Abstract
Environmental genotoxic factors pose a challenge to the genomic integrity of epithelial cells at barrier surfaces that separate host organisms from the environment. They can induce mutations that, if they occur in epithelial stem cells, contribute to malignant transformation and cancer development1,2,3. Genome integrity in epithelial stem cells is maintained by an evolutionarily conserved cellular response pathway, the DNA damage response (DDR). The DDR culminates in either transient cell-cycle arrest and DNA repair or elimination of damaged cells by apoptosis4,5. Here we show that the cytokine interleukin-22 (IL-22), produced by group 3 innate lymphoid cells (ILC3) and γδ T cells, is an important regulator of the DDR machinery in intestinal epithelial stem cells. Using a new mouse model that enables sporadic inactivation of the IL-22 receptor in colon epithelial stem cells, we demonstrate that IL-22 is required for effective initiation of the DDR following DNA damage. Stem cells deprived of IL-22 signals and exposed to carcinogens escaped DDR-controlled apoptosis, contained more mutations and were more likely to give rise to colon cancer. We identified metabolites of glucosinolates, a group of phytochemicals contained in cruciferous vegetables, to be a widespread source of genotoxic stress in intestinal epithelial cells. These metabolites are ligands of the aryl hydrocarbon receptor (AhR)6, and AhR-mediated signalling in ILC3 and γδ T cells controlled their production of IL-22. Mice fed with diets depleted of glucosinolates produced only very low levels of IL-22 and, consequently, the DDR in epithelial cells of mice on a glucosinolate-free diet was impaired. This work identifies a homeostatic network protecting stem cells against challenge to their genome integrity by AhR-mediated ‘sensing’ of genotoxic compounds from the diet. AhR signalling, in turn, ensures on-demand production of IL-22 by innate lymphocytes directly regulating components of the DDR in epithelial stem cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
RNA-seq data are available at ArrayExpress (http://www.ebi.ac.uk/arrayexpress/) under accession number E-MTAB-6723. All datasets generated and/or analysed during the current study are presented in this published article, the accompanying Source Data or Supplementary Information files, or are available from the corresponding author upon reasonable request.
References
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Wu, S., Powers, S., Zhu, W. & Hannun, Y. A. Substantial contribution of extrinsic risk factors to cancer development. Nature 529, 43–47 (2016).
Blokzijl, F. et al. Tissue-specific mutation accumulation in human adult stem cells during life. Nature 538, 260–264 (2016).
Blanpain, C., Mohrin, M., Sotiropoulou, P. A. & Passegué, E. DNA-damage response in tissue-specific and cancer stem cells. Cell Stem Cell 8, 16–29 (2011).
Halazonetis, T. D., Gorgoulis, V. G. & Bartek, J. An oncogene-induced DNA damage model for cancer development. Science 319, 1352–1355 (2008).
Bjeldanes, L. F., Kim, J. Y., Grose, K. R., Bartholomew, J. C. & Bradfield, C. A. Aromatic hydrocarbon responsiveness-receptor agonists generated from indole-3-carbinol in vitro and in vivo: comparisons with 2,3,7,8-tetrachlorodibenzo-p-dioxin. Proc. Natl Acad. Sci. USA 88, 9543–9547 (1991).
Fu, D., Calvo, J. A. & Samson, L. D. Balancing repair and tolerance of DNA damage caused by alkylating agents. Nat. Rev. Cancer 12, 104–120 (2012).
Greten, F. R. et al. IKKβ links inflammation and tumorigenesis in a mouse model of colitis-associated cancer. Cell 118, 285–296 (2004).
Martincorena, I. et al. Universal patterns of selection in cancer and somatic tissues. Cell 171, 1029–1041 (2017).
Bailey, M. H. et al. Comprehensive characterization of cancer driver genes and mutations. Cell 173, 371–385 (2018).
Huber, S. et al. IL-22BP is regulated by the inflammasome and modulates tumorigenesis in the intestine. Nature 491, 259–263 (2012).
Lindemans, C. A. et al. Interleukin-22 promotes intestinal-stem-cell-mediated epithelial regeneration. Nature 528, 560–564 (2015).
Zenewicz, L. A. et al. IL-22 deficiency alters colonic microbiota to be transmissible and colitogenic. J. Immunol. 190, 5306–5312 (2013).
