Abstract
The genome is partitioned into topologically associated domains and genomic compartments with shared chromatin valence. This architecture is constrained by the DNA polymer, which precludes interactions between genes on different chromosomes. Here we report a marked divergence from this pattern of nuclear organization that occurs in mouse olfactory sensory neurons. Chromatin conformation capture using in situ Hi-C on fluorescence-activated cell-sorted olfactory sensory neurons and their progenitors shows that olfactory receptor gene clusters from 18 chromosomes make specific and robust interchromosomal contacts that increase with differentiation of the cells. These contacts are orchestrated by intergenic olfactory receptor enhancers, the ‘Greek islands’, which first contribute to the formation of olfactory receptor compartments and then form a multi-chromosomal super-enhancer that associates with the single active olfactory receptor gene. The Greek-island-bound transcription factor LHX2 and adaptor protein LDB1 regulate the assembly and maintenance of olfactory receptor compartments, Greek island hubs and olfactory receptor transcription, providing mechanistic insights into and functional support for the role of trans interactions in gene expression.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All Figures include publicly available data. All ChIP–seq and RNA-seq data reported in this paper (see Supplementary Information 1, 2) are available from GEO under accession number GSE112153. Additional data (mOSN RNA-seq, mOSN LHX2 ChIP–seq, mOSN EBF ChIP–seq and OLFR1507+ ATAC-seq) were previously described7 and are available from GEO under accession number GSE93570. All Hi-C data generated in this study are publicly available at https://data.4dnucleome.org/ under the following accession numbers: 4DNESH4UTRNL, 4DNESNYBDSLY, 4DNES54YB6TQ, 4DNESRE7AK5U, 4DNES425UDGS and 4DNESEPDL6KY.
References
Buck, L. & Axel, R. A novel multigene family may encode odorant receptors: a molecular basis for odor recognition. Cell 65, 175–187 (1991).
Magklara, A. et al. An epigenetic signature for monoallelic olfactory receptor expression. Cell 145, 555–570 (2011).
Chess, A., Simon, I., Cedar, H. & Axel, R. Allelic inactivation regulates olfactory receptor gene expression. Cell 78, 823–834 (1994).
Monahan, K. & Lomvardas, S. Monoallelic expression of olfactory receptors. Annu. Rev. Cell Dev. Biol. 31, 721–740 (2015).
Lyons, D. B. et al. An epigenetic trap stabilizes singular olfactory receptor expression. Cell 154, 325–336 (2013).
Markenscoff-Papadimitriou, E. et al. Enhancer interaction networks as a means for singular olfactory receptor expression. Cell 159, 543–557 (2014).
Monahan, K. et al. Cooperative interactions enable singular olfactory receptor expression in mouse olfactory neurons. eLife 6, e28620 (2017).
Clowney, E. J. et al. Nuclear aggregation of olfactory receptor genes governs their monogenic expression. Cell 151, 724–737 (2012).
Armelin-Correa, L. M., Gutiyama, L. M., Brandt, D. Y. & Malnic, B. Nuclear compartmentalization of odorant receptor genes. Proc. Natl Acad. Sci. USA 111, 2782–2787 (2014).
Spilianakis, C. G. & Flavell, R. A. Molecular biology. Managing associations between different chromosomes. Science 312, 207–208 (2006).
Beagrie, R. A. et al. Complex multi-enhancer contacts captured by genome architecture mapping. Nature 543, 519–524 (2017).
Apostolou, E. & Thanos, D. Virus infection induces NF-κB-dependent interchromosomal associations mediating monoallelic IFN-β gene expression. Cell 134, 85–96 (2008).
Maass, P. G., Barutcu, A. R., Weiner, C. L. & Rinn, J. L. Inter-chromosomal contact properties in live-cell imaging and in Hi-C. Mol. Cell 70, 188–189 (2018).
Maass, P. G., Barutcu, A. R. & Rinn, J. L. Interchromosomal interactions: a genomic love story of kissing chromosomes. J. Cell Biol. jcb.201806052 (2018).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Nagano, T. et al. Comparison of Hi-C results using in-solution versus in-nucleus ligation. Genome Biol. 16, 175 (2015).
