Abstract
Regulatory T cells (Treg cells), a distinct subset of CD4+ T cells, are necessary for the maintenance of immune self-tolerance and homeostasis1,2. Recent studies have demonstrated that Treg cells exhibit a unique metabolic profile, characterized by an increase in mitochondrial metabolism relative to other CD4+ effector subsets3,4. Furthermore, the Treg cell lineage-defining transcription factor, Foxp3, has been shown to promote respiration5,6; however, it remains unknown whether the mitochondrial respiratory chain is required for the T cell-suppression capacity, stability and survival of Treg cells. Here we report that Treg cell-specific ablation of mitochondrial respiratory chain complex III in mice results in the development of fatal inflammatory disease early in life, without affecting Treg cell number. Mice that lack mitochondrial complex III specifically in Treg cells displayed a loss of T cell-suppression capacity without altering Treg cell proliferation and survival. Treg cells deficient in complex III showed decreased expression of genes associated with Treg function, whereas Foxp3 expression remained stable. Loss of complex III in Treg cells increased DNA methylation as well as the metabolites 2-hydroxyglutarate (2-HG) and succinate that inhibit the ten-eleven translocation (TET) family of DNA demethylases7. Thus, Treg cells require mitochondrial complex III to maintain immune regulatory gene expression and suppressive function.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All RNA-seq and DNA methylation data have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE120452. All other data from the manuscript are available from the corresponding author on request.
References
Lu, L., Barbi, J. & Pan, F. The regulation of immune tolerance by FOXP3. Nat. Rev. Immunol. 17, 703–717 (2017).
Josefowicz, S. Z., Lu, L.-F. & Rudensky, A. Y. Regulatory T cells: mechanisms of differentiation and function. Annu. Rev. Immunol. 30, 531–564 (2012).
Gerriets, V. A. et al. Metabolic programming and PDHK1 control CD4+ T cell subsets and inflammation. J. Clin. Invest. 125, 194–207 (2014).
Michalek, R. D. et al. Cutting edge: distinct glycolytic and lipid oxidative metabolic programs are essential for effector and regulatory CD4+ T cell subsets. J. Immunol. 186, 3299–3303 (2011).
Gerriets, V. A. et al. Foxp3 and Toll-like receptor signaling balance Treg cell anabolic metabolism for suppression. Nat. Immunol. 17, 1459–1466 (2016).
Angelin, A. et al. Foxp3 reprograms T cell metabolism to function in low-glucose, high-lactate environments. Cell Metab. 25, 1282–1293 (2017).
Pastor, W. A., Aravind, L. & Rao, A. TETonic shift: biological roles of TET proteins in DNA demethylation and transcription. Nat. Rev. Mol. Cell Biol. 14, 341–356 (2013).
Sena, L. A. et al. Mitochondria are required for antigen-specific T cell activation through reactive oxygen species signaling. Immunity 38, 225–236 (2013).
Rubtsov, Y. P. et al. Regulatory T cell-derived interleukin-10 limits inflammation at environmental interfaces. Immunity 28, 546–558 (2008).
Brunkow, M. E. et al. Disruption of a new forkhead/winged-helix protein, scurfin, results in the fatal lymphoproliferative disorder of the scurfy mouse. Nat. Genet. 27, 68–73 (2001).
Liston, A. et al. Lack of Foxp3 function and expression in the thymic epithelium. J. Exp. Med. 204, 475–480 (2007).
Fontenot, J. D., Gavin, M. A. & Rudensky, A. Y. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat. Immunol. 4, 330–336 (2003).
Delgoffe, G. M. et al. Stability and function of regulatory T cells is maintained by a neuropilin-1–semaphorin-4a axis. Nature 501, 252–256 (2013).
Zhang, B., Chikuma, S., Hori, S., Fagarasan, S. & Honjo, T. Nonoverlapping roles of PD-1 and FoxP3 in maintaining immune tolerance in a novel autoimmune pancreatitis mouse model. Proc. Natl Acad. Sci. USA 113, 8490–8495 (2016).
Sauer, A. V. et al. Alterations in the adenosine metabolism and CD39/CD73 adenosinergic machinery cause loss of Treg cell function and autoimmunity in ADA-deficient SCID. Blood 119, 1428–1439 (2012).