Barker, N. et al. Crypt stem cells as the cells-of-origin of intestinal cancer. Nature 457, 608–611 (2009).
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Snippert, H. J. et al. Intestinal crypt homeostasis results from neutral competition between symmetrically dividing Lgr5 stem cells. Cell 143, 134–144 (2010).
Sanos, S. L. et al. RORγt and commensal microflora are required for the differentiation of mucosal interleukin 22-producing NKp46+ cells. Nat. Immunol. 10, 83–91 (2009).
Falck, J., Coates, J. & Jackson, S. P. Conserved modes of recruitment of ATM, ATR and DNA-PKcs to sites of DNA damage. Nature 434, 605–611 (2005).
Barry, S. P. et al. STAT3 modulates the DNA damage response pathway. Int. J. Exp. Pathol. 91, 506–514 (2010).
Pickert, G. et al. STAT3 links IL-22 signaling in intestinal epithelial cells to mucosal wound healing. J. Exp. Med. 206, 1465–1472 (2009).
Rogakou, E. P., Pilch, D. R., Orr, A. H., Ivanova, V. S. & Bonner, W. M. DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. J. Biol. Chem. 273, 5858–5868 (1998).
Fischer, M. Census and evaluation of p53 target genes. Oncogene 36, 3943–3956 (2017).
Merritt, A. J. et al. The role of p53 in spontaneous and radiation-induced apoptosis in the gastrointestinal tract of normal and p53-deficient mice. Cancer Res. 54, 614–617 (1994).
Qiu, W. et al. PUMA regulates intestinal progenitor cell radiosensitivity and gastrointestinal syndrome. Cell Stem Cell 2, 576–583 (2008).
Hernandez, P., Gronke, K. & Diefenbach, A. A catch-22: interleukin-22 and cancer. Eur. J. Immunol. 48, 15–31 (2018).
Schumacher, F. et al. A secondary metabolite of Brassicales, 1-methoxy-3-indolylmethyl glucosinolate, as well as its degradation product, 1-methoxy-3-indolylmethyl alcohol, forms DNA adducts in the mouse, but in varying tissues and cells. Arch. Toxicol. 88, 823–836 (2014).
Kiss, E. A. et al. Natural aryl hydrocarbon receptor ligands control organogenesis of intestinal lymphoid follicles. Science 334, 1561–1565 (2011).
Li, Y. et al. Exogenous stimuli maintain intraepithelial lymphocytes via aryl hydrocarbon receptor activation. Cell 147, 629–640 (2011).
Lee, J. S. et al. AHR drives the development of gut ILC22 cells and postnatal lymphoid tissues via pathways dependent on and independent of Notch. Nat. Immunol. 13, 144–151 (2011).
Metidji, A. et al. The environmental sensor AHR protects from inflammatory damage by maintaining intestinal stem cell homeostasis and barrier integrity. Immunity 49, 353–362 (2018).
Eberl, G. & Littman, D. R. Thymic origin of intestinal αβ T cells revealed by fate mapping of RORγt+ cells. Science 305, 248–251 (2004).
Kreymborg, K. et al. IL-22 is expressed by Th17 cells in an IL-23-dependent fashion, but not required for the development of autoimmune encephalomyelitis. J. Immunol. 179, 8098–8104 (2007).
Alonzi, T. et al. Essential role of STAT3 in the control of the acute-phase response as revealed by inducible gene inactivation in the liver. Mol. Cell. Biol. 21, 1621–1632 (2001).
El Marjou, F. et al. Tissue-specific and inducible Cre-mediated recombination in the gut epithelium. Genesis 39, 186–193 (2004).
Schmidt, J. V., Su, G. H., Reddy, J. K., Simon, M. C. & Bradfield, C. A. Characterization of a murine Ahr null allele: involvement of the Ah receptor in hepatic growth and development. Proc. Natl Acad. Sci. USA 93, 6731–6736 (1996).
Walisser, J. A., Glover, E., Pande, K., Liss, A. L. & Bradfield, C. A. Aryl hydrocarbon receptor-dependent liver development and hepatotoxicity are mediated by different cell types. Proc. Natl Acad. Sci. USA 102, 17858–17863 (2005).