Johanson, T. M. et al. Genome-wide analysis reveals no evidence of trans chromosomal regulation of mammalian immune development. PLoS Genet. 14, e1007431 (2018).
Fuss, S. H., Omura, M. & Mombaerts, P. Local and cis effects of the H element on expression of odorant receptor genes in mouse. Cell 130, 373–384 (2007).
Le Gros, M. A. et al. Soft X-ray tomography reveals gradual chromatin compaction and reorganization during neurogenesis in vivo. Cell Reports 17, 2125–2136 (2016).
Hanchate, N. K. et al. Single-cell transcriptomics reveals receptor transformations during olfactory neurogenesis. Science 350, 1251–1255 (2015).
Saraiva, L. R. et al. Hierarchical deconstruction of mouse olfactory sensory neurons: from whole mucosa to single-cell RNA-seq. Sci. Rep. 5, 18178 (2015).
Tan, L., Li, Q. & Xie, X. S. Olfactory sensory neurons transiently express multiple olfactory receptors during development. Mol. Syst. Biol. 11, 844 (2015).
Ahmed, S. et al. DNA zip codes control an ancient mechanism for gene targeting to the nuclear periphery. Nat. Cell Biol. 12, 111–118 (2010).
Hewitt, S. L. et al. Association between the Igk and Igh immunoglobulin loci mediated by the 3′ Igk enhancer induces ‘decontraction’ of the Igh locus in pre-B cells. Nat. Immunol. 9, 396–404 (2008).
Gadye, L. et al. Injury activates transient olfactory stem cell states with diverse lineage capacities. Cell Stem Cell 21, 775–790 (2017).
Lin, B. et al. Injury induces endogenous reprogramming and dedifferentiation of neuronal progenitors to multipotency. Cell Stem Cell 21, 761–774 (2017).
Agulnick, A. D. et al. Interactions of the LIM-domain-binding factor Ldb1 with LIM homeodomain proteins. Nature 384, 270–272 (1996).
Bach, I. The LIM domain: regulation by association. Mech. Dev. 91, 5–17 (2000).
Krivega, I. & Dean, A. LDB1-mediated enhancer looping can be established independent of mediator and cohesin. Nucleic Acids Res. 45, 8255–8268 (2017).
Lee, J., Krivega, I., Dale, R. K. & Dean, A. The LDB1 complex co-opts CTCF for erythroid lineage-specific long-range enhancer interactions. Cell Reports 19, 2490–2502 (2017).
Deng, W. et al. Controlling long-range genomic interactions at a native locus by targeted tethering of a looping factor. Cell 149, 1233–1244 (2012).
Caputo, L. et al. The Isl1/Ldb1 complex orchestrates genome-wide chromatin organization to instruct differentiation of multipotent cardiac progenitors. Cell Stem Cell 17, 287–299 (2015).
Bronstein, R. et al. Transcriptional regulation by CHIP/LDB complexes. PLoS Genet. 6, e1001063 (2010).
Matthews, J. M. & Visvader, J. E. LIM-domain-binding protein 1: a multifunctional cofactor that interacts with diverse proteins. EMBO Rep. 4, 1132–1137 (2003).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320 (2017).
Schwarzer, W. et al. Two independent modes of chromatin organization revealed by cohesin removal. Nature 551, 51–56 (2017).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Pederson, T. The nucleolus. Cold Spring Harb. Perspect. Biol. 3, a000638 (2011).
Larson, A. G. et al. Liquid droplet formation by HP1α suggests a role for phase separation in heterochromatin. Nature 547, 236–240 (2017).
Strom, A. R. et al. Phase separation drives heterochromatin domain formation. Nature 547, 241–245 (2017).
Hnisz, D., Shrinivas, K., Young, R. A., Chakraborty, A. K. & Sharp, P. A. A phase separation model for transcriptional control. Cell 169, 13–23 (2017).
Sabari, B. R. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 361, aar3958 (2018).
Lomvardas, S. et al. Interchromosomal interactions and olfactory receptor choice. Cell 126, 403–413 (2006).