Joller, N. et al. Treg cells expressing the coinhibitory molecule TIGIT selectively inhibit proinflammatory Th1 and Th17 cell responses. Immunity 40, 569–581 (2014).
Shalev, I. et al. Targeted deletion of fgl2 leads to impaired regulatory T cell activity and development of autoimmune glomerulonephritis. J. Immunol. 180, 249–260 (2008).
Ohkura, N. et al. T cell receptor stimulation-induced epigenetic changes and Foxp3 expression are independent and complementary events required for Treg cell development. Immunity 37, 785–799 (2012).
Ohkura, N., Kitagawa, Y. & Sakaguchi, S. Development and maintenance of regulatory T cells. Immunity 38, 414–423 (2013).
McGrath-Morrow, S. A. et al. DNA methylation regulates the neonatal CD4+ T-cell response to pneumonia in mice. J. Biol. Chem. 293, 11772–11783 (2018).
Kitagawa, Y. et al. Guidance of regulatory T cell development by Satb1-dependent super-enhancer establishment. Nat. Immunol. 18, 173–183 (2016).
Ansó, E. et al. The mitochondrial respiratory chain is essential for haematopoietic stem cell function. Nat. Cell Biol. 19, 614–625 (2017).
Oldham, W. M., Clish, C. B., Yang, Y. & Loscalzo, J. Hypoxia-mediated increases in l-2-hydroxyglutarate coordinate the metabolic response to reductive stress. Cell Metab. 22, 291–303 (2015).
Intlekofer, A. M. et al. Hypoxia induces production of l-2-hydroxyglutarate. Cell Metab. 22, 304–311 (2015).
Mullen, A. R. et al. Oxidation of alpha-ketoglutarate is required for reductive carboxylation in cancer cells with mitochondrial defects. Cell Reports 7, 1679–1690 (2014).
Tyrakis, P. A. et al. S-2-hydroxyglutarate regulates CD8+ T-lymphocyte fate. Nature 540, 236–241 (2016).
Xu, W. et al. Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of α-ketoglutarate-dependent dioxygenases. Cancer Cell 19, 17–30 (2011).
Xiao, M. et al. Inhibition of α-KG-dependent histone and DNA demethylases by fumarate and succinate that are accumulated in mutations of FH and SDH tumor suppressors. Genes Dev. 26, 1326–1338 (2012).
Collison, L. W. & Vignali, D. A. A. In vitro Treg suppression assays. Methods Mol. Biol. 707, 21–37 (2011).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative C T method. Nat. Protoc. 3, 1101–1108 (2008).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Hill, J. A. et al. Foxp3 transcription-factor-dependent and -independent regulation of the regulatory T cell transcriptional signature. Immunity 27, 786–800 (2007).
Meissner, A. et al. Reduced representation bisulfite sequencing for comparative high-resolution DNA methylation analysis. Nucleic Acids Res. 33, 5868–5877 (2005).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Feng, H., Conneely, K. N. & Wu, H. A Bayesian hierarchical model to detect differentially methylated loci from single nucleotide resolution sequencing data. Nucleic Acids Res. 42, e69 (2014).
Workman, C. J. et al. In vivo Treg suppression assays. Methods Mol. Biol. 707, 119–156 (2011).
Acknowledgements
We thank Robert H. Lurie Cancer Center Flow Cytometry facility and Metabolomics Core, and the High Throughput RNA-Seq Center, within the Division of Pulmonary and Critical Care and the Mouse Histology and Phenotyping Laboratory at Northwestern University. We thank K. Nam for processing of RNA sequencing samples. This research was supported in part through the computational resources and staff contributions provided by Quest high performance computing cluster. This work was supported by the NIH (R35CA197532, 5P01AG049665, 5P01HL071643) to N.S.C., and NIH (T32 T32HL076139) to S.E.W. B.D.S was supported by NIH K08HL128867 and the Francis Family Foundation’s Parker B. Francis Research Opportunity Award. E.M.S is a Cancer Research Institute Irvington Fellow supported by the Cancer Research Institute.
