Abstract
The ability of cytoskeletal motors to move unidirectionally along filamentous tracks is central to their role in cargo transport, motility and cell division. Kinesin and myosin motor families have a subclass that moves towards the opposite end of the microtubule or actin filament with respect to the rest of the motor family1,2, whereas all dynein motors that have been studied so far exclusively move towards the minus end of the microtubule3. Guided by cryo-electron microscopy and molecular dynamics simulations, we sought to understand the mechanism that underpins the directionality of dynein by engineering a Saccharomyces cerevisiae dynein that is directed towards the plus end of the microtubule. Here, using single-molecule assays, we show that elongation or shortening of the coiled-coil stalk that connects the motor to the microtubule controls the helical directionality of dynein around microtubules. By changing the length and angle of the stalk, we successfully reversed the motility towards the plus end of the microtubule. These modifications act by altering the direction in which the dynein linker swings relative to the microtubule, rather than by reversing the asymmetric unbinding of the motor from the microtubule. Because the length and angle of the dynein stalk are fully conserved among species, our findings provide an explanation for why all dyneins move towards the minus end of the microtubule.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The generated yeast strains and the data that support the findings of this study are available from the corresponding author upon request.
References
Bryant, Z., Altman, D. & Spudich, J. A. The power stroke of myosin VI and the basis of reverse directionality. Proc. Natl Acad. Sci. USA 104, 772–777 (2007).
Sablin, E. P. et al. Direction determination in the minus-end-directed kinesin motor ncd. Nature 395, 813–816 (1998).
Roberts, A. J., Kon, T., Knight, P. J., Sutoh, K. & Burgess, S. A. Functions and mechanics of dynein motor proteins. Nat. Rev. Mol. Cell Biol. 14, 713–726 (2013).
Reck-Peterson, S. L. et al. Single-molecule analysis of dynein processivity and stepping behavior. Cell 126, 335–348 (2006).
Gennerich, A., Carter, A. P., Reck-Peterson, S. L. & Vale, R. D. Force-induced bidirectional stepping of cytoplasmic dynein. Cell 131, 952–965 (2007).
King, S. M. (ed.) Dyneins, Dynein Mechanics, Dysfunction and Disease Vol. 2, 2nd edn (Academic, London, 2017).
Gee, M. A., Heuser, J. E. & Vallee, R. B. An extended microtubule-binding structure within the dynein motor domain. Nature 390, 636–639 (1997).
Roberts, A. J. et al. AAA+ ring and linker swing mechanism in the dynein motor. Cell 136, 485–495 (2009).
Schmidt, H., Gleave, E.S. & Carter, A.P. Insights into dynein motor domain function from a 3.3-Å crystal structure. Nat. Struct. Mol. Biol. 19, 492–497 (2012).
Kon, T. et al. The 2.8 Å crystal structure of the dynein motor domain. Nature 484, 345–350 (2012).
Schmidt, H., Zalyte, R., Urnavicius, L. & Carter, A. P. Structure of human cytoplasmic dynein-2 primed for its power stroke. Nature 518, 435–438 (2015).
Carter, A. P. et al. Structure and functional role of dynein’s microtubule-binding domain. Science 322, 1691–1695 (2008).
Lin, J., Okada, K., Raytchev, M., Smith, M. C. & Nicastro, D. Structural mechanism of the dynein power stroke. Nat. Cell Biol. 16, 479–485 (2014).
Imai, H. et al. Direct observation shows superposition and large scale flexibility within cytoplasmic dynein motors moving along microtubules. Nat. Commun. 6, 8179 (2015).
Lippert, L. G. et al. Angular measurements of the dynein ring reveal a stepping mechanism dependent on a flexible stalk. Proc. Natl Acad. Sci. USA 114, E4564–E4573 (2017).
Cleary, F. B. et al. Tension on the linker gates the ATP-dependent release of dynein from microtubules. Nat. Commun. 5, 4587 (2014).
Nicholas, M. P. et al. Cytoplasmic dynein regulates its attachment to microtubules via nucleotide state-switched mechanosensing at multiple AAA domains. Proc. Natl Acad. Sci. USA 112, 6371–6376 (2015).
