Abstract
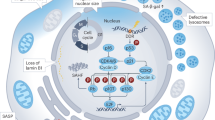
Replicative crisis is a senescence-independent process that acts as a final barrier against oncogenic transformation by eliminating pre-cancerous cells with disrupted cell cycle checkpoints1. It functions as a potent tumour suppressor and culminates in extensive cell death. Cells rarely evade elimination and evolve towards malignancy, but the mechanisms that underlie cell death in crisis are not well understood. Here we show that macroautophagy has a dominant role in the death of fibroblasts and epithelial cells during crisis. Activation of autophagy is critical for cell death, as its suppression promoted bypass of crisis, continued proliferation and accumulation of genome instability. Telomere dysfunction specifically triggers autophagy, implicating a telomere-driven autophagy pathway that is not induced by intrachromosomal breaks. Telomeric DNA damage generates cytosolic DNA species with fragile nuclear envelopes that undergo spontaneous disruption. The cytosolic chromatin fragments activate the cGAS–STING (cyclic GMP-AMP synthase–stimulator of interferon genes) pathway and engage the autophagy machinery. Our data suggest that autophagy is an integral component of the tumour suppressive crisis mechanism and that loss of autophagy function is required for the initiation of cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information files). Reagents are available from J.K. upon reasonable request.
References
Wei, W. & Sedivy, J. M. Differentiation between senescence (M1) and crisis (M2) in human fibroblast cultures. Exp. Cell Res. 253, 519–522 (1999).
Hayashi, M. T., Cesare, A. J., Rivera, T. & Karlseder, J. Cell death during crisis is mediated by mitotic telomere deprotection. Nature 522, 492–496 (2015).
Maciejowski, J. & de Lange, T. Telomeres in cancer: tumour suppression and genome instability. Nat. Rev. Mol. Cell Biol. 18, 175–186 (2017).
Artandi, S. E. & DePinho, R. A. A critical role for telomeres in suppressing and facilitating carcinogenesis. Curr. Opin. Genet. Dev. 10, 39–46 (2000).
Steinberg, M. L. & Defendi, V. Transformation and immortalization of human keratinocytes by SV40. J. Invest. Dermatol. 81, 131s–136s (1983).
Le Poole, I. C. et al. Generation of a human melanocyte cell line by introduction of HPV16 E6 and E7 genes. In Vitro Cell. Dev. Biol. Anim. 33, 42–49 (1997).
Romanov, S. R. et al. Normal human mammary epithelial cells spontaneously escape senescence and acquire genomic changes. Nature 409, 633–637 (2001).
Kiyono, T. et al. Both Rb/p16INK4a inactivation and telomerase activity are required to immortalize human epithelial cells. Nature 396, 84–88 (1998).
Pankiv, S. et al. p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J. Biol. Chem. 282, 24131–24145 (2007).
Yamamoto, A. et al. Bafilomycin A1 prevents maturation of autophagic vacuoles by inhibiting fusion between autophagosomes and lysosomes in rat hepatoma cell line, H-4-II-E cells. Cell Struct. Funct. 23, 33–42 (1998).
Bampton, E. T., Goemans, C. G., Niranjan, D., Mizushima, N. & Tolkovsky, A. M. The dynamics of autophagy visualized in live cells: from autophagosome formation to fusion with endo/lysosomes. Autophagy 1, 23–36 (2005).
Tanida, I. et al. Consideration about negative controls for LC3 and expression vectors for four colored fluorescent protein-LC3 negative controls. Autophagy 4, 131–134 (2008).
Takai, H., Smogorzewska, A. & de Lange, T. DNA damage foci at dysfunctional telomeres. Curr. Biol. 13, 1549–1556 (2003).
van Steensel, B., Smogorzewska, A. & de Lange, T. TRF2 protects human telomeres from end-to-end fusions. Cell 92, 401–413 (1998).
Cho, N. W., Dilley, R. L., Lampson, M. A. & Greenberg, R. A. Interchromosomal homology searches drive directional ALT telomere movement and synapsis. Cell 159, 108–121 (2014).
