Abstract
Wood, a type of xylem tissue, originates from cell proliferation of the vascular cambium. Xylem is produced inside, and phloem outside, of the cambium1. Morphogenesis in plants is typically coordinated by organizer cells that direct the adjacent stem cells to undergo programmed cell division and differentiation. The location of the vascular cambium stem cells and whether the organizer concept applies to the cambium are currently unknown2. Here, using lineage-tracing and molecular genetic studies in the roots of Arabidopsis thaliana, we show that cells with a xylem identity direct adjacent vascular cambial cells to divide and function as stem cells. Thus, these xylem-identity cells constitute an organizer. A local maximum of the phytohormone auxin, and consequent expression of CLASS III HOMEODOMAIN-LEUCINE ZIPPER (HD-ZIP III) transcription factors, promotes xylem identity and cellular quiescence of the organizer cells. Additionally, the organizer maintains phloem identity in a non-cell-autonomous fashion. Consistent with this dual function of the organizer cells, xylem and phloem originate from a single, bifacial stem cell in each radial cell file, which confirms the classical theory of a uniseriate vascular cambium3. Clones that display high levels of ectopically activated auxin signalling differentiate as xylem vessels; these clones induce cell divisions and the expression of cambial and phloem markers in the adjacent cells, which suggests that a local auxin-signalling maximum is sufficient to specify a stem-cell organizer. Although vascular cambium has a unique function among plant meristems, the stem-cell organizer of this tissue shares features with the organizers of root and shoot meristems.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Gene accession numbers of all the Arabidopsis genes analysed in this study are listed above. Source Data related to Fig. 2b, c, 3a, b, d and Extended Data Figs. 3d, 4a, c, d, 6a–e, 7b, f, g, 8a–e, 9f are provided with the paper. All lines and other data supporting the findings of this study are available from the corresponding author upon request.
References
Larson, P. R. The Vascular Cambium: Development and Structure (Springer, Berlin, 1994).
Greb, T. & Lohmann, J. U. Plant stem cells. Curr. Biol. 26, R816–R821 (2016).
Sanio, C. Anatomie der gemeinen Kiefer (Pinus silvestris L.) ll. Entwickelungsgeschichte der Holzzellen. Jahrb. Wiss. Bot. 9, 50–128 (1873).
Evert, R. F. & Eichhorn, S. E. Esau’s plant anatomy: Meristems, Cells, and Tissues of the Plant Body—their Structure, Function and Development, 3rd edn (John Wiley & Sons, Hoboken, 2006).
Chaffey, N., Cholewa, E., Regan, S. & Sundberg, B. Secondary xylem development in Arabidopsis: a model for wood formation. Physiol. Plant. 114, 594–600 (2002).
Wunderling, A. et al. A molecular framework to study periderm formation in Arabidopsis. New Phytol. 219, 216–229 (2018).
Atta, R. et al. Pluripotency of Arabidopsis xylem pericycle underlies shoot regeneration from root and hypocotyl explants grown in vitro. Plant J. 57, 626–644 (2009).
Carlsbecker, A. et al. Cell signalling by microRNA165/6 directs gene dose-dependent root cell fate. Nature 465, 316–321 (2010).
Ohashi-Ito, K. & Fukuda, H. HD-Zip III homeobox genes that include a novel member, ZeHB-13 (Zinnia)/ATHB-15 (Arabidopsis), are involved in procambium and xylem cell differentiation. Plant Cell Physiol. 44, 1350–1358 (2003).
Baima, S. et al. The Arabidopsis ATHB-8 HD-zip protein acts as a differentiation-promoting transcription factor of the vascular meristems. Plant Physiol. 126, 643–655 (2001).
Robischon, M., Du, J., Miura, E. & Groover, A. The Populus class III HD ZIP, popREVOLUTA, influences cambium initiation and patterning of woody stems. Plant Physiol. 155, 1214–1225 (2011).
Siligato, R. et al. MultiSite gateway-compatible cell type-specific gene-inducible system for plants. Plant Physiol. 170, 627–641 (2016).
Mallory, A. C. et al. MicroRNA control of PHABULOSA in leaf development: importance of pairing to the microRNA 5′ region. EMBO J. 23, 3356–3364 (2004).