Reviewer information
Nature thanks R. Johnson and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
S.E.W., E.M.S., M.M.M., and I.M.-R. carried out most of the experiments. B.D.S. and K.A.H. performed the mRRBS experiments and analysis. L.A.S. provided technical expertise and carried out some of the initial experiments. S.E.W., C.A.M. and H.A.-V. performed the RNA sequencing and analysis. S.E.W. and P.G. conducted and analysed metabolomics. P.T.S. generated the RISP KO mice. S.E.W., B.D.S., E.M.S., L.A.T. and N.S.C. provided intellectual input and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Loss of RISP in Treg cells results in T cell proliferation, activation and immune infiltration into multiple organs.
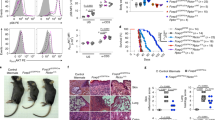
a, Representative images of skin changes observed in RISP KO (top) mice compared to RISP wild-type mice. b, c, Weight (b) and total cellularity (c) of the spleen (RISP wild type, n = 10; RISP KO, n = 9), lymph nodes (RISP wild type, n = 10; RISP KO n = 9) and thymus (RISP wild type, n = 5; RISP KO n = 5) in RISP wild-type and RISP KO mice at 3 weeks of age. d, Representative images from three-week-old RISP wild-type and RISP KO mice. The RISP KO thymuses show thinning of the thymic cortex secondary to lymphoid depletion. There is marked perivascular inflammation in the lung and liver, and dermal thickening and profound inflammation in the skin. e, f, CD4+ (e; RISP wild type, n = 10; RISP KO; n = 9) and CD8+ (f; RISP wild type, n = 10; RISP KO, n = 9) T cell numbers in the spleen and lymph nodes in three-week-old mice. g, h, Percentage of naive (CD62L+CD44−) and effector (CD62L−CD44+) cells in the total CD4+ (g; RISP wild type, n = 5; RISP KO, n = 5) and CD8+ (h; RISP wild type, n = 5; RISP KO, n = 5) T cell populations. Images are representative of at least three mice collected on independent days. Data are mean ± s.d. and were analysed with two-tailed t-test (b, c; **P < 0.001, P values in Source Data) or with multiple two-tailed t-tests using a two-stage linear step-up procedure of Benjamini, Krieger and Yekutieli, with Q = 0.01(e–h). Each cell type was analysed individually, without assuming a consistent s.d. (**Q < 0.001, Q values in Source Data). All data points on graphs represent individual mice isolated and analysed on at least two separate days.
Extended Data Fig. 2 Mice with regulatory T cells deficient in RISP do not display thymic dysfunction early in life.
a, b, Thymic weight (a) and total thymocyte number (b) observed in 10-day-old RISP KO (n = 5) and RISP wild-type (n = 5) mice. c, Absolute cell numbers of double-negative (DN; CD4−CD8−), double-positive (DP; CD4+CD8+), CD8 single-positive (CD8 SP; CD4−CD8+) and CD4 single-positive (CD4 SP; CD4+CD8−) populations from the thymuses of 10-day-old RISP KO (n = 5) and RISP wild-type (n = 5) mice. d, Foxp3–YFP+ and Foxp3–YFP+ CD4 SP absolute cell numbers in the thymuses of 10-day-old RISP KO (n = 5) and RISP wild-type (n = 5) mice. e, Representative contour plot of the Treg cell compartment in the superficial lymph nodes at three weeks of age. Contour plots are representative of at least three independent experiments with a total of at least five mice. Numbers in dot plot quadrants indicate percentage of cells. Data are mean ± s.d.; two-tailed t-test (a, b; P values in Source Data); multiple two-tailed t-tests using a two-stage linear step-up procedure of Benjamini, Krieger and Yekutieli, with Q = 0.01 (c, d). Each cell type was analysed individually, without assuming a consistent s.d. (Q values in Source Data). All data points on graphs represent individual mice isolated and analysed on at least two separate days.