Furuta, A. et al. Creating biomolecular motors based on dynein and actin-binding proteins. Nat. Nanotechnol. 12, 233–237 (2017).
Can, S., Dewitt, M. A. & Yildiz, A. Bidirectional helical motility of cytoplasmic dynein around microtubules. eLife 3, e03205 (2014).
DeWitt, M. A., Chang, A. Y., Combs, P. A. & Yildiz, A. Cytoplasmic dynein moves through uncoordinated stepping of the AAA+ ring domains. Science 335, 221–225 (2012).
Qiu, W. et al. Dynein achieves processive motion using both stochastic and coordinated stepping. Nat. Struct. Mol. Biol. 19, 193–200 (2012).
Redwine, W. B. et al. Structural basis for microtubule binding and release by dynein. Science 337, 1532–1536 (2012).
Gibbons, I. R. et al. The affinity of the dynein microtubule-binding domain is modulated by the conformation of its coiled-coil stalk. J. Biol. Chem. 280, 23960–23965 (2005).
Kon, T. et al. Helix sliding in the stalk coiled coil of dynein couples ATPase and microtubule binding. Nat. Struct. Mol. Biol. 16, 325–333 (2009).
Belyy, V., Hendel, N. L., Chien, A. & Yildiz, A. Cytoplasmic dynein transports cargos via load-sharing between the heads. Nat. Commun. 5, 5544 (2014).
Niekamp, S., Coudray, N., Zhang, N., Vale, R. D. & Bhabha, G. Stalk-mediated communication in the dynein motor domain. Preprint at https://www.biorxiv.org/content/early/2018/04/27/309179 (2018).
Chrétien, D., Kenney, J. M., Fuller, S. D. & Wade, R. H. Determination of microtubule polarity by cryo-electron microscopy. Structure 4, 1031–1040 (1996).
Gell, C. et al. Purification of tubulin from porcine brain. Methods Mol. Biol. 777, 15–28 (2011).
Yildiz, A. et al. Myosin V walks hand-over-hand: single fluorophore imaging with 1.5-nm localization. Science 300, 2061–2065 (2003).
Neuman, K. C. & Block, S. M. Optical trapping. Rev. Sci. Instrum. 75, 2787–2809 (2004).
Dogan, M. Y., Can, S., Cleary, F. B., Purde, V. & Yildiz, A. Kinesin’s front head is gated by the backward orientation of its neck linker. Cell Rep. 10, 1967–1973 (2015).
Schlitter, J., Engels, M. & Krüger, P. Targeted molecular dynamics: a new approach for searching pathways of conformational transitions. J. Mol. Graph. 12, 84–89 (1994).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Best, R. B. et al. Optimization of the additive CHARMM all-atom protein force field targeting improved sampling of the backbone φ, ψ and side-chain χ1 and χ2 dihedral angles. J. Chem. Theory Comput. 8, 3257–3273 (2012).
Kellogg, E. H. et al. Insights into the distinct mechanisms of action of taxane and non-taxane microtubule stabilizers from cryo-EM structures. J. Mol. Biol. 429, 633–646 (2017).
Alushin, G. M. et al. High-resolution microtubule structures reveal the structural transitions in αβ-tubulin upon GTP hydrolysis. Cell 157, 1117–1129 (2014).
Acknowledgements
We thank F. B. Cleary, H. Schmidt, C. Cypranowska, V. Belyy and L. Ferro for initiation of this project; S. Chen, C. Savva, G. Cannone, J. Grimmett, T. Darling and L. Kellogg for technical assistance with cryo-EM; M. Mofrad and M. Mehrbod for coarse-grained simulations; and the Savio Cluster for molecular dynamics simulations. This work was funded by grants from the NIH (GM094522), and NSF (MCB-1055017, MCB-1617028) to A.Y., the Wellcome Trust (WT100387) and the Medical Research Council, UK (MC_UP_A025_1011) to A.P.C., and TUBITAK (215Z398) and ITU BAP (38777) to M.G.
Reviewer information
Nature thanks W. Hancock and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
A.Y. and A.P.C. initiated the project. S.C. engineered dynein mutants and performed motility assays. S.L. performed cryo-EM imaging. M.G. performed molecular dynamics simulations. A.Y., S.C., A.P.C. and M.G. prepared the manuscript. A.Y., A.P.C. and M.G. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Mechanochemical cycle of dynein.