Hirakawa, Y., Kondo, Y. & Fukuda, H. TDIF peptide signaling regulates vascular stem cell proliferation via the WOX4 homeobox gene in Arabidopsis. Plant Cell 22, 2618–2629 (2010).
Fisher, K. & Turner, S. PXY, a receptor-like kinase essential for maintaining polarity during plant vascular-tissue development. Curr. Biol. 17, 1061–1066 (2007).
Lee, J. Y. et al. Transcriptional and posttranscriptional regulation of transcription factor expression in Arabidopsis roots. Proc. Natl Acad. Sci. USA 103, 6055–6060 (2006).
Miyashima, S. et al. Mobile PEAR transcription factors integrate positional cues to prime cambial growth. Nature https://doi.org/10.1038/s41586-018-0839-y (2019).
Ursache, R. et al. Tryptophan-dependent auxin biosynthesis is required for HD-ZIP III-mediated xylem patterning. Development 141, 1250–1259 (2014).
Donner, T. J., Sherr, I. & Scarpella, E. Regulation of preprocambial cell state acquisition by auxin signaling in Arabidopsis leaves. Development 136, 3235–3246 (2009).
Uggla, C., Moritz, T., Sandberg, G. & Sundberg, B. Auxin as a positional signal in pattern formation in plants. Proc. Natl Acad. Sci. USA 93, 9282–9286 (1996).
Immanen, J. et al. Cytokinin and auxin display distinct but interconnected distribution and signaling profiles to stimulate cambial activity. Curr. Biol. 26, 1990–1997 (2016).
Weijers, D. & Wagner, D. Transcriptional responses to the auxin hormone. Annu. Rev. Plant Biol. 67, 539–574 (2016).
Przemeck, G. K., Mattsson, J., Hardtke, C. S., Sung, Z. R. & Berleth, T. Studies on the role of the Arabidopsis gene MONOPTEROS in vascular development and plant cell axialization. Planta 200, 229–237 (1996).
Brackmann, K. et al. Spatial specificity of auxin responses coordinates wood formation. Nat. Commun. 9, 875 (2018).
Berleth, T. & Jurgens, G. The role of the monopteros gene in organising the basal body region of the Arabidopsis embryo. Development 118, 575–587 (1993).
Mähönen, A. P. et al. PLETHORA gradient formation mechanism separates auxin responses. Nature 515, 125–129 (2014).
Krogan, N. T., Ckurshumova, W., Marcos, D., Caragea, A. E. & Berleth, T. Deletion of MP/ARF5 domains III and IV reveals a requirement for Aux/IAA regulation in Arabidopsis leaf vascular patterning. New Phytol. 194, 391–401 (2012).
Kubo, M. et al. Transcription switches for protoxylem and metaxylem vessel formation. Genes Dev. 19, 1855–1860 (2005).
Sabatini, S. et al. An auxin-dependent distal organizer of pattern and polarity in the Arabidopsis root. Cell 99, 463–472 (1999).
Nieminen, K., Blomster, T., Helariutta, Y. & Mähönen, A. P. Vascular cambium development. Arabidopsis Book 13, e0177 (2015).
Emery, J. F. et al. Radial patterning of Arabidopsis shoots by class III HD-ZIP and KANADI genes. Curr. Biol. 13, 1768–1774 (2003).
Heidstra, R., Welch, D. & Scheres, B. Mosaic analyses using marked activation and deletion clones dissect Arabidopsis SCARECROW action in asymmetric cell division. Genes Dev. 18, 1964–1969 (2004).
Wu, M. F., Tian, Q. & Reed, J. W. Arabidopsis microRNA167 controls patterns of ARF6 and ARF8 expression, and regulates both female and male reproduction. Development 133, 4211–4218 (2006).
Karimi, M., Depicker, A. & Hilson, P. Recombinational cloning with plant gateway vectors. Plant Physiol. 145, 1144–1154 (2007).
Prigge, M. J. et al. Class III homeodomain-leucine zipper gene family members have overlapping, antagonistic, and distinct roles in Arabidopsis development. Plant Cell 17, 61–76 (2005).
Okushima, Y. et al. Functional genomic analysis of the AUXIN RESPONSE FACTOR gene family members in Arabidopsis thaliana: unique and overlapping functions of ARF7 and ARF19. Plant Cell 17, 444–463 (2005).