Khan, M., Vaes, E. & Mombaerts, P. Regulation of the probability of mouse odorant receptor gene choice. Cell 147, 907–921 (2011).
Nishizumi, H., Kumasaka, K., Inoue, N., Nakashima, A. & Sakano, H. Deletion of the core-H region in mice abolishes the expression of three proximal odorant receptor genes in cis. Proc. Natl Acad. Sci. USA 104, 20067–20072 (2007).
Guo, Y. et al. CRISPR inversion of CTCF sites alters genome topology and enhancer/promoter function. Cell 162, 900–910 (2015).
Noordermeer, D. et al. Variegated gene expression caused by cell-specific long-range DNA interactions. Nat. Cell Biol. 13, 944–951 (2011).
Shykind, B. M. et al. Gene switching and the stability of odorant receptor gene choice. Cell 117, 801–815 (2004).
Rock, J. R. et al. Basal cells as stem cells of the mouse trachea and human airway epithelium. Proc. Natl Acad. Sci. USA 106, 12771–12775 (2009).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Vassalli, A., Rothman, A., Feinstein, P., Zapotocky, M. & Mombaerts, P. Minigenes impart odorant receptor-specific axon guidance in the olfactory bulb. Neuron 35, 681–696 (2002).
Eggan, K. et al. Mice cloned from olfactory sensory neurons. Nature 428, 44–49 (2004).
Mangale, V. S. et al. Lhx2 selector activity specifies cortical identity and suppresses hippocampal organizer fate. Science 319, 304–309 (2008).
Zhao, Y. et al. LIM-homeodomain proteins Lhx1 and Lhx5, and their cofactor Ldb1, control Purkinje cell differentiation in the developing cerebellum. Proc. Natl Acad. Sci. USA 104, 13182–13186 (2007).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3, 95–98 (2016).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Droettboom, M. matplotlib/matplotlib v2.2.2 https://zenodo.org/record/1202077 (2018).
Waskom, M. mwaskom/seaborn: v0.8.1 https://zenodo.org/record/883859 (2017).
McKinney, W. Data Structures for Statistical Computing in Python. Proc. 9th Python Sci. Conf. 1697900, 51–56 (2010).
Freese, N. H., Norris, D. C. & Loraine, A. E. Integrated genome browser: visual analytics platform for genomics. Bioinformatics 32, 2089–2095 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Klein, F. A. et al. FourCSeq: analysis of 4C sequencing data. Bioinformatics 31, 3085–3091 (2015).
Pedregosa, F. Scikit-learn: machine learning in python. J. Mach. Learn. Res. 12, 2825–2830 (2012).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Ross-Innes, C. S. et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481, 389–393 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Benjamini, Y. & Speed, T. P. Summarizing and correcting the GC content bias in high-throughput sequencing. Nucleic Acids Res. 40, e72 (2012).
Yan, J. et al. Histone H3 lysine 4 monomethylation modulates long-range chromatin interactions at enhancers. Cell Res. 28, 387 (2018).
Bonev, B. et al. Multiscale 3D genome rewiring during mouse neural development. Cell 171, 557–572 (2017).
Acknowledgements
We thank A. Mumbay-Wafula for assistance with mouse colony management, I. Schieren for assistance with fluorescence-activated cell sorting, P. Love for sharing the Ldb1fl/fl mice and E. Monuki for sharing the Lhx2fl/fl mice. A.H. was funded by F31 post-doctoral fellowship DC016785 (NIH) and K.M. was funded by F32 post-doctoral fellowship GM108474 (NIH). S.L. acknowledges support from the NIH Common Fund 4D Nucleome Program (grant 1U01DA040582). In addition, this project was funded by R01DC013560, R01DC015451 (NIH) and the HHMI Faculty Scholar Award. Research reported in this publication was also performed in the CCTI Flow Cytometry Core, supported in part by the Office of the Director, National Institutes of Health under award S10OD020056. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Author information
Authors and Affiliations
Contributions
K.M, A.H., and S.L. designed the study. K.M. performed in situ Hi-C in Lhx2 and Ldb1 knockout mice, performed ChIP–seq in wild-type and Lhx2 knockout mOSNs, and performed RNA-seq in Ldb1 knockout and control mOSNs and methimazole-treated cells from the MOE. A.H. performed in situ Hi-C in mOSNs, INPs, HBCs, and OLFR1507, OLFR16, and OLFR17-expressing cells. Both K.M and A.H. analysed data with input from S.L. S.L. wrote the manuscript with input from K.M. and A.H.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Hi-C on FAC-sorted primary cells from the MOE reveals extensive interchromosomal interactions between OR clusters.