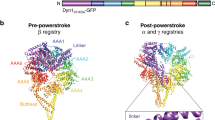
a, The AAA+ ring of a dynein monomer lies parallel to the microtubule, and the stalk is tilted towards the plus end at its base. In the absence of a nucleotide (apo) at AAA1, dynein is tightly bound to microtubules and the linker has a straight post-power stroke conformation, exiting the ring at AAA4. ATP binding to AAA1 (yellow) triggers microtubule release through a shift in the registry of a coiled-coil stalk, and the linker undergoes the priming stroke. At this pre-power stroke conformation, the linker is bent by a flexible hinge towards the middle of the ring and exits the ring at AAA2. The LSV is aligned with the long axis of the microtubule, and moves the MTBD towards the minus end. After ATP hydrolysis, the dynein head re-binds to the microtubule, and releases the inorganic phosphate (Pi). In the ADP-bound state, the linker undergoes a force-generating power stroke by moving back to its straight conformation. This pulls the cargo towards the minus end (black arrow). After ADP release, dynein returns back to the apo state for the next cycle.
Extended Data Fig. 2 Engineering the directionality of dynein motility.
Schematic of the helices (CC1 and CC2) at the stalk of yeast cytoplasmic dynein shows the heptad repeat hydrophobic contacts (black lines) in the core of the coiled-coil, when dynein is in a low-microtubule-affinity (β) state. Conserved proline residues at the base of the Dyn stalk are highlighted by magenta arrows. The three heptads deleted from the stalk of Dyn−3hep are highlighted in green in Dyn. The three- and seven heptad repeats inserted into Dyn+3hep, Dyn+7hep and DynRK+7hep are highlighted in red. The inserted sequences were taken from the Drosophila melanogaster cytoplasmic dynein12. Point mutations inserted into DynRK and DynRK+7hep are highlighted in cyan.
Extended Data Fig. 3 Estimated orientation of the LSV relative to a microtubule as a function of stalk length.
a, A Dyn monomer was manually docked onto a tubulin dimer (PDB 3VKG10, 4RH711, 3J6G36 and 5SYF35). The LSV was defined as the displacement vector of the N terminus of the linker from pre- (V1258 of PDB 4RH7, red bead) to post-power stroke (A1526 of PDB 3VKG, green bead) conformation. The stalk axis was defined as the vector that connects S3100 to S3248 (PDB 4RH7), which lies in the same plane with LSV of Dyn. b, Definition of the principal axes. Dyn was manually docked onto a microtubule. The longitudinal axis (PA1) is directed towards the minus end of the long axis of the microtubule. The radial axis (PA2) is directed from the microtubule centre-of-mass towards the pre-power stroke conformation of the linker (V1258 of PDB 4RH7, red bead). The tangential axis (PA3) is perpendicular to PA1 and PA2, as shown. c, The LSV (black arrow) of Dyn is aligned with the microtubule axis and is parallel to PA1. The expected orientations of Dyn+3hep and Dyn−3hep were modelled by alignment of the coiled-coils after insertions and deletions into the stalk. Insertion of three heptads into the stalk (Dyn+3hep) is expected to reorient the ring and rotate the LSVshort clockwise with respect to the minus end of the microtubule. Shortening the stalk by three heptads (Dyn−3hep) is expected to rotate the LSVshort anticlockwise. d, Velocity analysis of dynein-driven beads around microtubule bridges. All of the beads moved towards the minus end of the microtubule. n = 24, 20, 19 and 22 beads, from left to right. Centre line and error bars represent the mean and 5–95% confidence intervals. e, The comparison of the average LSV angles from molecular dynamics simulations (n = 1,680 conformations from 3 different simulations) and the average pitch angles from helical rotation of dynein-driven beads (nbeads = 24, 20 and 19, and nrotations = 99, 59 and 72 for Dyn, Dyn+3hep, and Dyn−3hep, respectively) reveals that the LSVshort determines the helical directionality of dynein. Error bars represent s.d. In d and e, P values are calculated from a two-sided t-test.