Sawchuk, M. G., Edgar, A. & Scarpella, E. Patterning of leaf vein networks by convergent auxin transport pathways. PLoS Genet. 9, e1003294 (2013).
Friml, J. et al. Efflux-dependent auxin gradients establish the apical–basal axis of Arabidopsis. Nature 426, 147–153 (2003).
Liao, C. Y. et al. Reporters for sensitive and quantitative measurement of auxin response. Nat. Methods 12, 207–210 (2015).
Baesso, B. et al. Transcription factors PRE3 and WOX11 are involved in the formation of new lateral roots from secondary growth taproot in A. thaliana. Plant Biol. 20, 426–432 (2018).
Suer, S., Agusti, J., Sanchez, P., Schwarz, M. & Greb, T. WOX4 imparts auxin responsiveness to cambium cells in Arabidopsis. Plant Cell 23, 3247–3259 (2011).
Vidaurre, D. P., Ploense, S., Krogan, N. T. & Berleth, T. AMP1 and MP antagonistically regulate embryo and meristem development in Arabidopsis. Development 134, 2561–2567 (2007).
Wilmoth, J. C. et al. NPH4/ARF7 and ARF19 promote leaf expansion and auxin-induced lateral root formation. Plant J. 43, 118–130 (2005).
Bonke, M., Thitamadee, S., Mähönen, A. P., Hauser, M. T. & Helariutta, Y. APL regulates vascular tissue identity in Arabidopsis. Nature 426, 181–186 (2003).
Zhong, R., Lee, C., Zhou, J., McCarthy, R. L. & Ye, Z. H. A battery of transcription factors involved in the regulation of secondary cell wall biosynthesis in Arabidopsis. Plant Cell 20, 2763–2782 (2008).
Baima, S. et al. The expression of the Athb-8 homeobox gene is restricted to provascular cells in Arabidopsis thaliana. Development 121, 4171–4182 (1995).
Idänheimo, N. et al. The Arabidopsis thaliana cysteine-rich receptor-like kinases CRK6 and CRK7 protect against apoplastic oxidative stress. Biochem. Biophys. Res. Commun. 445, 457–462 (2014).
Kurihara, D., Mizuta, Y., Sato, Y. & Higashiyama, T. ClearSee: a rapid optical clearing reagent for whole-plant fluorescence imaging. Development 142, 4168–4179 (2015).
Kotogany, E., Dudits, D., Horvath, G. V. & Ayaydin, F. A rapid and robust assay for detection of S-phase cell cycle progression in plant cells and tissues by using ethynyl deoxyuridine. Plant. Methods 6, 5 (2010).
Randall, R. S. et al. AINTEGUMENTA and the D-type cyclin CYCD3;1 regulate root secondary growth and respond to cytokinins. Biol. Open 4, 1229–1236 (2015).
Liebsch, D. et al. Class I KNOX transcription factors promote differentiation of cambial derivatives into xylem fibers in the Arabidopsis hypocotyl. Development 141, 4311–4319 (2014).
Czechowski, T., Stitt, M., Altmann, T., Udvardi, M. K. & Scheible, W. R. Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol. 139, 5–17 (2005).
Vandesompele, J. et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 3, RESEARCH0034 (2002).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔC T method. Methods 25, 402–408 (2001).
Yang, W., Wightman, R. & Meyerowitz, E. M. Cell cycle control by nuclear sequestration of CDC20 and CDH1 mRNA in plant stem cells. Mol. Cell 68, 1108–1119 (2017).
Mähönen, A. P. et al. Cytokinin signaling and its inhibitor AHP6 regulate cell fate during vascular development. Science 311, 94–98 (2006).
Colón-Carmona, A., You, R., Haimovitch-Gal, T. & Doerner, P. Technical advance: spatio-temporal analysis of mitotic activity with a labile cyclin–GUS fusion protein. Plant J. 20, 503–508 (1999).
Bishopp, A. et al. A mutually inhibitory interaction between auxin and cytokinin specifies vascular pattern in roots. Curr. Biol. 21, 917–926 (2011).
De Rybel, B. et al. Integration of growth and patterning during vascular tissue formation in Arabidopsis. Science 345, 1255215 (2014).
Ohashi-Ito, K. et al. A bHLH complex activates vascular cell division via cytokinin action in root apical meristem. Curr. Biol. 24, 2053–2058 (2014).