Skarnes, W. C. et al. A conditional knockout resource for the genome-wide study of mouse gene function. Nature 474, 337–342 (2011).
Farley, F. W., Soriano, P., Steffen, L. S. & Dymecki, S. M. Widespread recombinase expression using FLPeR (flipper) mice. Genesis 28, 106–110 (2000).
Gronke, K., Kofoed-Nielsen, M. & Diefenbach, A. Isolation and flow cytometry analysis of innate lymphoid cells from the intestinal lamina propria. Methods Mol. Biol. 1559, 255–265 (2017).
Cossarizza, A. et al. Guidelines for the use of flow cytometry and cell sorting in immunological studies. Eur. J. Immunol. 47, 1584–1797 (2017).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Trapnell, C. et al. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Quaroni, A., Wands, J., Trelstad, R. L. & Isselbacher, K. J. Epithelioid cell cultures from rat small intestine. Characterization by morphologic and immunologic criteria. J. Cell Biol. 80, 248–265 (1979).
Takahashi, M., Nakatsugi, S., Sugimura, T. & Wakabayashi, K. Frequent mutations of the β-catenin gene in mouse colon tumors induced by azoxymethane. Carcinogenesis 21, 1117–1120 (2000).
Schumacher, F., Herrmann, K., Florian, S., Engst, W. & Glatt, H. Optimized enzymatic hydrolysis of DNA for LC–MS/MS analyses of adducts of 1-methoxy-3-indolylmethyl glucosinolate and methyleugenol. Anal. Biochem. 434, 4–11 (2013).
Schumacher, F. et al. Detection of DNA adducts originating from 1-methoxy-3-indolylmethyl glucosinolate using isotope-dilution UPLC–ESI–MS/MS. Anal. Chem. 84, 6256–6262 (2012).
Speckmann, B. et al. Selenium increases hepatic DNA methylation and modulates one-carbon metabolism in the liver of mice. J. Nutr. Biochem. 48, 112–119 (2017).
Acknowledgements
We thank M. Di Virgilio, B. Epe, T. Nikolova, J. Ommer and the members of the Diefenbach laboratory for valuable discussions; K. Oberle for technical assistance; the IMB Flow Cytometry Core Facility (J. Hartwig, I. Schäfer and S. Möckel); the IMB Microscopy Core Facility (S. Ritz and J. Schwirz); the Deep Sequencing Facility at the MPI Immunobiology & Epigenetics Freiburg; and EUCOMM for providing targeted ES cells. The work was supported by the European Research Council (ERC StG No. 311377 to A.D.), the European Union’s Horizon 2020 programme (Marie Skłodowska-Curie Grant Agreement No. 744257 to J.Z.), Deutsche Forschungsgemeinschaft (TR-SFB156/A02, TR-SFB241/A01, SPP1937-DI764/9 to A.D. and KL2963/1-1 to C.S.N.K.), Einstein Foundation Berlin (Einstein Professorship to A.D.), the German Rheumatism Research Centre (to A.D. and A.T.) and the Max-Planck Society (IMPRS-MCB, MPI Freiburg to K.G., M.K.-B. and P.P.H.).
Reviewer information
Nature thanks R. Locksley, W. Ouyang and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
K.G. and P.P.H. carried out most of the experiments and analysed the data. J.Z., M.K.-B., F.G., M.W., C.T., L.A. and A.T. helped with experiments and analysis. C.S.N.K. generated Il22ra1fl/fl mice. F.S. and H.G. provided 1-MIM-OH and performed LC–MS/MS analyses. K.G., P.P.H. and A.D. designed the experiments and analysed data. A.D. directed the research and wrote the manuscript with input from co-authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Increased inflammation in Il22−/− mice during AOM–DSS carcinogenesis.