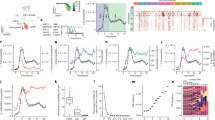
a, Table summarizing all Hi-C experiments in this manuscript separated by biological replicates. The total number of Hi-C contacts in each replicate and the total number of interchromosomal (trans) Hi-C contacts are shown. b–d, Hi-C contact curves for wild-type conditions (b), for wild-type and mutant MOE populations (c), and for cells sorted on the basis of expression of specific OR genes (d). All panels present pooled data from two independent biological replicates that yielded similar results when analysed separately. e, Graphs showing the proportion of trans Hi-C contacts between replicates of each genotype and cell type. Pooled data from publicly available datasets is shown for ES cells72, B cells15 and cortical neurons73. f, As in e, but showing the median fraction of Hi-C contacts made to trans OR clusters for OR cluster regions divided into 50-kb bins. g, Machine-learning HMM score for a given number of compartments (see Methods). Nine compartments were used for further analysis. h, From the nine HMM-derived compartments, one includes predominantly OR clusters (magenta, bottom) and overlaps with OR compartments defined by biased analysis of trans OR contacts (black, top). OR gene clusters depicted in red. Scale on the biased analysis represents the percentage of Hi-C contacts mapped to trans OR clusters (pooled data from two biological replicates). Scale in the HMM-derived compartments represents the average value of a given locus in a given compartment. i, Circos plots depicting the strongest 1,000 interchromosomal interactions genome-wide at 1-Mb resolution in mOSNs. Red lines represent OR-to-OR contacts and black lines non-OR-to-non-OR contacts. Line thickness increases with contact frequency. Chromosome numbers given at the periphery of the circle.
Extended Data Fig. 2 Extensive interchromosomal contacts form between OR gene clusters over OSN differentiation.
a–i, In situ Hi-C contact matrices of chromosomes 2 and 9, APAs, and Circos plots depicting the strongest 1,000 interchromosomal interactions genome-wide for mOSNs (a–c), INPs (d–f) and HBCs (g–i). All three sets of analyses reveal an increase in trans OR cluster interactions over the course of differentiation. j, For OR gene clusters (divided into 50-kb bins, n = 768 bins) the frequency of cis short (<5 Mb distance, including self), cis long (>5 Mb) and trans contacts with OR clusters is shown, expressed as the fraction of total Hi-C contacts mapped to each bin. k, Number of Hi-C contacts, normalized to a library size of one billion Hi-C contacts genome-wide, observed for each OR cluster region (divided into 50-kb bins, n = 768 bins) in HBCs, INPs, mOSNs, ES cells, B cells and cortical neurons. l–n, For OR cluster regions (divided into 50-kb bins, n = 768 bins), the fraction of total Hi-C contacts that are made to ORs clusters located in short range cis (l), long range cis (m) and trans (n). o, The six most distinct HMM-derived compartments of chromosome 2 in HBCs (green, left), INPs (blue, middle) and mOSNs (magenta, right). OR clusters emerge as distinct compartment in INPs and strengthen in mOSNs. For all box plots, box indicates median, upper, and lower quartiles; whiskers indicate 1.5 × the interquartile range. All panels present pooled data from two independent biological replicates that yielded similar results when analysed separately.