Matsumoto-Kitano, M. et al. Cytokinins are central regulators of cambial activity. Proc. Natl Acad. Sci. USA 105, 20027–20031 (2008).
Acknowledgements
We thank T. Berleth, A. Carlsbecker, T. Greb, P. Ramachandran, E. Scarpella and D. Weijers for sharing published material; A.-M. Bågman, M. Herpola and J. Smetana for technical assistance; W. Yang for helping with the in situ protocol; and S. el-Showk, P. Etchells, Y. Helariutta, S. Otero and B. Scheres for constructive comments. This work was supported by the Academy of Finland (grants #132376, #266431, #271832), University of Helsinki HiLIFE fellowship (O.S., R.M., M.L., A.S.G., F.S.R., M.L.G., R.S., T.B., S.B. and A.P.M.), Erasmus+ (F.S.R. and M.L.G.), and US National Science Foundation grant IOS-0744874 (M.-F.W. and J.W.R.).
Reviewer information
Nature thanks T. Beeckman, H. Fukuda and J. Lohmann for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
O.S. and A.P.M. designed the experiments. O.S. carried out all the experiments, except R.M. and M.L. conducted qRT–PCR, amiMP and inducible axr3 work; F.S.R. analysed HD-ZIP III mutants and S.B. carried out RNA in situ hybridization and lineage tracing with 6-benzylaminopurine. M.-F.W. and J.W.R generated and tested the amiMP plasmid. A.S.-G. and M.L.G. provided data on the inducible miR165a line. A.A. conducted the statistical analyses. R.S., S.M., P.R. and T.B. provided transgenic lines for the study. O.S. and A.P.M. wrote the manuscript with the help of co-authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Ontogeny of different cell types after activation of secondary growth in Arabidopsis root.
a, Bright-field microscopy of root cross-sections of an early phloem marker (PEAR1), a marker of differentiating phloem expressed in phloem-companion cells (APL), an early xylem marker (AtHB8) and a marker specific to differentiating xylem vessels (VND6) at different times after secondary growth activation. b, Histological analysis of different cell types found in 14-day-old root that expresses a phloem differentiation marker pAPL:GUS (top) and in 30-day-old wild-type root (bottom). Cambium (red) typically consists of 2–4 cells (14-day-old root) or 7–9 cells (30-day-old root). It is located between fully differentiated phloem elements (purple) and xylem vessels (blue), and is characterized by frequent cell divisions, detected histologically as a recently made thin cell wall. Xylem- and phloem-parenchyma cells are quiescent cells located between xylem vessels, or rarely dividing cells located between differentiated phloem cells, respectively. Data shown are representative of at least three independent experiments. Scale bars, 50 μm.
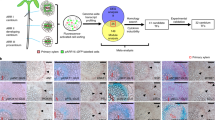
Extended Data Fig. 2 A lineage-tracing tool for developmental studies in plants.
a, A schematic of two clonal activation systems. Cre recombinase was fused to the N-terminal part of CYCB1;157 to make CYCB1;1–Cre stable only in meristematic cells that are transiting between the early G2 and M phases of the cell cycle. Heat-shock induction of pHS:CYCB1;1-cre causes recombination of two loxP sites, which activates expression of 35S:GUS (labelled (i)) or 35S:GAL4 (labelled (ii)), respectively. In the second step, the 35S:GAL4 transactivates pUAS:erGFP (labelled (iii)) and—if desired—another gene of interest cloned under pUAS in a third construct (labelled (iv)). b, Confocal microscopy of the root (RAM) and shoot (SAM; dashed line, top view) apical meristems, lateral root primordium (LRP) and procambium (PC) containing GFP clones 15 h after induction, in 5-day-old seedlings containing pHS:CYCB1;1-cre and pCB132 constructs. Because of the instability provided by the CYCB1;1-Cre fusion and the apparent non-synchronicity of the cell cycle, a high frequency of single-cell-descent clones was obtained almost exclusively in the dividing cells in the apical and lateral meristems. c, Examples of the three different GUS sector categories: basic sectors originate from a single-cell clone induced before the activation of secondary growth (red, 75%), large sectors originate from a cluster of clones simultaneously induced in several neighbouring cells (black, 19%), and half sectors originate from a single-cell clone induced after the activation of secondary growth (purple, 6%). A total of 176 sectors was analysed. d, A schematic of vascular cambium (red) and phellogen (blue) ontogeny. Under normal conditions (top) both meristems originate from XPP cells. When roots are pre-treated with cytokinin (CK), cells in the protoxylem position (black arrowheads) remain procambial cells56, and are thus capable of contributing to secondary growth (bottom, CK). In these experimental conditions, separation of vascular cambium and phellogen lineages has already occurred during primary growth, and connection with the xylem remains the determining factor for the origin of vascular cambium. e, Examples of GUS clones 24 h after heat-shock in cytokinin (BAP) pre-treated roots. Arrowheads, procambial cells in place of protoxylem; asterisks, pericycle cells that have lost their xylem connection. f, Correlation of different shapes of the GUS sector with the presence (left) or absence (centre, right) of protoxylem (insets, arrowheads). The numbers represent the frequency of these sectors. g, XPP-cell fate map of roots pre-treated with BAP. XPP cells that lose their xylem connection behave similar to pericycle cells (light purple, compare with Fig. 1b); cells in the protoxylem position behave similar to xylem-procambial cells, making wide sectors (red, compare with Fig. 1b). In b, e, f, experiments were repeated at least two times with similar results. Scale bars, 50 μm.