Caron, P. et al. Non-redundant functions of ATM and DNA-PKcs in response to DNA double-strand breaks. Cell Reports 13, 1598–1609 (2015).
Harding, S. M. et al. Mitotic progression following DNA damage enables pattern recognition within micronuclei. Nature 548, 466–470 (2017).
Fumagalli, M. et al. Telomeric DNA damage is irreparable and causes persistent DNA-damage-response activation. Nat. Cell Biol. 14, 355–365 (2012).
Gisselsson, D. et al. Telomere dysfunction triggers extensive DNA fragmentation and evolution of complex chromosome abnormalities in human malignant tumors. Proc. Natl Acad. Sci. USA 98, 12683–12688 (2001).
Chen, Y. A. et al. Extrachromosomal telomere repeat DNA is linked to ALT development via cGAS–STING DNA sensing pathway. Nat. Struct. Mol. Biol. 24, 1124–1131 (2017).
Doksani, Y. & de Lange, T. Telomere-internal double-strand breaks are repaired by homologous recombination and PARP1/Lig3-dependent end-joining. Cell Reports 17, 1646–1656 (2016).
Ishikawa, H., Ma, Z. & Barber, G. N. STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity. Nature 461, 788–792 (2009).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Comb, W. C., Cogswell, P., Sitcheran, R. & Baldwin, A. S. IKK-dependent, NF-κB-independent control of autophagic gene expression. Oncogene 30, 1727–1732 (2011).
Mathew, R. et al. Autophagy suppresses tumor progression by limiting chromosomal instability. Genes Dev. 21, 1367–1381 (2007).
Karantza-Wadsworth, V. et al. Autophagy mitigates metabolic stress and genome damage in mammary tumorigenesis. Genes Dev. 21, 1621–1635 (2007).
Sarbassov, D. D., Guertin, D. A., Ali, S. M. & Sabatini, D. M. Phosphorylation and regulation of Akt/PKB by the rictor-mTOR complex. Science 307, 1098–1101 (2005).
N’Diaye, E.-N. et al. PLIC proteins or ubiquilins regulate autophagy-dependent cell survival during nutrient starvation. EMBO Rep. 10, 173–179 (2009).
Hahn, W. C. et al. Enumeration of the simian virus 40 early region elements necessary for human cell transformation. Mol. Cell. Biol. 22, 2111–2123 (2002).
Hatch, E. M., Fischer, A. H., Deerinck, T. J. & Hetzer, M. W. Catastrophic nuclear envelope collapse in cancer cell micronuclei. Cell 154, 47–60 (2013).
Arnoult, N. et al. Regulation of DNA repair pathway choice in S and G2 phases by the NHEJ inhibitor CYREN. Nature 549, 548–552 (2017).
O’Sullivan, R. J., Kubicek, S., Schreiber, S. L. & Karlseder, J. Reduced histone biosynthesis and chromatin changes arising from a damage signal at telomeres. Nat. Struct. Mol. Biol. 17, 1218–1225 (2010).
Cesare, A. J. et al. Spontaneous occurrence of telomeric DNA damage response in the absence of chromosome fusions. Nat. Struct. Mol. Biol. 16, 1244–1251 (2009).
Karlseder, J., Smogorzewska, A. & de Lange, T. Senescence induced by altered telomere state, not telomere loss. Science 295, 2446–2449 (2002).
Geigl, J. B., Uhrig, S. & Speicher, M. R. Multiplex-fluorescence in situ hybridization for chromosome karyotyping. Nat. Protocols 1, 1172–1184 (2006).
Acknowledgements
Data are archived at the Salk Institute. We thank P. Adams for discussions and U. Manor and L. Andrade in the Waitt Advanced Biophotonic Core for transmission electron microscopy experiments. J.N. was supported by EMBO (ALTF213-2016) and the Hewitt Foundation. R.R. was supported by the Paul F. Glenn Center for Biology of Aging Research. The Salk Institute Cancer Center Core Grant (P30CA014195), the NIH (R01CA227934, GM087476, R01CA174942), the Donald and Darlene Shiley Chair and the Helmsley, Auen and Highland Street Foundations support J.K.
