthesis (2024)
-
The forefront of chemical engineering research
Nature Chemical Engineering (2024)
-
De novo biosynthesis of the hops bioactive flavonoid xanthohumol in yeast
Nature Communications (2024)
-
Reconstitution of early paclitaxel biosynthetic network
Nature Communications (2024)
-
The NLRP3 inflammasome: a vital player in inflammation and mediating the anti-inflammatory effect of CBD
Inflammation Research (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.