Extended Data Fig. 3 Loss of RISP does not impair expression of classic Treg cell markers, activation markers and proliferation, but impairs Treg suppressive function.
a, Expression of CD25 on CD4+Foxp3–YFP+ cells isolated from the spleen and lymph nodes of three-week-old RISP KO and RISP wild-type mice. b, Surface expression of CTLA-4 and GITR on CD4+Foxp3–YFP+CD25+ cells from three-week-old RISP KO and RISP wild-type mice. c, Cellular expression of EOS and Helios from CD4+Foxp3+CD25+ cells isolated from the spleen and lymph nodes. d, Surface expression of activation makers CD44, CD69, CD103, ICOS and OX40 on CD4+Foxp3–YFP+CD25+ cells isolated from the spleen and lymph nodes. e, Ki-67 expression in CD4+Foxp3+CD25+ cells from the spleen and lymph nodes. f, Percentage of central Treg (cTreg) and effector Treg (eTreg) cells among Treg cells isolated from the spleen and lymph nodes of three-week-old RISP KO (n = 4 for both tissues) and RISP wild-type (spleen, n = 6, lymph nodes, n = 8) mice with representative contour plot. g, Representative histograms of CD69, CD73 and NRP1 expression on cTreg and eTreg cells isolated from lymph nodes of three-week-old RISP KO and RISP wild-type mice. h, Cell trace violet (CTV) dilution in CD8+ effector T cells stimulated to proliferate with varying ratios of RISP wild-type and RISP KO CD4+Foxp3–YFP+CD25+ cells isolated from the lymph nodes of three-week-old mice. Data are representative of five mice, performed on at least three separate days. Numbers in top left corner represent the division index of each histogram. i, Representative images of the colons from RAG1-deficient mice, one month following adoptive transfer. Images are representative of at least three mice collected on independent days. Contour plots are representative at least three independent experiments with at least four mice. Numbers in dot plot quadrants indicate percentage of cells. Histograms are representative of at least three independent experiments with at least five mice in total. In graphs: blue, RISP wild type; red, RISP KO. Numbers on histograms represent mean fluorescence intensity (MFI) of depicted samples; y axis of all histograms represents percentage of maximum. Data in f analysed with multiple two-tailed t-tests using a two-stage linear step-up procedure of Benjamini, Krieger and Yekutieli, with Q = 0.01; Q values in Source Data. Each cell type was analysed individually, without assuming a consistent s.d. All data points on graphs represent individual mice isolated and analysed on at least two separate days.
Extended Data Fig. 4 Loss of complex III subunit QPC in Treg cells gives rise to a lethal inflammatory disorder.
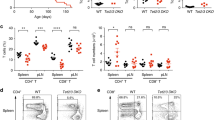
a, Scheme illustrating the strategy used to generate Uqcrq floxed and excised alleles. b, Expression of Uqcrq mRNA in CD4+Foxp3–YFP+CD25+ cells from three-week-old QPC KO (n = 4) relative to QPC wild-type (n = 4) mice. Uqcrq expression was normalized to Rpl19 expression. c, d, OCR (c) and ECAR (d) of CD4+Foxp3–YFP+CD25+ cells isolated from three-week-old QPC wild-type (n = 3) and QPC KO (n = 3) mice e, Survival of QPC wild-type (n = 8) and QPC KO (n = 9) mice (P = 0.0002 using one-sided log-rank test). f–h, Weight of spleens (f), lymph nodes (g) and thymuses (h) from three-week-old QPC wild-type (n = 4) and QPC KO (n = 5) mice. i–k, Total cellularity of the spleen (i), lymph nodes (j) and thymuses (k) from three-week-old QPC wild-type (n = 4) and KO (n = 5) mice. l, m, CD4+ (l) and CD8+ (m) T cell numbers in the spleen and lymph nodes in three-week-old QPC wild-type (n = 4) and QPC KO (n = 5) mice. n, o, Percentage of naive (CD62L+CD44−) and effector (CD62L−CD44+) cells of the total CD4+ (n) and CD8+ (o) T cells in three-week-old QPC wild-type (n = 4) and QPC KO (n = 5) mice. p, Absolute number of Treg cells in the spleen and lymph nodes from three-week-old QPC wild-type (n = 4) and QPC KO (n = 5) mice. Data are mean ± s.d.; two-tailed t-test (b, f–k; *P < 0.01, **P < 0.001, P values in Source Data); multiple two-tailed t-tests using a two-stage linear step-up procedure of Benjamini, Krieger and Yekutieli, with Q = 0.01 (c, d, l–p; *Q < 0.01, **Q <0.001, Q values in Source Data). Each cell type was analysed individually, without assuming a consistent s.d. All data points on graphs represent individual mice isolated and analysed on at least two separate days.