Extended Data Fig. 4 Alignment of the CC1 of the stalk region in 67 dynein heavy chains.
The sequences are oriented from the N terminus to the C terminus in these alignments. Isoforms of dynein used in the alignment were grouped on the basis of the type and the organism (cytoplasmic (cyt1), axonemal outer arm (22Sab, 22Sg), axonemal inner arm (IA) and intra-flagellar transport (cyt2) dyneins). The α and β registry of the stalk coiled-coils is shown on top. Stalk length is conserved among dyneins. Fully conserved proline residues at the base of the MTBD that cause tilting of the stalk coiled-coils towards the plus end of the microtubule are highlighted in green. Other residues that are conserved at over 90% are highlighted in yellow.
Extended Data Fig. 5 Alignment of the CC2 of the stalk region in 67 dynein heavy-chains.
The sequences are oriented from the N terminus to the C terminus in these alignments. Isoforms of dynein used in the alignment were grouped on the basis of the type and the organism (cytoplasmic (cyt1), axonemal outer arm (22Sab, 22Sg), axonemal inner arm (IA) and intra-flagellar transport (cyt2) dyneins). The α and β registry of the stalk coiled-coils is shown on top. Stalk length is conserved among dyneins. Fully conserved proline residues at the base of the MTBD that cause tilting of the stalk coiled-coils towards the plus end of the microtubule are highlighted in green. Other residues that are conserved at over 90% are highlighted in yellow.
Extended Data Fig. 6 Calculation of stalk and LSV angles by molecular dynamics simulations.
a, Changes of the stalk angle in three independent molecular dynamics simulations of Dyn and DynRK, and one simulation of DynRK+7hep. In the DynRK1 and DynRK3 simulations, the stalk angle sharply increases around 50 ns and 400 ns, respectively and remains pointed towards the minus end after its reversal; 180° represents the tilting of the stalk towards the minus end. b, Left, stalk angle distribution for the DynRK simulations; 180° represents the tilting of the stalk towards the minus end of the microtubule. Right, the length of the LSV unit vector projected onto the long axis of the microtubule after the reversal of the stalk in DynRK1; −1 corresponds to LSV pointed towards the minus end. c, Stalk length distributions from molecular dynamics simulations and cryo-EM experiments (mean ± s.d., n = 2,400, 2,400, 7,263, 7,263, 392 and 421 conformations, from left to right). Centre line and error bars represents the mean and 5–95% confidence intervals. P values are calculated from a two-sided t-test.
Extended Data Fig. 7 Cryo-EM image analysis of dynein monomers on microtubules.
a, A representative cryo-electron micrograph of Dyn monomers bound to microtubules with single monomers indicated with arrows, and enlarged insets. Scale bar, 100 nm. n = 98 micrographs from 1 grid for Dyn, and 235 micrographs from 2 grids for DynRK+7hep. b, A representative cryo-electron micrograph of DynRK+7hep monomers bound to microtubules with single monomers indicated with arrows, and enlarged insets. Scale bar, 100 nm. c, A simplified depiction of the power spectrum of a microtubule as a method to determine microtubule polarity. Microtubules with different numbers of protofilaments have different degrees of protofilament skew around the long axis. This causes differences in the Moiré patterns produced in cryo-EM images. These visual changes result in changes to the position of certain reflections in the microtubule power spectrum. As a result, right- and left-handed helix architectures can be differentiated by the relative positions of the JS (light green) and JN–S (dark green) reflections. For right-handed helices, the JN–S (RH, pink) reflection is further from the equator (blue) than the JS reflection, whereas for left-handed helices the JN–S (LH, purple) reflection is closer to the equator than the JS reflection. When the microtubule is Fourier-filtered to only include information from the equator (blue), a characteristic arrowhead pattern is formed from the Moiré patterns. For right-handed helix architectures, this points towards the plus end; for left-handed helix architectures, this points towards the minus end. Thirteen protofilament microtubules were not included in the analysis because they have no protofilament skew. d, An example power spectrum of an microtubule determined to be a right-handed helix. The enlargement on the right shows that the JS reflection is closer to the equator than the JN–S reflection. e, A Fourier-filtered image produced from the equatorial reflections from the power spectrum in d (blue box). For right-handed helix architectures, the arrowheads point towards the plus end. Scale bar, 100 nm. f, An example power spectrum of an microtubule determined to be a left-handed helix. The enlargement on the right shows that the JS reflection is further from the equator than the JN–S reflection. g, A Fourier-filtered image produced from the equatorial reflections from the power spectrum in f (blue box). For left-handed helix architectures, the arrowheads point towards the minus end. Scale bar, 100 nm. h, Orthogonal views of an atomic model of a dynein motor in an arrangement corresponding to the wild-type view, and synthetic projections produced from them. In this arrangement, the motor appears as an even ring. i, As in h, but with the model tilted 30° around the indicated axis. In this arrangement, the projection creates a crescent shape similar to that seen in the DynRK+7hep class (Fig. 2e), albeit with the ring and stalk unflipped; this suggests that the ring of DynRK+7hep is slightly tilted in relation to the microtubule.