Extended Data Fig. 3 In vitro and in silico Hi-C experiments show that OR Hi-C contacts are generated by unique sequences that do not map to other OR clusters.
a, Contact matrices from in vitro Hi-C (top) using a 165-kb BAC plasmid containing seven OR genes from an OR cluster from chromosome 1 and in situ Hi-C from mOSNs (bottom). Hi-C contacts in the BAC Hi-C are restricted to the coordinates of the BAC plasmid and do not extend to two OR genes from this cluster that are absent from the BAC. b, Virtual 4C from the 165-kb BAC region to chromosome 2, which contains the highest number of OR genes. Top, virtual 4C from the BAC in vitro Hi-C shows that no reads mapped to ORs from chromosome 2, whereas the same 165-kb regions makes abundant trans contacts with these ORs in mOSNs. c, Of all the BAC Hi-C contacts, 99.3% map within the BAC, whereas in mOSNs only 21.7% of the BAC region Hi-C contacts map within the BAC. d, In silico Hi-C analysis shows complete absence of mis-mapped reads corresponding to OR clusters under the mapping conditions used throughout the manuscript (removing mapq < 30). Each OR cluster was subjected to intracluster in silico Hi-C (g) and then the Hi-C contacts of the 69 OR clusters were mapped in aggregate to the whole genome. As seen in the contact matrix from chromosomes 2 and 9 (d), the in silico reads only map within clusters, with no mis-mapped reads that would erroneously be interpreted as intercluster cis or trans contacts. e, For reference, the corresponding in situ Hi-C from mOSNs. f, Aggregate analysis for all 69 OR gene clusters shows that our mapping protocol does not mis-map any Hi-C contacts to the wrong OR cluster. g, Brief description of the pipeline used for the in silico analysis.
Extended Data Fig. 4 Greek islands make differentiation-dependent contacts with other Greek islands in trans that are stronger than cis contacts with LHX2 and EBF peaks.
a, b, Heat maps and 3D projections of Hi-C contacts between a pair of OR gene clusters in cis (a) and trans (b) reveal a local maximum of in situ Hi-C interactions between Greek island loci (arrowheads) in mOSNs. c–f, As in a, b but for INPs and HBCs. g, For chromosome 2, fraction of all Hi-C contacts made to trans Greek islands in mOSNs (top), INPs (middle) and HBCs (bottom). h, For each Greek island, the distribution of Hi-C contacts—expressed as contacts per billion—made to individual Greek islands located in trans for HBCs, INPs and mOSNs. Boxes indicate median, upper, and lower quartiles; whiskers indicate 1.5 × the interquartile range. For each Greek island, the number of trans Greek islands is listed. i, Left, comparison of the total fraction of Hi-C contacts made by each Greek island to intergenic LHX2- and EBF-bound peaks present in cis versus Greek islands present in trans for HBCs, INPs and mOSNs. For each category, we compare roughly equal numbers of peaks (number of trans Greek islands for each island versus number of cis LHX2/EBF sites for each island, mean ± s.d.). Right, mean fraction of Hi-C contacts across all Greek islands (two-sided, paired Wilcoxon signed-rank test, n = 59). Contacts with trans Greek islands (red) constitute a higher fraction of Hi-C contacts than short-range cis (dark blue) or long-range cis (light blue) contacts with intergenic LHX2/EBF peaks. All panels present pooled data from two independent biological replicates that yielded similar results when analysed separately.
Extended Data Fig. 5 Greek islands and LHX2 are required for OR compartmentalization in developing OSNs.