Extended Data Fig. 5 Loss of QPC in Treg cells impairs suppressive function without altering Foxp3 expression.
a, Representative images of the spleen, superficial lymph nodes, mesenteric lymph nodes and thymuses from QPC iKO mice compared to QPC wild-type mice treated with tamoxifen for 28 days. b, Cellularity of the spleen and lymph nodes from QPC wild-type (n = 4) and QPC iKO (n = 4) mice treated with tamoxifen for 28 days. c, Histology of the skin from QPC iKO and QPC wild-type mice. d, Percentage of CD4+ T cells (CD45+CD4+), CD8+ T cells (CD45+CD8+), Treg cells (CD45+CD4+Foxp3–GFP+) and non-T leukocytes (CD45+CD4−CD8−) from the spleen and lymph nodes of QPC wild-type (n = 4) and QPC iKO (n = 4) mice, that express tdTomato–RFP after two weeks with tamoxifen treatment every third day. e, Percentage of thymic epithelial cells (CD45−EpCAM+) expressing tdTomato–RFP from QPC wild-type (n = 4) and QPC iKO (n = 4) mice after two weeks with tamoxifen treatment every third day. f, g, Total cellularity of the spleen (f) and lymph nodes (g) from QPC wild-type (n = 4) and QPC iKO (n = 4) mice, three months after three doses of tamoxifen. h, Representative dot plots of the splenic Treg cell compartment in QPC iKO and QPC wild-type mice, three months after tamoxifen treatment. Numbers in quadrants indicate percentage of cells. Images are representative of at least three mice collected on three different days. Data are mean ± s.d. (b, c, e–h); two-tailed t-tests (b, c, f–h; *P < 0.05, **P < 0.01, P values in Source Data); multiple two-tailed t-tests using a two-stage linear step-up procedure of Benjamini, Krieger and Yekutieli (e), with Q = 0.01. Each cell type was analysed individually, without assuming a consistent s.d. (**Q < 0.001, Q values in Source Data). All data points on graphs represent individual mice isolated and analysed on at least two separate days.
Extended Data Fig. 6 Loss of RISP in Treg cells results in major alterations in gene expression including downregulation of known Treg cell suppressive genes in three-week-old mice.
a, Hierarchical clustering showing changes in gene expression in CD4+Foxp3–YFP+CD25+ cells isolated from three-week-old RISP KO (n = 4) and RISP wild-type (n = 4) mice. b, Volcano plot showing differential gene expression in CD4+Foxp3–YFP+CD25+ cells isolated from three-week-old RISP KO (n = 4) and RISP wild-type (n = 4) mice. Uqcrsf1 (encoding for RISP) is not shown on the plot (log2(expression in RISP KO/expression in RISP WT) = –4.83, −log10(P value) = 203). c, Normalized enrichment scores from gene set enrichment analysis from the hallmark gene set in the molecular signatures database v.6.0 (http://software.broadinstitute.org/gsea/msigdb) comparing the gene set generated from a. d, Heat map of Treg cell-signature genes that are differentially expressed (adjusted P < 0.01) in RISP KO (n = 4) versus RISP wild-type (n = 4) mice. e, Surface expression of CD73, NRP1, PD1, and TIGIT on CD4+Foxp3–YFP+CD25+ cells isolated from the spleen and lymph nodes of three-week-old RISP KO and RISP wild-type mice. Histograms are representative at least two independent experiments with a total of at least four mice. Numbers above histograms represent MFI of depicted samples. Data in a–d are from one RNA sequencing experiment. CD4+Foxp3–YFP+CD25+ cells from RISP wild-type (n = 4) and RISP KO (n = 4) mice were isolated on different days. cDNA library production and sequencing was performed once and all eight RNA samples were prepared together.