Extended Data Fig. 8 Nucleotide- and force-induced release of Dyn and DynRK+7hep monomers from microtubules.
a, The normalized intensity of 100 nM GFP-tagged Dyn and DynRK+7hep monomers on sea urchin axonemes under given nucleotide conditions. Similar to Dyn, DynRK+7hep released from microtubules in the ADP–Pi state, mimicked by ATP and vanadate (Vi, n = 40 axonemes from three independent measurements, mean ± s.d.). P values are calculated from a two-sided t-test. b, A model of the dynein–microtubule interaction shows two distinct binding modes in the apo state, with k1 and k2 representing force-induced release rates from the weak and strong states, respectively. The slow rate (k2) represents strong binding of the motor to its tubulin binding site, whereas the fast rate (k1) represents transient or nonspecific interactions of the motor with the microtubule. c, Cumulative probability distributions (solid circles) of the microtubule-bound time of Dyn monomers at given force ranges. The release rates (k1 and k2) were calculated by a two-exponential-decay fit (solid curves). d, Calculated k1 values from the exponential fit (± 95% confidence intervals) to the DynRK+7hep dwell time has similar force-dependence to Dyn. Each bin contains 120 dwells from 2 independent measurements.
Extended Data Fig. 9 DynRK+7hep monomers exhibit robust plus-end directionality in microtubule gliding and bead-motility assays.
a, Top, schematic of the microtubule gliding assay with monomeric dynein. Bottom, images from time-lapse recordings show gliding of polarity-marked microtubules by Dyn and DynRK+7hep monomers. Dyn glides microtubules with their plus end ahead, whereas DynRK+7hep glides microtubules towards the opposite direction. n = 3 biological replicates. b, Microtubule gliding velocity and directionality of Dyn and DynRK+7hep in the presence and absence of 100 mM KCl. Negative velocities correspond to minus-end directionality. n = 45, 47, 27 and 70 from left to right, from two independent measurements. c, Schematic of the bead-motility assay with monomeric dynein (not to scale). N-terminally GFP-tagged monomers are attached to 860-nm diameter GFP-antibody-coated beads from their tail. d, Velocities of the beads driven by Dyn and DynRK+7hep monomers. n = 29 and 24 from left to right, from three independent experiments. In b and d, the centre line and edges represent the mean and 5–95% confidence intervals, respectively. P values are calculated from a two-sided t-test.
Supplementary information
Video 1: Helical movement of cargo beads carried by dynein.
Example recordings of 500 nm diameter cargo beads driven by Dyn, Dyn+3hep and Dyn-3hep on MT bridges on 2 µm diameter beads coated with SRS-MTBD85:82. The movies were acquired at 10 Hz under bright-field illumination. A fluorescent image of Cy3-labeled MTs (arrow) has been superimposed in the first frame. White arrowhead indicates the monodisperse cargo beads that walk on the bridges. The xy positions of the beads were determined by Gaussian fitting and the z position was determined from their intensities (from left to right, nbeads = 24, 20, 19 and nrotations = 99, 59, 72 from two independent experiments).
Video 2: MD simulations show the reversal of dynein’s stalk by altering the position of the proline residues at its base.