a–f, Mice were given one dose of AOM intraperitoneally, followed by one week of 2% DSS in drinking water. a, Tumour development in Il22+/+ and Il22−/− mice. Scale bar, 1 cm. Tumour number (b) (Il22+/+, n = 9; Il22−/−, n = 8; mean ± s.e.m.; P = 0.022) and regional distribution of tumours (c) (Il22+/+, n = 7; Il22−/−, n = 10; mean ± s.e.m.; rectum, P = 0.0007; total, P = 0.0078). d, Body weight as percentage of initial weight (Il22+/+, n = 7; Il22−/−, n = 10; mean ± s.e.m.). e, Expression of the indicated inflammatory and tumour-promoting genes (n = 4–6; mean ± s.e.m.) in colonic mucosa was assessed by qRT–PCR two days after DSS treatment was stopped. f, Expression of the indicated genes was analysed by qRT–PCR within tumours (n = 3–9, mean ± s.e.m.). g, Schematic representation of the CAC treatment in mice with sporadic inactivation of the Il22ra1 gene (Il22ra1fl/fl;Lgr5creERT2/+;R26R-Confetti and Il22ra1fl/+;Lgr5creERT2/+;R26R-Confetti). Data are representative of two biologically independent experiments.
Extended Data Fig. 2 Generation and validation of Il22ra1flox mice.
a, Schematic representation of the Il22ra1tm1a(EUCOMM)Wtsi allele in targeted ES cells. The position of PCR primers and Southern blot probes is indicated. b, Verification of construct integration into JM8.N4 ES cells by genomic PCR. DNA from untargeted ES cells (Bruce4) served as a control. c, Southern blot after BamHI digestion, confirming the correct integration of the construct, which introduced a novel BamHI recognition site into the genomic DNA. d, Germline transmission was assessed by genomic PCR in the F1 generation. e, LacZ reporter activity was assessed in F1 mice. f, Gene expression of the IL22RA1 binding partners in colonic epithelial cells of the indicated mouse strains was assessed by qRT–PCR (Il22+/+, n = 5; Il22−/−, n = 4; Il22ra1fl/fl;Vil1-cre, n = 4; Ahrfl/fl;Rorc(γt)-cre, n = 3; mean ± s.e.m.). g, Representative immunofluorescence of IL22RA1 (red) in colonic epithelial cells. Scale bars, 50 µm. h, i, Il22ra1fl/fl;Lgr5creERT2/+;R26R-Confetti or Il22ra1fl/+;Lgr5creERT2/+;R26R-Confetti mice were fed with a tamoxifen-containing diet for two weeks. h, Labelling efficiency of colon crypts at different time points after stopping the tamoxifen treatment (day 1: Il22ra1fl/fl, n = 8; Il22ra1fl/+, n = 10; day 7: Il22ra1fl/fl, n = 7; Il22ra1fl/+, n = 8; day 14: Il22ra1fl/fl, n = 7; Il22ra1fl/+, n = 7 visual fields; mean ± s.e.m.). i, Fraction of Confetti+ crypts (mean) with one, two or three colours. j, The indicated mouse strains were fed with tamoxifen-containing diets and subjected to AOM–DSS treatment as indicated. Body weight is shown as percentage of initial weight (Il22ra1fl/+, n = 6; Il22ra1fl/fl, n = 10; mean ± s.e.m.). Data in f–j are representative of two biologically independent experiments.
Extended Data Fig. 3 IL22RA1 expression in tumours from Il22ra1fl/fl;Lgr5creERT2/+;R26R-Confetti mice.
a–d, Representative IL22RA1 immunofluorescence (violet) by Confetti-negative (a, c) and Confetti-positive (b, d) colon tumours from Il22ra1fl/+;Lgr5creERT2/+;R26R-Confetti (a, b) and Il22ra1fl/fl;Lgr5creERT2/+;R26R-Confetti (c, d) mice. e, MFI of IL22RA1 fluorescence across tumour area of the indicated mouse strains with and without prior Cre activation (n = 7, except Il22ra1fl/fl Confetti-negative, n = 8; mean ± s.e.m.).
Extended Data Fig. 4 Identification of IL-22-producing lymphocytes.
a–c, Mice of the indicated genotypes or strains were injected with AOM or PBS (control). Twenty-four hours later, IL-22+ cells among the indicated lymphocyte subsets and absolute numbers of IL-22+ cells were determined. a, Colon of C57BL/6 mice (n = 6; mean ± s.e.m.; P = 0.0154). b, Small intestine of C57BL/6 mice (n = 6; mean ± s.e.m.). c, Colon of C57BL/6 mice (n = 2; mean ± s.e.m.). Data are representative of two biologically independent experiments.