Extended Data Fig. 3 Transcriptional and mutant analyses of HD-ZIP III transcription factors.
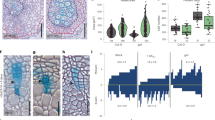
a, In situ RNA hybridization of sense- and anti-sense probes in cross-sections of 21-day-old, Col-0 plants, showing expression of HD-ZIP III transcription factors in the cambium. Insets are 2× magnifications of the cambial region of the same photograph. b, Confocal microscopy of root cross section of 14-day-old pAtHB8:erYFP plants. Cell wall staining in grey. Red arrowheads, recent cell divisions in cambium. c, Bright-field microscopy of primary and secondary vasculature shows patterning defects in higher-order HD-ZIP III mutants 7 days and 10 days after germination. Col-0 and cna-2 phb-13 phv-11 show normal differentiation of the primary xylem axis and two phloem poles, whereas the majority of athb8-11 cna-2 phb-13 phv-11 roots did not have a fully differentiated xylem axis, or showed a triarch pattern with three phloem poles. The fraction in top left corner denotes the frequency of the presented phenotype. Asterisks, primary phloem poles; arrowheads, extremities of the primary xylem. d, Confocal images of EdU-stained (green) nuclei in cna-2 phb-13 phv-1 and athb8-11 cna-2 phb-13 phv-11 mutants and the miR165a overexpressor (top). miR165a was induced for 12 days (starting from 4-day-old plants), and the mutants were 14 days old. Inhibition of HD-ZIP III transcription factors leads to cambium disorganization (histograms, bottom). Even though AtHB8, CNA, PHB and PHV—and perhaps also REV—are redundantly required for secondary xylem formation and normal cambium organization, it is possible that individual members of the family have distinct function in the cambial stem cells and their organizers. Histograms of the relative distances (RD) of EdU-positive nuclei and secondary xylem vessels in the radial direction from the root midpoint (xylem axis, x = 0; root surface, x = 1, with the scale shown in green in the Col-0 panel). KLD indicates the mean Kullback–Leibler distances between EdU-positive nuclei and vessels. nc = number of cross-sections analysed; nn = total number of nuclei. In a and b, data shown are representative of two (a) or four (b) independent experiments. Scale bars, 50 μm.
Extended Data Fig. 4 Phenotypic characterization of inducible miR165a line.
a, qRT–PCR analysis of transcription of HD-ZIP III transcription factors after miR165a induction for 24 h (in 9-day-old plants). b, miR165a-overexpressing plants grown on induction medium for 14 days (from germination onwards) resemble the quintuple HD-ZIP III mutant8. c, Quantification of secondary xylem vessel formation after miR165a induction for 2 days (in 8-day-old plants). Phenotypic changes on secondary vessel number start to appear within two days of the induction. A two-tailed Wilcoxon test (n = 25 independent roots). d, qRT–PCR analysis for marker-gene expression after a 2-day miR165a induction (in 8-day-old plants). e, Bright-field microscopy of miR165a overexpressor, double stained with Safranin and Alcian blue to highlight secondary phloem (insets) after miR165a induction for 12 days (starting from 4-day-old plants). Red dots, secondary phloem; asterisk, primary phloem; arrowheads, extremities of the primary xylem. P values in a and d are from a one-tailed t-test. Data are from three experiments (n = 10 roots per treatment). In b and e, the fraction denotes the frequency of the presented phenotype. Box-and-whisker plots show median and interquartile range; whiskers show range, and dots show outliers. Scale bars, 50 μm.