a, Pairwise Hi-C contacts between all pairs of Greek islands ordered by genomic position in control (left) and Greek island triple KO (right) mOSNs. The 50-kb regions containing the deleted Greek islands are marked with arrowheads. Plotting the log2[fold difference] in Hi-C contacts (right) reveals that consistent strong reductions are observed for the deleted islands. Colour bar depicts chromosome. b, c, The genomic regions exhibiting the most significant reductions in Hi-C contacts with trans OR Greek islands (b) or trans OR clusters (c), in triple KO mOSNs relative to control mOSNs, are mostly located within the three OR clusters containing the deleted Greek islands (two biological replicates per condition, see Methods). d, Genetic and experimental strategy for early Lhx2 deletion. Tamoxifen induction with Krt5-creER deletes Lhx2 in HBCs and then methimazole treatment ablates INPs and mOSNs, leading to regeneration from Lhx2-deleted HBCs. e, Fluorescent labelling of the HBC-derived cells upon methimazole induction reveals two major populations, bright and dim. f, RNA-seq shows that the dim population expresses markers of INPs and mOSNs; the bright population expresses markers of HBCs. Counts are normalized by row. g, Three-dimensional projection of Hi-C contacts between OR clusters located on different chromosomes in control mOSNs, INPs, early Lhx2 KO cells and late Lhx2 KO cells (from left to right). A Hi-C hotspot between interacting Greek islands is observed only in control mOSNs (arrowhead). In addition, a strong reduction in the surrounding OR–OR contacts relative to mOSNs or INPs is observed in the early Lhx2 KO cells. h, Circos plots depicting the strongest 1,000 interchromosomal interactions genome-wide at 1-Mb resolution in (left to right) mOSNs, INPs, early Lhx2 KO cells, and late Lhx2 KO cells. Red lines represent OR–OR contacts and black lines non-OR–non-OR contacts. Line thickness increases with contact frequency. Chromosome numbers depicted at the periphery of the circle. i, Genetic strategy for late Lhx2 deletion and fluorescent marking of Lhx2 KO mOSNs. j, Left, for each Greek island, the fraction of total Hi-C contacts made to other Greek islands located in cis at short range (<5 Mb apart, grey), long range (>5 Mb apart, blue), and in trans (red). Top, control mOSNs; bottom, late Lhx2 KO cells. Right, the effect of late Lhx2 KO on the mean fraction of Hi-C contacts across all Greek islands (two-sided, paired Wilcoxon signed-rank test, n = 59).
Extended Data Fig. 6 LDB1 expression and genomic distribution in mOSNs.
a, Transcript level, expressed as fragments per kilobase per million mapped reads (FPKM), of the two LDB family members in mOSN RNA-seq datasets (n = 5 biological replicates). b, Sections of olfactory epithelium stained for LDB1 (green) and ADCY3 (magenta), a marker for mOSNs. Nuclei are labelled with DAPI (blue). Scale bar, 25 μm. Similar results were obtained from four independent experiments. c, LDB1 and LHX2 ChIP–seq signal in mOSNs across the OR gene cluster containing the Greek island Lipsi. OR genes are red and all other genes are blue. Plot shows pooled data from two biological replicates for LHX2 and three biological replicates for LDB1, each of which yielded similar results when analysed separately. Values are counts per 10 million reads. d, Extensive overlap between consensus LHX2 and LDB1 ChIP–seq peak sets. e, Linear relationship between normalized LHX2 ChIP signal and LDB1 ChIP signal. Any peak observed in at least two of the five experiments (two for LHX2 and three for LDB1) was included (n = 26,667) and plotted together with a best fit line obtained by linear regression with y-intercept set to 0. f, LDB1 ChIP signal over Greek islands in mOSNs and late Lhx2 KO mOSNs. Heat map shows pooled data from three biological replicates for mOSNs and two biological replicates for late Lhx2 KO cells, each of which yielded similar results when analysed separately. Values are counts per 10 million reads. g, Normalized LDB1 ChIP–seq signal is greater for LDB1 peaks that overlap Greek islands than for peaks that do not (P < 2.2 × 10–16, two-sided Wilcoxon rank sum test, n = 63 for Greek islands, n = 22,728 for non-island peaks). Violin plots are scaled to the same area and show density for the full set of points over the full range. h, Normalized LDB1 ChIP–seq signal is greater for LDB1 peaks that overlap EBF ChIP peaks than for peaks that do not (P < 2.2 × 10–16, two-sided Wilcoxon rank sum test, n = 8,779 for LDB1 peaks that overlap EBF peaks, n = 14,012 for non-EBF peaks). Violin plots are scaled to the same area and show density for the full set of points over the full range. i, mOSN ChIP–seq for LDB1, CTCF and RAD21 across the OR gene cluster containing the Greek island Lipsi. OR genes are red and all other genes are blue. Plot shows pooled data from three biological replicates for LDB1 and two biological replicates for CTCF and RAD21. Values are counts per 10 million reads. Analysing each replicate separately yielded similar results. j, mOSN ChIP signal over Greek island and non-Greek island ChIP–seq peaks. For ChIP–seq peaks, the heat map shows 500 randomly selected peaks and the plot shows data from the full consensus set of peaks (n = 22,791 for LDB1, n = 24,883 for CTCF, and n = 9,882 for RAD21). Plots show pooled data; similar results were obtained with each replicate (n = 3 for LDB1 ChIP–seq and n = 2 for CTCF and RAD21 ChIP–seq). Units are counts per 10 million reads. k, As in j, but showing LDB1 ChIP signal over OR genes (n = 1,255) in mOSNs. l, LDB1 ChIP–seq from control mOSNs (top) and OLFR1507-expressing cells (middle). Strong signal is observed on the Greek island H, in both populations, but only a very weak signal on the Olfr1507 promoter when it is transcriptionally engaged. Pooled data from three biological replicates is shown for the mOSNs. One of two biological replicates is shown for OLFR1507+ OSNs; the other replicate yielded similar results but with lower enrichment in peaks. ATAC-seq from the OLFR1507-expressing cells (bottom) shows that the promoter of Olfr1507 has similar accessibility to the H element. ATAC-seq plot shows pooled data from two biological replicates that yielded similar results.