MD simulations of Dyn1 and DynRK-1 (blue and cyan, respectively) are shown over 72 ns. The pre-powerstroke conformation of dynein was selected for the starting point of the simulations. 0 ns represents the minimized and equilibrated structure. Reverse tilt transformation of DynRK takes place during time interval 20-50 ns. Conformers were sampled each 0.1 ns. Each conformer was docked on the MT after the MD simulations. Trajectories were smoothed over a window size of 5. Water and ions were not shown for clarity (n = 3 independent simulations for Dyn and DynRK).
Video 3: Variation in stalk angle from CryoEM images.
For both Dyn and DynRK+7hep, particles were picked and aligned to each other in Relion. Particles were sorted with ascending stalk angles and averaged as sets of 15 frames to increase contrast. Particles have a broad distribution of stalk angles, centered on a pivot at the base of the stalk. The striking difference is the complete reversal of stalk angle of DynRK+7hep compared to Dyn, such that most particles point in the opposite direction. Scale bar corresponds to 20 nm (n = 392 particles from 1 grid in Dyn and 421 particles from 2 grids in DynRK+7hep).
Video 4: Example recordings of MT gliding assays at 0 mM KCl.
GFP-tagged motors were immobilized to the glass surface from their tails via the GFP-antibody. The minus-end of MTs was polarity marked with bright Cy5-labeled NEM tubulin seeds. The assays were performed at 1 mM ATP without additional salt (0 mM KCl). The movies were recorded at 1 Hz under TIRF illumination. Only MTs with a single bright end were analyzed. Dyn glides MTs with their dark (plus) ends in the lead, whereas DynRK+7hep glides MTs with their bright (minus) ends in the lead (n = 38, 49, 59, 49 MTs for Dyn, Dyn+7hep, DynRK and DynRK+7hep, respectively).
Video 5: Example recordings of MT gliding assays with DynRK+7hep at various salt concentrations.
GFP-tagged motors were immobilized to the glass surface from their tails via the GFP-antibody. The assays were performed at 1 mM ATP with 0, 100 and 175 mM additional KCl in the assay buffer. The movies were recorded at 1 Hz under TIRF illumination. The minus end of MTs was polarity marked with bright Cy5-labeled NEM tubulin seeds. Only MTs with a single bright end were analyzed (n = 49, 83 and 53 MTs from left to right).
Video 6: Example recordings of single-molecule motility assays.
Full-length GFP-DynRK+7hep was labeled at its N-terminus with anti-GFP coated 655-nm QDs. At the beginning of the movie, Cy3-labeled polarity marked MTs were imaged under 532 nm excitation to determine the MT polarity. Later, excitation was switched to 488 nm for QD imaging at 1 Hz under TIRF illumination. GFP and QD fluorescence were superimposed after data collection (n = 52 motors).
Video 7: Example recordings of bead motility driven by monomeric dyneins.
500 nm diameter cargo beads were driven by multiple GFP-Dyn or GFP-DynRK+7hep monomers on surface immobilized axonemes. Motors are attached to GFP-antibody coated beads through the GFP tag at their N-termini. MT polarity was determined from the directionality of single TMR-labeled native dynein dimers (GST-Dyn-TMR) added to the assay chamber. GFP (green) and TMR (red) signals were simultaneously recorded at 1 Hz under TIRF illumination. White and yellow arrows indicate the motility of beads and TMR-dynein dimers, respectively, on the same MT (n = 29 and 24 from top to bottom from three independent experiments).
Rights and permissions
About this article
Cite this article
Can, S., Lacey, S., Gur, M. et al. Directionality of dynein is controlled by the angle and length of its stalk. Nature 566, 407–410 (2019). https://doi.org/10.1038/s41586-019-0914-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-0914-z
This article is cited by
-
Lis1 slows force-induced detachment of cytoplasmic dynein from microtubules
Nature Chemical Biology (2024)
-
Membrane-bound myosin IC drives the chiral rotation of the gliding actin filament around its longitudinal axis
Scientific Reports (2023)
-
Regulatory mechanisms of the dynein-2 motility by post-translational modification revealed by MD simulation
Scientific Reports (2023)
-
Tau Toxicity in Neurodegeneration
Molecular Neurobiology (2022)
-
In situ ultrastructures of two evolutionarily distant apicomplexan rhoptry secretion systems
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.