Extended Data Fig. 5 Lineage tracing in active cambium using two different lineage-tracing systems.
a–d, GUS sectors obtained after clone induction in 15-day-old (a) and 30-day-old (b–d) plants harbouring pHS:d-Box-cre and 35S:lox-GUS constructs (see Extended Data Fig. 2a). a, b, Middle panels are lateral view of roots with GUS sectors before sectioning. c, d, Lineage tracing in 30-day-old cambium revealed that bifacial stem cells are present, even in wide cambium in which 7–9-cell layers could be identified histologically (see Extended Data Fig. 1b). Quantification was performed as in Fig. 1d. e–h, Lineage tracing in ANT-expression domain. e, A schematic of clonal activation system used to induce GUS clones in the ANT-expression domain. After induction, cre-directed recombination leads to clonal activation of the GUS reporter. f, GUS sectors obtained after clone induction in 17-day-old plants containing pANT:XVE >> cre and 35S:lox-GUS. Inset is a 2.5× magnification of the cambial zone of the same image. g, h, Lineage tracing in active cambium showed ANT as a stem-cell marker. Six days after induction (in 17-day-old plants), growing GUS sectors had either a xylem (green) or phloem (yellow) fate but the majority (76%) had a cambial fate (red), on the basis of the position of thinnest cell wall (arrow) in radial direction (which represents the most recent periclinal cell division in cambium) (y = 0). d, h, Each bar represents the size and position of an individual sector (n = 131); bars with extensions denote sectors beginning or ending with a xylem vessel. a, b, f, Red arrows, sectors originating from cambium stem cells; yellow arrow, sectors on the phloem side; green arrow, sectors on the xylem side; brown arrow, examples of sectors excluded from the quantification analysis. GUS staining is weak or absent in differentiated xylem vessels. Data represents sectors collected across at least 15 (a, f–h) or 4 (b–d) independent roots. Experiments were repeated at least twice with similar results. Scale bars, 50 μm (unless otherwise specified).
Extended Data Fig. 6 Phenotypic characterization of inducible AtHB8d line.
a, qRT–PCR analysis of AtHB8 expression after AtHB8d induction for 24 h (in 9-day-old plants). b, Confocal microscopy of roots after EdU staining of S-phase nuclei reveal a centrifugal shift of the cell division domain after a 2-day AtHB8d induction (in 8-day-old plants). Asterisks mark cambium. Histograms showing the relative distances of EdU-positive nuclei in the radial direction from the root midpoint (xylem axis, x = 0, root surface, x = 1), as depicted in mock. nn, total number of nuclei. c, Quantification of secondary xylem vessel formation after AtHB8d induction for 2 days (in 8-day-old plants). d, Confocal microscopy of fluorescent markers after a 2-day AtHB8d induction (in 8-day-old plants). Cell wall staining in grey. Dashed lines, borders of the secondary tissues seen from the lateral view. Graph is quantification of the relative width of marker expression domains. e, Bright-field microscopy of roots after AtHB8 induction for 4 days (in 8-day-old plants) and quantification of secondary xylem vessel numbers. Red dots, secondary xylem vessels; yellow dashed line, cambium position in mock-treated roots. a, d, One-tailed t-test. Data are from three experiments (in a, n = 10 roots per treatment; in d, n = denominator of the fraction indicated in left panels). c, e, Two-tailed Wilcoxon test. n, independent roots. Box-and-whisker plots show median and interquartile range; whiskers show range, and dots show outliers. Scale bars, 50 μm.