Extended Data Fig. 5 Initiation of the DDR is impaired in Il22−/− mice.
a–c, Il22+/+;Lgr5eGFP and Il22−/−;Lgr5eGFP mice (n = 3 each) were irradiated with 8 Gy (DNA damage) or left untreated (steady-state). LGR5(GFP)+ colon epithelial stem cells were highly purified and gene expression was analysed by RNA-seq. a, Sorting strategy for colonic LGR5+ stem cells. Stem cells were highly purified (>98%) for RNA-seq analysis. b, Modified Venn diagram. All expressed genes with fold change greater than 2 are represented. The bottom half of the diagram represents genes differentially expressed between Il22+/+ and Il22−/− mice at the steady state. ‘Core expression’ contains all genes with detectable transcripts that were not differentially expressed. The top half represents genes differentially expressed 24 h after irradiation in comparison to untreated mice. Of these, 596 genes differed in both genotypes, whereas 638 genes were only regulated in stem cells of Il22+/+ mice and 392 genes were regulated in stem cells of in Il22−/− mice. c, GSEA was performed on all expressed transcripts represented in LGR5+ colon stem cells from Il22+/+ (n = 3) and Il22−/− (n = 3) mice at the steady state. Nominal P value (Kolmogorov–Smirnov) was calculated for ‘Double strand break processing’ (P = 0.01). d, Mice were irradiated with 8 Gy. Expression of DDR effector genes p21 and Puma in colonic epithelial cells was determined by qRT–PCR at the indicated time points after irradiation (n = 3; mean ± s.e.m.). e, List of genes included in the GSEA gene set ‘DDR effector genes’4 (Fig. 2b). f, Mice were irradiated with 8 Gy. Gene expression of Il22bp (also known as Il22ra2) in lamina propria or colon epithelial cells was determined by qRT–PCR at the indicated time points (n = 3; mean ± s.e.m.). g, h, Expression of the indicated genes in colon epithelial cells was determined by qRT–PCR at steady state: MRN complex genes (g; Il22+/+, n = 7; Il22−/−, n = 6; mean ± s.e.m.); DDR transducer genes (h; Il22+/+, n = 10, except Atmin, n = 7 and Parp1, n = 5; Il22−/−, n = 7, except Atmin, n = 4 and Parp1, n = 5; mean ± s.e.m.).
Extended Data Fig. 6 IL-22 regulates ATM expression and formation of γH2AX.
a, Immunoblot analysis of ATM from colon epithelial cells. Densitometric quantification of signal intensity normalized to loading controls (n = 3; mean ± s.e.m.; P = 0.004). b, Schematic representation of STAT3-binding sites in the Atm gene locus. c, STAT3 ChIP was performed to determine specific pulldown at the transcription factor-binding sites (start, mid, end) in the promoter regions of the Atm gene and at a control site within the Atm gene (cold). Genomic DNA from colon epithelial cells of Il22+/+ and Il22−/− mice before and after in vivo stimulation with recombinant IL-22. Results represent specific pulldown relative to a DNA input sample (start, n = 14; mid, n = 7; end, n = 12; cold, n = 8; mean ± s.e.m.). d, e, Mice of the indicated genotypes were irradiated with 8 Gy. d, Immunofluorescence analysis for γH2AX (red) and EpCam (blue) in the entire colon was performed 8 h after irradiation. Scale bars, 500 µm. e, Quantification (control, 4 h and 24 h, n = 2; 8h, n = 3; mean ± s.e.m.) of γH2AX+ nuclei per colon before and at the indicated time points after irradiation. f, Immunoblot analysis of H2AX protein and densitometric quantification of signal intensity (normalized to loading controls) in primary colon epithelial cells of Il22+/+ and Il22−/− mice (mean ± s.e.m.; n = 2). g, h, Mice were irradiated with 8 Gy or injected intraperitoneally with AOM. Immunofluorescence analysis of colon sections was performed 8 h after damage. Quantification (g; n = 3, mean ± s.e.m.) of γH2AX+EpCam+cells and representative immunofluorescence images (h) after γH2AX (red) and EpCam (blue) staining. Scale bars, 25 µm.