Reviewer information
Nature thanks M. Narita and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Experiments were designed by J.N., R.J.S. and J.K. Experiments were performed by J.N. (all except those outlined below), R.R. (mCherry–GFP–LC3(G120A) construct in Fig. 1c, irradiation in Fig. 4b, fusions with fibroblasts in Extended Data Fig. 1h–i, DNA extraction in Extended Data Fig. 5a, fusions in Extended Data Figs. 9e, 8b and imaging in Extended Data Fig. 8c), A.C. (Fig. 2d, Extended Data Fig. 4b, telomere dysfunction-induced damage foci with fibroblasts in Extended Data Figs. 1f, g, 8c, d, telomere dysfunction-induced damage foci in Extended Data Fig. 9d), J.M.F. (Extended Data Fig. 5a), B.S. and A.J. (fluorescence in situ hybridization and analysis in Fig. 2e, f, Supplementary Table 1). J.N. and J.K. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Characteristic features of fibroblasts and epithelial cells in crisis.
a, Growth curves of IMR90 and WI38 lung fibroblasts expressing empty vector or vectors encoding SV40-LT or HPV-16 E6E7. Senescence (sen) and crisis plateaus are indicated. b, Immunoblotting of IMR90 and WI38 lung fibroblasts upon expression of SV40-LT or HPV-16 E6E7 with GAPDH as loading control. Two independent experiments were performed. c, Growth curves of HMECs (left) and PrECs (right) expressing vectors encoding P53(DD) and CDK4(R24C). NI, non-infected PrECs. Senescence (sen) and crisis plateaus are indicated. d, Immunoblotting of HMECs that spontaneously bypass senescence (sen) and enter crisis with GAPDH as loading control. Two independent experiments were performed. e, Immunoblotting of PrECs upon expression of P53(DD) and CDK4(R24C) with GAPDH as loading control. Two independent experiments were performed. f, Metaphase chromosomes of growing (PD97) and pre-crisis (PD103) IMR90E6E7 cells. DAPI staining in blue, telomeres in green and γH2AX in red. Two independent experiments were performed. g, Scatter plot showing the number of telomeric γH2AX foci per metaphase. Centre line, mean; error bars, ± s.d. n shows number of metaphases analysed. Two independent experiments were performed. One-way ANOVA; ns, not significant, ∗∗∗P < 0.001. h, Metaphase chromosomes of growing (PD72) and pre-crisis (PD79) WI38SV40 cells. DAPI staining in blue, telomeres in green and centromeres in red. Two independent experiments were performed. i, Scatter plot showing the number of fused chromosomes per metaphase. n shows number of metaphases analysed. Centre line, mean; error bars, ± s.d. n shows number of metaphases analysed. Two independent experiments were performed. One-way ANOVA; ns, not significant, ∗∗∗P < 0.001. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 2 Cells in crisis show characteristics of autophagy and not apoptosis.
a, Cells at the indicated population doublings were stained with annexin V and PI and the percentages of double-positive cells were measured by flow cytometry. Scatter plot with bars showing the percentage of dying cells approaching crisis. Bars represent mean ± s.d. n indicates number of samples analysed. One experiment was performed. One-way ANOVA; ns, not significant, ∗∗∗P < 0.001. b, Representative phase contrast and fluorescence images of growing, crisis, and staurosporine-treated growing cells (1 μM for 6 h). DAPI staining in blue. One experiment was performed. Scale bar, 50 μm. c, Representative electron micrographs of autophagy-related structures present in crisis fibroblasts and epithelial cells. The images show double-membraned autophagosomes and simple-membraned autolysosomes with cargo at different stages of digestion. Two experiments were performed. d, Box and whisker plots showing the number of autophagic vacuoles per field of view. Centre line, median; box limits, first and third quartiles; whiskers, minimum and maximum. n shows number of images analysed. One-way ANOVA; ∗∗∗P < 0.001. AVs, autophagic vacuoles. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 3 Autophagy activation in crisis cells.