Extended Data Fig. 7 Auxin signalling and ARF reporters.
a, Bright-field microscopy of DR5rev:GUS root cross-section of a 14-day-old plant. b, Confocal microscopy of DR5v2:nGFP (heat map) root cross-sections. Nuclear fluorescence signal (examples of regions of interest are indicated with a circle) was quantified in three cell layers from the most recent cell division (arrowheads) towards xylem (positions −1, −2 and −3) and phloem (positions 1, 2 and 3). Fourteen cross-sections were analysed; nn = total number of nuclei. c, Bright-field microscopy of 17-day-old roots expressing class A ARF GUS markers. d, Confocal microscopy of 14-day-old plant expressing pAtHB8:erRFP and pMP:erYFP. e, Confocal and bright-field microscopy of pMP:erYFP and DRrev:GUS marker width after a 2-day AtHB8d or miR165a induction in 8-day-old plants. Nuclear localization of AtHB8d-YFP is shown in Extended Data Fig. 6d. Cell walls in pMP:erYFP roots were stained with calcofluor (grey). The fraction in the bottom left corner denotes the frequency of the presented phenotype. White bars, width of the marker domain. Dashed lines, borders of the secondary tissues. f, qRT–PCR analysis of MP expression after a 2-day miR165a induction in 8-day-old plants. Data are from three experiments (n = 10 roots per treatment). g, Quantification of the relative width of marker expression domains after pANT:XVE >> AtHB8d-YFP induction shown in e (right panels). Sample size for each group is equal to denominator of the fraction indicated in e. b, f, g, One-tailed t-test for the comparison between different positions (b) or between mock and induced sample (f, g). Box-and-whisker plots show median and interquartile range; whiskers show range, and dots show outliers (g) or individual data points (b). a, c, d, Representative images of at least three independent observations. b, d, e, Cell wall staining in grey. p, phloem; x, xylem. Scale bars, 50 μm.
Extended Data Fig. 8 Phenotypic characterization inducible amiMP and axr3-1 lines.
a, The arf7 arf19 double mutant shows no difference in secondary xylem production, when compared to wild-type (Col-0) and analysed 14 days after germination, (nCol-0 = 17 and narf7 arf19 = 15 independent roots). b, Quantification of secondary xylem vessel formation after amiMP induction for 7 days (in 5-day-old plants) in wild-type (nCol-0 = 7 independent roots) and arf7 arf19 (narf7 arf19 = 34 independent roots) plants analysed 15 mm below the root–hypocotyl junction. The reduction in xylem vessel formation was less apparent when amiMP was induced in the wild-type background, which indicates that ARF7 and/or ARF19 are also required for this process. c, Plots of regressions between MP transcript levels and vasculature diameter (left), secondary xylem vessel formation (centre) and cell divisions (right) analysed 5 mm (top) and 15 mm (bottom) below the root–hypocotyl junction. Each point represents an individual T1 transformant or arf7 arf19 plant (namiMP = 33; narf7 arf19 = 6) grown for 5 days on a basta selection plate or 1/2 GM and then induced for 7 days. Because the residuals were not normal, a one-way Wilcoxon test for the similarity of the means of the arf7 arf19 and amiMP groups on MP transcription level was performed. A negligible P value of 3.065 × 10−7 confirms that the arf7 arf19 mean is significantly greater than the amiMP mean. To assess the linear relationship between MP transcript levels and the observed phenotypes, we performed regression analyses. The plotted blue line indicates the fitted regression model to both groups combined, and the P value indicating the goodness of this fit is plotted on the top of each respective graph. d, qRT–PCR analysis of the behaviour of HD-ZIP III transcription factors after a 24-h and 72-h induction of axr3-1 (in 8-day-old plants). Data are from four experiments (n = 16 roots per treatment). e, Bright-field microscopy and secondary xylem vessel quantification (bottom) of roots after axr3-1 induction for 48 and 72 h (in 8-day-old plants). Insets are 1.5× magnifications of the cambial region of the same photo. Arrowhead, recent cell division. Sample size for each group is equal to denominator of the fraction indicated in top panels. f, Bright-field microscopy of roots after axr3-1 induction for 7 days (starting from 5-day-old plants). Box-and-whisker plots show median and interquartile range; whiskers show range, and dots show outliers. a, b, Two-tailed Wilcoxon test. d, e, One-tailed t-test. In e, f, the fraction in top left corner denotes the frequency of the presented phenotype. In a–c, arf7, 19 refers to arf7 arf19. Scale bars, 50 μm.