Extended Data Fig. 7 Mice that contain a Treg cell compartment with chimeric RISP deficiency do not develop inflammatory disease but display cell-autonomous impairment in expression of a Treg-specific transcription profile.
a, b, Total number of cells in the spleen (a) and lymph nodes (b) of adult (8–12-week-old) RISP chimeric KO (n = 4) and chimeric wild-type (n = 4) mice. c, CD4+ and CD8+ T cell numbers in the spleen and lymph nodes of adult RISP chimeric KO (n = 4) and chimeric wild-type (n = 4) mice. d, Percentage of total CD4+ cells that are either CD4+Foxp3–YFP+CD25+ (chimeric Treg cells) and CD4+Foxp3+CD25+ (total Treg cells) from adult RISP chimeric KO (n = 4) and RISP chimeric wild-type (n = 4) mice. e, Percentage of chimeric Treg cells out of all Treg cells in adult RISP chimeric KO (n = 4) and chimeric wild-type (n = 4) mice. f, g, Ratio of surface expression of CD44, ICOS (f) and CTLA-4 (g) on RISP chimeric KO Treg cells (CD4+Foxp3–YFP+CD25+) relative to RISP chimeric wild-type Treg cells (CD4+Foxp3–YFP−CD25+) from pooled samples of the spleen and lymph node in adult RISP chimeric KO (n = 4) and RISP chimeric wild-type mice (n = 4). h, Hierarchical clustering showing changes in gene expression in CD4+Foxp3–YFP+CD25+ cells isolated from adult RISP chimeric KO (n = 4) and RISP chimeric wild-type mice (n = 4). i, Normalized enrichment scores from gene set enrichment analysis from the hallmark gene set in the molecular signatures database v.6.0 on the dataset shown in h. j, Ratio of surface expression of CD73 and NRP1 on RISP chimeric Treg cells relative to RISP wild-type Treg cells from pooled samples of the spleen and lymph node in adult RISP chimeric KO (n = 4) and RISP chimeric wild-type (n = 4) mice. k, Venn diagram showing overlap of differentially expressed genes (adjusted P < 0.01) from Treg cells (CD4+Foxp3–YFP+CD25+) isolated from three-week-old RISP KO (n = 4) and RISP wild-type (n = 4) mice (RISP three-week-old) versus chimeric Treg cells (CD4+Foxp3–YFP+CD25+) isolated from adult RISP chimeric KO (n = 4) and RISP chimeric wild-type (n = 4) mice. P values were calculated using hypergeometric similarity measure. In a–f, g, j, data are mean ± s.d.; two-tailed t-test (a, b, f, g, j; *P < 0.05, **P < 0.01, P values in Source Data); multiple two-tailed t-tests using a two-stage linear step-up procedure of Benjamini, Krieger and Yekutieli, with Q = 0.01 (c–e). Each cell type was analysed individually, without assuming a consistent s.d. (**Q <0.001, Q values in Source Data). All data points on graphs represent individual mice isolated and analysed on at least 2 separate days. Data in h, i are from one RNA sequencing experiment. CD4+Foxp3–YFP+CD25+ cells from RISP chimeric wild-type (n = 4) and RISP chimeric KO (n = 4) mice were isolated on different days. cDNA library production and sequencing was performed once and all eight RNA samples were prepared together.
Extended Data Fig. 8 Loss of RISP in Treg cells alters DNA methylation without affecting the Foxp3 locus in three-week-old mice.
a, b, CpG methylation around the TSS and transcriptional end site (TES) (a) and the start (S) and end (E) of Treg cell-specific super-enhancer (SE) elements with 20 kb of flanking sequence (b) in CD4+Foxp3–YFP+CD25+ cells from three-week-old RISP KO (n = 4) and RISP wild-type (n = 4) mice. c, DNA methylation status of the super-enhancer associated with the Foxp3 locus in RISP KO and RISP wild-type Treg cells. d, e, Percentage of methylated CpG sites around the TSS of differentially downregulated (d) and upregulated (e) genes (log2(expression in RISP KO/expression in RISP wild type) ≥ 2, FDR Q < 0.05) from Extended Data Fig. 6. In a, d, e, mRRBS data were smoothed with a sliding window size of 20 CpGs and a step of 10 CpGs for sites with better than 5× coverage. In b, Treg cell-specific super-enhancer data were normalized over 1-kb windows. Average CpG methylation of n = 3 biological replicates per group is shown. f, Venn diagram analysis partitioning differentially methylated loci (DML) and differentially expressed genes (DEG). g, Cumulative distribution function of differentially methylated CpGs within 100 kilobases of 87 differentially expressed genes that were downregulated (log2(expression in RISP KO/expression in RISP wild type) ≤ 0) in RISP chimeric KO Treg cells. Data points represent average methylation at each gene locus. h, Heat map displaying the methylation state (beta) of differentially methylated CpGs at gene loci that were both hypermethylated and differentially downregulated (adjusted P < 0.01) in the RISP chimeric KO Treg cells. Data in c are mean ± s.e.m. and were analysed by two-tailed t-test. In a–e, CD4+Foxp3–YFP+CD25+ cells from RISP wild-type (n = 3) and RISP KO (n = 3) mice were isolated on different days. Further processing and sequencing was performed with all six samples simultaneously. In f–h, CD4+Foxp3–YFP+CD25+ cells from RISP chimeric wild-type (n = 4) and RISP chimeric KO (n = 4) mice were isolated on different days. Further processing and sequencing was performed with all eight samples together.