a, Representative confocal microscopy images of growing fibroblasts and epithelial cells expressing wild-type mCherry–GFP–LC3 (mCherry-GFP-LC3WT). Cells were grown in complete growth medium (CTR) or Earle’s balanced salt solution (EBSS) (1 h). One experiment was performed. Scale bar, 10 μm. b, LC3 immunofluorescence performed on growing and crisis fibroblasts and epithelial cells. Representative confocal microscopy images. One experiment was performed. Scale bar, 10 μm. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 4 Inhibition of autophagy promotes crisis bypass.
a, Immunoblotting of HMECs and IMR90E6E7 cells expressing non-targeting control shRNA or shRNA against ATG3, ATG5 or ATG7 with GAPDH as loading control. Two experiments were performed. b, Measurement of cell proliferation rates in growing HMECs (PD22) and IMR90E6E7 cells (PD90) expressing non-targeting control shRNA or shRNA against ATG3, ATG5 or ATG7. Cells were stained with CytoLabelling Green Reagent dye and the fluorescence intensity was measured by flow cytometry at day 0 and day 3 post-labelling. Left, plots showing the difference in fluorescence intensity between day 0 and day 3. Right, scatter plots with bars showing the difference in median fluorescence intensity between day 0 and day 3 in HMECs and IMR90E6E7 cells. Bars represent mean ± s.d. n shows number of independent experiments. One-way ANOVA; ns, not significant. c, Dot plots of cell cycle distribution of growing HMECs (PD22) and IMR90E6E7 cells (PD90) expressing non-targeting control shRNA or shRNA against ATG3, ATG5 or ATG7. Three experiments were performed. d, Scatter plots with bars showing the mean percentage of cells in G1, S and G2/M cell cycle phases. Bars represent mean ± s.d. n shows number of independent experiments. One-way ANOVA; ns, not significant. e, Quantification of crystal violet staining. Scatter plot with bars showing the optic density of crystal violet solutions. Bars represent mean ± s.d. n shows number of replicates. One experiment was performed. OD, optical density. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 5 Crisis bypass is associated with DNA damage signalling.
a, TRF analysis of IMR90E6E7 cells expressing non-targeting control shRNA or shRNA against ATG3, ATG5 or ATG7. Genomic DNA was prepared from parental cells (day 0) or cells before crisis (day 40). Two experiments were performed. b, Scatter plots showing the number of telomeric γH2AX foci per metaphase in HMECs and IMR90E6E7 cells expressing non-targeting control shRNA or shRNA against ATG3, ATG5 or ATG7. Centre line, mean; error bars, ± s.d. Samples were taken at the indicated days. n shows number of metaphases analysed. One experiment was performed. One-way ANOVA; ns, not significant, ∗P < 0.05, ∗∗∗P < 0.001. c, Scatter plots showing the number of fused chromosomes per metaphase in HMECs and IMR90E6E7 cells expressing non-targeting control shRNA or shRNA against ATG3, ATG5 or ATG7. Centre line, mean; error bars, ± s.d. Samples were taken at the indicated days. n shows number of metaphases analysed. Two experiments were performed. One-way ANOVA; ns, not significant, ∗∗∗P < 0.001. d, Scatter plots showing the number of non-telomeric γH2AX foci per metaphase in HMECs and IMR90E6E7 cells expressing non-targeting control shRNA or shRNA against ATG3, ATG5 or ATG7. Centre line, mean; error bars, ± s.d. Samples were taken at the indicated days. n shows number of metaphases analysed. One experiment was performed. One-way ANOVA; ns, not significant, ∗P < 0.05, ∗∗∗P < 0.001. For gel source data see Supplementary Fig. 1.