Extended Data Fig. 7 Effects of conditional Ldb1 deletion on Greek island interactions and OR expression.
a, Schematic of the genetic strategy used to generate Ldb1 KO mOSNs that are fluorescently labelled. b, In Ldb1fl/fl;OMP-cre mice, LDB1 (green) is lost from mOSNs but retained in basal immature cells. Nuclei are stained with DAPI (magenta). Scale bar, 20 μm. Similar results were obtained from three independent experiments. c, Hi-C contacts between a pair of OR clusters located on different chromosomes in control (top) and Ldb1 KO (bottom) mOSNs. A Hi-C hotspot between interacting Greek islands in control mOSNs (arrowheads) is absent in Ldb1 KO OSNs. d, Three-dimensional projection of the same OR cluster pair in control and Ldb1 KO OSNs. e, Trans interactions of each Greek island (n = 59) with the other Greek islands as fraction of the total Hi-C contacts in mOSNs versus Ldb1 KO cells. Greek islands that changed more than twofold are red. f, For each Greek island, the mean number of cis long-range (left) and trans (right) Hi-C contacts per billion made to every non-OR sequence (at 50-kb resolution), intergenic LHX2- and EBF-bound peak (outside OR clusters), or Greek island. Boxes indicate median, upper and lower quartiles; whiskers indicate 1.5 × the interquartile range. g, As in e but for trans contacts between OR gene clusters (n = 67). Clusters that changed more than 1.5-fold are red. h, Circos plots depicting the strongest 1,000 interchromosomal interactions genome-wide at 1-Mb resolution in control mOSNs (left) and Ldb1 KO mOSNs (right). Red lines represent OR–OR contacts and black lines non-OR–non-OR contacts. Line thickness increases with contact frequency. Chromosome numbers depicted at the periphery of the circle. i, Transcript levels of Greek island-binding factors in RNA-seq data from control mOSNs and Ldb1 KO mOSNs. Transcript levels of Ebf3 are reduced approximately twofold (P = 0.031 for greater than 1.5-fold change, DESeq2 normalized Wald test with n = 5 for control mOSNs and n = 4 Ldb1 KO). The expression of other factors is not significantly different between conditions.
Extended Data Fig. 8 Long-range interactions in homogeneous OSN subpopulations.
a–c, Circos plots representing the 1,000 strongest trans contacts in OLFR16+ (a), OLFR17+ (b) and OLFR1507+ (c) OSNs. d, Left, comparison of the frequency of local cis (grey), long-range cis (blue) and trans (red) Greek island interactions in mixed mOSNs and OSNs expressing specific OR genes. Right, mean values for OLFR16+, OLFR17+ and OLFR1507+ cells are not significantly different from those for mixed mOSNs (P > 0.05 for all comparisons, two-tailed paired Wilcoxon signed-rank test). e, In situ Hi-C contact matrices from OLFR16+, OLFR17+ and OLFR1507+ cells focused on the Olfr16 gene locus. Arrowhead points to specific long-range contacts between Olfr16 and the Greek island Astypalea that occur only in OLFR16+ cells. Grey arrowheads mark Greek island–Greek island contacts that also differ between cell types. f, g, Similar analysis for the Olfr16 locus in OLFR17+ and OLFR1507+ cells. h–j, As in e–g, but for the Olfr17 locus. k–m, As in e–g, but for the Olfr1507 locus.