Extended Data Fig. 7 Impairment of p53 activation and DDR effector genes in Il22−/− mice.
a–g, Mice were irradiated with 8 Gy or injected with AOM. a, b, Immunoblot analysis of the indicated proteins from primary colon epithelial cells 24 h after irradiation. Densitometric quantification of signal intensity (mean ± s.e.m.; n = 2). Phosphorylated (p)-p53 and actin, p-SMC and SMC, PUMA and actin, and p21 immunoblots were derived from one gel each. Band densities were normalized to loading (p-p53, PUMA) or sample-processing (p-SMC, SMC, p21) controls. c, d, Fold induction of p21 and Puma gene expression in colon epithelial cells 24 h after irradiation or AOM treatment was determined by qRT–PCR (AOM, n = 10; 8 Gy, n = 5; Stat3fl/fl, n = 4; Stat3fl/fl;Vil1-cre, n = 5; mean ± s.e.m.). e, Apoptosis (red, cCaspase-3) in LGR5+ (blue) colon stem cells (arrows). Representative immunofluorescence images 8 h after irradiation. f, g, Quantification of cCaspase-3+EpCam+ colonic epithelial cells 8 h after treatment with AOM or 8 Gy irradiation (f; n = 3; mean ± s.e.m.) and representative immunofluorescence images (g; cCaspase-3, red; EpCam, blue). Scale bars, 25 µm. Data are representative of two (d–g) or three (a–c) biologically independent experiments.
Extended Data Fig. 8 Impairment of DDR-triggered apoptosis in Il22−/− mice.
a–c, Mice of the indicated genotypes were irradiated with 8 Gy and analysed 8 h later. a, Immunofluorescence for cCaspase-3 (red) and EpCam (blue). Scale bars, 50 µm. Quantification of cCaspase-3+ cells by histology (b; Il22+/+, n = 3; Il22−/−, n = 6; Il22ra1fl/fl;Vil1-cre, n = 3; Ahrfl/fl;Rorc(γt)-cre, n = 4, mean ± s.e.m.) or by flow cytometry (c; n = 3; mean ± s.e.m.; P = 0.003). d, IL-22 neutralizing antibody or control IgG was injected intraperitoneally 36 h and 12 h before, and 12 h after AOM application. Representative immunofluorescence images of the colon 24 h after AOM treatment, stained for LGR5 (blue) and cCaspase-3 (red; arrows indicate apoptotic stem cells). Right, quantification of cCaspase-3+ crypts (n = 3; mean ± s.e.m.; P = 0.0039). e, IL-22-responsive IEC6 cells were cultured with or without recombinant IL-22. Cells were irradiated with 2 Gy or left untreated and cultured in the presence of cytochalasin B. DNA was visualized with DAPI 48 h after irradiation and number of cells containing one or more micronuclei was counted (n = 4, mean ± s.e.m.). Representative images of untreated (ctrl) or irradiated (2 Gy) IEC6 cells. Arrow indicates a micronucleus. f, Mice of the indicated genotypes were irradiated with 8 Gy. Quantification of BrdU+ cells in colonic crypts (n = 3; mean ± s.e.m.). Data are representative of two biologically independent experiments.
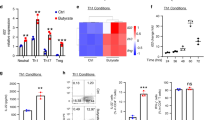
Extended Data Fig. 9 Dietary AhR ligands promote IL-22 production.
a, Wild-type mice were given 1-MIM-OH or carrier control (ctrl) by gavage. Quantification of DNA adducts at the indicated time points (control 4 h, n = 2; control 8 h, n = 4; control 24 h, n = 2; 1-MIM-OH 4 h, n = 4; 1-MIM-OH 8 h, n = 8; 1-MIM-OH 24 h, n = 6; mean ± s.e.m.). b, Schematic representation of the different treatment regimens. c–g, Mice of the indicated genotypes were exposed to 1-MIM-OH (c, f) or I3C (d, e, g) by gavage and analysed 8 h later. c, d, Representative flow cytometry analysis of IL-22 production by the indicated lymphocyte populations after gavage with 1-MIM-OH (c) or I3C (d). CD4+ T cells were defined as CD45+ lineage marker (Lin)+CD4+ cells, ILC3 as CD45+Lin−RORγt+ cells, and γδ T cells as CD45+Lin+γδTCR+ cells. Numbers indicate percentage of cells in gate. e, Representative flow cytometry analysis and quantification of IL-22 and IL-10 production by Lin+CD4+FOXP3+ Treg cells 8 h after gavage with I3C (control, n = 4; I3C, n = 4; mean ± s.e.m.). f, g, Expression of the IL-22 response gene Reg3g in primary colon epithelial cells was determined by qRT–PCR 8 h after gavage with 1-MIM-OH (f; Il22+/+ control, n = 5; Il22+/+ 1-MIM-OH, n = 4; Il22−/− control, n = 3; Il22−/− 1-MIM-OH, n = 6; mean ± s.e.m.) or I3C (g; n = 5; mean ± s.e.m.). h, Percentage of IL-22+ cells among the indicated lymphocyte subsets 8 h after 1-MIM-OH treatment of Ahr−/− mice (control, n = 3; 1-MIM-OH, n = 4; mean ± s.e.m.). i, Numbers of IL-22+ ILC3 cells at different time points after 1-MIM-OH gavage (control, n = 2; 1-MIM-OH, n = 4; 1-MIM-OH 24 h, n = 6; mean ± s.e.m.). Data are representative of two biologically independent experiments.