Extended Data Fig. 6 Telomere dysfunction activates autophagy.
a, Metaphase chromosomes of post-senescent HMECs (PD22) expressing non-targeting control shRNA or shRNA against TRF1 or TRF2. Metaphases were prepared from cells at day 6 post-transduction. Mock represents non-transduced cells. DAPI staining in blue, telomeres in green and γH2AX in red. Arrowheads indicate chromosome fusion events. Two independent experiments were performed. b, Left, scatter plot showing the mean number of telomeric γH2AX foci per metaphase at day 6 post-transduction. Right, scatter plot showing the number of fused chromosomes per metaphase at day 6 post-transduction. Centre line, mean. n shows number of metaphases analysed. Two independent experiments were performed. One-way ANOVA; ns, not significant, ∗∗∗P < 0.001. c, LC3-II and P62 turnover assays. HMECs and IMR90E6E7 cells expressing non-targeting control shRNA or shRNA targeting TRF2 at day 6 post-transduction were treated for bafilomycin A1 (50 nM for 24 h) or MG132 (10 μM for 24 h). Top, experimental timeline. Bottom, immunoblotting of HMECs and IMR90E6E7 cells at day 6 post-transduction with GAPDH as loading control. One experiment was performed. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 7 Telomere dysfunction activates autophagy.
a, Box and whisker plots showing the number of telomeric and non-telomeric γH2AX foci per cell upon increasing doses of ionizing radiation (IR), bleocin, or Shield1-based AsiSI, catalytically inactive TRF1–FokI(450A), or wild-type TRF1–FokI 1 h after damage induction. Centre line, median; box limits, first and third quartiles; whiskers, 10th and 90th percentiles. Cells used are post-senescent HMECs (PD23). Two independent experiments were performed. n shows number of cells analysed. One-way ANOVA; ns, not significant, ∗P < 0.05, ∗∗P < 0.01, ∗∗∗P < 0.001. b, Post-senescent HMECs (PD23) were transfected with non-targeting siRNA or siRNA targeting DNAPKcs or ligase IV 48 h before damage induction. Representative confocal images of cells before damage, and 1 h or 48 h post-damage. DAPI staining in blue, telomeres in green and γH2AX in red. Two independent experiments were performed. c, Box and whisker plots showing the number of γH2AX foci per cell at 1, 12, 24 and 48 h after damage induction, as in a. d, Immunoblotting of IMR90E6E7 cells (PD40) at 12, 24 and 48 h post-induction of TRF1–FokI(450A) or wild-type TRF1–FokI. Mock represents non-transduced cells. GAPDH loading control. Two independent experiments were performed. e, LC3-II and P62 turnover assays. Control (non-induced) and wild-type TRF1–FokI-expressing cells were treated with bafilomycin A1 (50 nM for 24 h) or MG132 (10 μM for 24 h). Top, experimental timeline. Bottom, immunoblotting of HMECs and IMR90E6E7 cells before and 48 h after induction of wild-type TRF1–FokI. GAPDH loading control. One experiment was performed. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 8 Crisis cells display cytosolic DNA species.
a, Top, representative confocal microscopy images of crisis IMR90E6E7 cells (PD108) expressing RFP–NLS or immunostained with antibodies against lamin A or lamin B1. Bottom, grouped stacked bars showing the ratio of positive and negative micronuclei for each of the indicated stains. n shows number of micronuclei analysed. b, Top, representative confocal microscopy images of crisis IMR90E6E7 cells (PD108) immunostained with mitotracker dye. Bottom, grouped stacked bars showing the ratio of positive and negative cytosolic DNA products for mitotracker staining. n shows number of cytosolic DNA products analysed. c, Top, representative confocal microscopy images of crisis IMR90E6E7 cells, growing IMR90E6E7 cells expressing shRNA targeting TRF2 (day 6 post-transduction) or growing IMR90E6E7 cells expressing wild-type TRF1–FokI (48 h post-induction). The corresponding population doublings are indicated. DAPI staining in blue, telomeres in green. Bottom, grouped stacked bars showing the ratio of positive and negative micronuclei for telomeres. n shows number of micronuclei analysed. d, Top, representative confocal image of U2OS cells displaying extrachromosomal telomeric repeat (ECTR) DNA. Bottom, scatter plot with bars showing the percentage of IMR90E6E7 and U2OS cells positive for ECTRs. Growing and crisis IMR90E6E7 cells, growing IMR90E6E7 cells expressing shRNA targeting TRF2 (day 6 post-transduction) and growing IMR90E6E7 cells expressing wild-type TRF1–FokI (48 h post-induction) were used. Bars represent mean. The corresponding population doublings are indicated. Two independent experiments were performed. n shows number of cells analysed. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 9 Telomere fusion is required for autophagy activation.