Extended Data Fig. 9 Metabolite alterations in complex III-deficient Treg cells.
a, NAD+/NADH ratio in Treg (CD4+Foxp3–GFP+tdTomato–RFP+) cells isolated from QPC wild-type (n = 5) or QPC iKO (n = 6) mice six weeks after receiving three doses of tamoxifen. b, Calculated intracellular concentration of 2-HG, succinate, α-KG and fumarate in Treg (CD4+Foxp3–GFP+tdTomato–RFP+) cells isolated from QPC wild-type (n = 5) or QPC iKO (n = 5) six weeks after receiving three doses of tamoxifen. c, d, Specific complex III-dependent (c) and TCA cycle (d) metabolites in CD4+Foxp3–YFP+CD25+ cells isolated from three-week-old RISP KO and RISP wild-type mice, quantified by LC–MS. Data are mean ± s.d.; two-tailed t-test (a, b;**P < 0.01; P values in Source Data); multiple two-tailed t-tests using a two-stage linear step-up procedure of Benjamini, Krieger and Yekutieli, with Q = 0.01 (c, d). Each metabolite was analysed individually, without assuming a consistent s.d. (*Q < 0.01, **Q < 0.001, Q values in Source Data). All data points on graphs represent individual mice isolated and analysed on at least two separate days.
Extended Data Fig. 10 Elevated glycolytic flux does not phenocopy complex III inhibition in regulatory T cells.
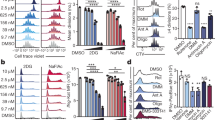
a, Venn diagram displaying overlap of differentially expressed genes (adjusted P < 0.01) from Treg cells (CD4+Foxp3–YFP+CD25+) isolated from 8–12-week-old RISP chimeric KO and RISP chimeric wild-type mice versus Treg cells overexpressing GLUT1 isolated from adult mice. P values were calculated using hypergeometric similarity measure. b, OCR of Treg (CD4+Foxp3–YFP+CD25+) cells treated with 1 µM piericidin, 500 µM 3-NPA or 1 µM antimycin A for 4 h. Data are mean ± s.d.; one-way ANOVA with Dunnett’s test for multiple comparisons; *adjusted P < 0.05, **adjusted P < 0.01, P values in Source Data. All data points on graphs represent individual mice isolated and analysed on at least two separate days.
Supplementary information
Supplementary Figure
This file contains uncropped images with size marker indications.
Source data
Rights and permissions
About this article
Cite this article
Weinberg, S.E., Singer, B.D., Steinert, E.M. et al. Mitochondrial complex III is essential for suppressive function of regulatory T cells. Nature 565, 495–499 (2019). https://doi.org/10.1038/s41586-018-0846-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0846-z
This article is cited by
-
Lipid metabolism in tumor-infiltrating regulatory T cells: perspective to precision immunotherapy
Biomarker Research (2024)
-
Ferritin heavy chain supports stability and function of the regulatory T cell lineage
The EMBO Journal (2024)
-
Identification of molecular subgroups and establishment of risk model based on the response to oxidative stress to predict overall survival of patients with lung adenocarcinoma
European Journal of Medical Research (2023)
-
A Treg-specific long noncoding RNA maintains immune-metabolic homeostasis in aging liver
Nature Aging (2023)
-
Metabolic landscape in cardiac aging: insights into molecular biology and therapeutic implications
Signal Transduction and Targeted Therapy (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.