Extended Data Fig. 9 Long-range cis and trans contacts between Greek islands and the active OR gene.
a, Hi-C contacts that span more than 80 Mb are observed between the Olfr16 locus and Greek islands in OLFR16+ cells. b, Close examination of the contacts (dashed box from a) reveals that Greek islands contact Olfr16 only in OLFR16+ cells (top, black arrowhead). Extremely long-range contacts between Greek islands (grey arrowheads), but not involving the Olfr16 locus, are also observed in OLFR17+ and OLFR1507+ cells (middle, bottom). c, Heat maps depicting interchromosomal contacts between Olfr16 (chromosome 1) and Greek islands from different chromosomes in in situ Hi-C from OLFR16+, OLFR17+ and OLF1507+ cells. d, Three-dimensional projection of APA between the Olfr16 locus and trans Greek islands in the three specific mOSN populations. e, Heat maps for contacts between Olfr16, Olfr17, or Olfr1507 and trans Greek islands reveals an accumulation of contacts centred around the active allele. f, APA for OR versus trans Greek islands shows the accumulation of contacts on the active allele at 10-kb resolution. The poor mappability of the Olfr17 locus and the lower sequencing depth perturbs the expected focal peak. For the Olfr1507 locus, the presence of the Greek island H, 50 kb from Olfr1507, results in Hi-C contacts spanning a broad area. g, h, Short, long and trans contacts with Greek islands across the OR gene clusters containing Olfr17 (g) and Olfr1507 (h) plotted as a fraction of the total Hi-C contacts mapped to each position (5-kb resolution). Top, contacts in cells in which Olfr17 and Olfr1507 are active; bottom, contacts in OLFR16+ cells in which Olfr17 and Olfr1507 are silent.
Extended Data Fig. 10 A model for specific OR compartmentalization and the generation of mutually exclusive phases that regulate OR gene choice.
a, b, Coincidence of LHX2 and LDB1 peaks with trimethylation of histone 3 lysine 9 (H3K9me3) enrichment may generate an OR-enriched molecular barcode that promotes specific interactions between OR gene clusters. c, In INPs, where OR compartments first form, Greek islands do not make specific contacts with each other. d, However, in mOSNs Greek islands specifically interact with each other through homotypic LDB1 interactions, forming a multi-enhancer hub that is segregated from the OR compartment. We hypothesize that OR compartments and Greek island hubs form incompatible liquid phases driven by HP1 proteins and the unstructured domains of LHX2 and LDB1, respectively. e, Upon deletion of Ldb1 (or Lhx2), the Greek island phase falls apart and the Greek islands become incorporated to the OR compartments, as in the INPs.
Supplementary information
Supplementary Information 1
This file contains a summary of ChIP-seq datasets. Metadata, sequencing statistics, and peak calling summary for ChIP-seq datasets presented in this manuscript.
Supplementary Information 2
This file contains a summary of RNA-seq datasets. Metadata and sequencing statistics for RNA-seq datasets presented in this manuscript.
Rights and permissions
About this article
Cite this article
Monahan, K., Horta, A. & Lomvardas, S. LHX2- and LDB1-mediated trans interactions regulate olfactory receptor choice. Nature 565, 448–453 (2019). https://doi.org/10.1038/s41586-018-0845-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0845-0
This article is cited by
-
RNA-mediated symmetry breaking enables singular olfactory receptor choice
Nature (2024)
-
Enhancer contacts during embryonic development show diverse interaction modes and modest yet significant increases upon gene activation
Nature Genetics (2024)
-
Simultaneous single-cell three-dimensional genome and gene expression profiling uncovers dynamic enhancer connectivity underlying olfactory receptor choice
Nature Methods (2024)
-
p53 rapidly restructures 3D chromatin organization to trigger a transcriptional response
Nature Communications (2024)
-
LiMCA: Hi-C gets an RNA twist
Nature Methods (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.