a, Immunoblotting of growing IMR90E6E7 cells (PD30) upon siRNA targeting DNAPKcs or ligase IV 48 h post-transfection (experiment in Fig. 4b). γTubulin loading control. Two independent experiments were performed. b, Experimental timeline for c–g. c, Metaphase chromosomes of growing IMR90E6E7 cells (PD55) upon shRNA targeting of TRF2 and siRNA targeting of ligase IV at day 6 post-transduction. Left, DAPI staining in blue, telomeres in green and centromeres in red. Arrowheads indicate chromosome fusion events. Two independent experiments were performed. d, Scatter plot showing the number of telomeric γH2AX foci per metaphase. Centre line, mean; error bars, ± s.d. n shows number of metaphases analysed. Two experiments were performed. One-way ANOVA, ns: not significant, ∗∗∗P < 0.001. e, Scatter plots showing the number of fused chromosomes per metaphase, as in d. f, Grouped stacked bars showing the percentage of IMR90E6E7 cells with nucleoplasmic bridges, micronuclei and cytoplasmic chromatin fragments upon shRNA targeting TRF2 and siRNA targeting ligase IV. Bars represent mean ± s.d. n shows number of cells analysed. Three experiments were performed. g, Immunoblotting of IMR90E6E7 cells upon shRNA targeting TRF2 and siRNA targeting ligase IV with γtubulin as loading control. One experiment was performed. h, Scatter plot showing the number of fused chromosomes per metaphase 48 h after induction of wild-type TRF1–FokI, as in d. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 10 cGAS–STING pathway is required for telomere-driven autophagy.
a, Quantification of crystal violet staining. Scatter plot with bars showing the optic density of crystal violet solutions. Bars represent mean ± s.d. n shows number of replicates. One experiment was performed. b, Representative confocal microscopy images of IMR90E6E7 cells expressing sh-scramble or shRNA targeting CGAS or STING with DAPI staining. Three independent experiments were performed. Scale bar, 10 μm. For gel source data, see Supplementary Fig. 1.
Supplementary information
Supplementary Figure 1
This file contains the uncropped images of all gels.
Supplementary Table 1
This file contains Supplementary Information Table 1: Chromosomal aberrations in IMR90 E6E7 analyzed by Multicolor fluorescence in situ hybridization (M-FISH). Each image represents one specific metaphase analyzed by M-FISH. The type of aberration and the total number of chromosome (n) are indicated. Abbreviations: del, deletion; t, translocation; i, isochromosome; der, derivative chromosome. Aberrations are highlighted in different colors: structural aberration (yellow), chromosome loss (blue), chromosome gain (green).
Source data
Rights and permissions
About this article
Cite this article
Nassour, J., Radford, R., Correia, A. et al. Autophagic cell death restricts chromosomal instability during replicative crisis. Nature 565, 659–663 (2019). https://doi.org/10.1038/s41586-019-0885-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-0885-0
This article is cited by
-
cGAS suppresses hepatocellular carcinoma independent of its cGAMP synthase activity
Cell Death & Differentiation (2024)
-
Harnessing innate immune pathways for therapeutic advancement in cancer
Signal Transduction and Targeted Therapy (2024)
-
Second messenger 2'3'-cyclic GMP-AMP (2'3'-cGAMP): the cell autonomous and non-autonomous roles in cancer progression
Acta Pharmacologica Sinica (2024)
-
Scrambling the genome in cancer: causes and consequences of complex chromosome rearrangements
Nature Reviews Genetics (2024)
-
The two sides of chromosomal instability: drivers and brakes in cancer
Signal Transduction and Targeted Therapy (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.