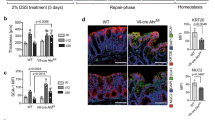
Extended Data Fig. 10 Diet-derived AhR-ligands control IL-22 production.
a–d, Mice were fed for 6 weeks with either normal, grain-based chow (CD), EDD− or EDD+. a, Representative flow cytometry analysis of IL-22 production by the indicated lymphocyte populations. Numbers indicate percentage of cells in gate. b, Expression of the IL-22 responsive gene Reg3g in primary colon epithelial cells was determined by qRT–PCR (n = 5; mean ± s.e.m.). c, d, Representative colon immunofluorescence images 8 h after irradiation stained for γH2AX (red, c) or cCaspase-3 (red, d) and EpCam (blue). Scale bars, 50 µm. Data are representative of three biologically independent experiments. e, Model of IL-22-mediated protection of epithelial stem cells against genotoxic stress. Epithelial stem cells at the intestinal barrier are continuously exposed to a variety of insults. Among them are threats to genomic integrity represented by genotoxins contained in diets or produced by bacteria. Genome integrity in stem cells requires a functioning DDR pathway to limit damage to the genetic code in order to prevent malignant transformation. Our data implicate a homeostatic regulatory loop that assesses the degree of ‘genotoxic danger’ by AhR-mediated sensing of nutrient-derived genotoxins that serve as AhR ligands. AhR signalling in ILC3 and γδ T cells controls IL-22 production. IL-22 signalling in intestinal epithelial stem cells in turn controls components of the DDR. In particular ATM, a PI3-kinase-like molecule and an important upstream transducer of the DDR, was upregulated by IL-22 signalling in stem cells. Following genotoxic stress, ATM controls downstream events of the DDR such as p53 activation and expression of the DDR effector molecules p21 and PUMA, which control DNA repair and apoptosis, respectively. Disturbances of this pathway in mice genetically lacking IL-22, the IL-22 receptor or the AhR led to an impaired DDR to various forms of genotoxic stress. Consequently, intestinal epithelial stem cells from mice with impairment of IL-22 signalling or production accumulated cancer-promoting mutations and showed accelerated development of cancer.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables 1-3. Full table legends are provided in the PDF file.
Supplementary Figures
This file contains the western blot source files.
Source data
Rights and permissions
About this article
Cite this article
Gronke, K., Hernández, P.P., Zimmermann, J. et al. Interleukin-22 protects intestinal stem cells against genotoxic stress. Nature 566, 249–253 (2019). https://doi.org/10.1038/s41586-019-0899-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-0899-7
This article is cited by
-
Group 3 innate lymphoid cells in intestinal health and disease
Nature Reviews Gastroenterology & Hepatology (2024)
-
Disease pathogenesis and barrier functions regulated by group 3 innate lymphoid cells
Seminars in Immunopathology (2024)
-
Hypoxic microenvironment in cancer: molecular mechanisms and therapeutic interventions
Signal Transduction and Targeted Therapy (2023)
-
Endothelial AHR activity prevents lung barrier disruption in viral infection
Nature (2023)
-
Unconventional immune cells in the gut mucosal barrier: regulation by symbiotic microbiota
Experimental & Molecular Medicine (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.