Extended Data Fig. 9 Clonal activation in the S17 expression domain.
a, A schematic of the clonal activation system to induce clones in phloem-pole pericycle cells and surrounding tissue. After induction, Cre-directed recombination leads to clonal activation of GUS or the gene-of-interest (GOI) (AtHB8d, MPΔ, or gVND6) tagged with a reporter. b, Bright-field microscopy of GUS sectors in S17 position analysed 1 day after clone induction (in 5-day-old plants). Induction of large sectors (consisting of several single-cell GUS clones in the root-tip region) recapitulated the previously described expression pattern of the S17 promoter, as being specific to phloem-pole pericycle cells16 (bottom left). Higher up in the root—where secondary growth has already started—GUS clones were obtained in primary phloem and surrounding tissue (right panels), which suggests that after secondary growth activation the S17 expression domain enlarges centripetally. The experiment was repeated twice with similar results. c, d, None of 21 35S:lox-cATHB8d-GUS transformants in pS17:XVE >> cre background showed ectopic vessel formation. c, Confocal (left) and bright-field microscopy (right) analysis of AtHB8-GUS clones in a cambial marker background. Marker expression was analysed 2 days after clone induction (in 5-day-old plants). The bright-field and confocal images are from different roots. n = number of entire roots screened for ectopic marker expression. d, Confocal analysis of AtHB8d-GUS clonal activation lines performed 7 days after AtHB8d-GUS clone induction (in 5-day-old plants), with calcofluor staining in grey. n = number of entire independent roots screened for ectopic vessel formation. e, f, MPΔ- and gVND6-induced ectopic vessels show different levels of induction of periclinal cell division in neighbouring cells. Bright-field microscopy of root cross-sections, showing ectopic vessels with periclinal cell division in neighbouring cells analysed 7 days after induction (in 5-day-old plants). Yellow dots, ectopic xylem vessel; red dots, cells that have recently undergone periclinal cell division. n = number of xylem vessels examined for each category (0–7 neighbouring cells dividing periclinally). Quantification in f. Data were collected across 16 independent roots for each genotype. g, Confocal analysis of marker expression within gVND6-YFP clones (top) and around gVND6-YFP-induced ectopic vessels (bottom). The fraction in top right corner denotes the frequency of the presented phenotype. Cell walls were stained with calcofluor (grey). Insets show merged YFP and RFP channels, except for insets of WOX4 that show only the YFP channel. Scale bars, 50 μm.
Extended Data Fig. 10 Model describing the dynamic nature of vascular cambium.
a, When the organizer cells differentiate into a xylem vessel, a new organizer is formed in adjacent cambial stem cell. Stem cells and their xylem-side daughters dynamically gain organizer-cell gene expression as they mature towards xylem identity. Only the events that lead to xylem production are illustrated in the model: the mechanism for phloem production from the bifacial stem cell is unknown, and therefore not presented in the scheme. Density of the dots corresponds to the level of signalling components shown in b. b, A model presenting a signalling framework that defines the stem-cell organizer of vascular cambium. Question marks refer to unknown factors that promote stem-cell or phloem identity in a non-cell-autonomous manner. These mobile stem-cell-promoting factors could, for example, be cytokinins, which—during primary development—move from primary xylem precursor cells (marked with high levels of auxin signalling) to promote cell division in the neighbouring procambial cells58,59,60. Supporting this scenario, cytokinins are required for cambial cell proliferation during secondary development61. Alternatively, WOX4 or other transcription factors expressed in the organizer could move to the adjacent cell to promote stem-cell identity, similar to WUS and WOX5 in the apical meristems2.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables S1-S8. Full table descriptions are found in separate Supplementary tables guide
Supplementary Information
This file contains the full Supplementary Tables Guide with descriptions
Rights and permissions
About this article
Cite this article
Smetana, O., Mäkilä, R., Lyu, M. et al. High levels of auxin signalling define the stem-cell organizer of the vascular cambium. Nature 565, 485–489 (2019). https://doi.org/10.1038/s41586-018-0837-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0837-0
This article is cited by
-
Single-cell transcriptomics unveils xylem cell development and evolution
Genome Biology (2023)
-
Genes expression profiles in vascular cambium of Eucalyptus urophylla × Eucalyptus grandis at different ages
BMC Plant Biology (2023)
-
Single-cell resolution analysis reveals the preparation for reprogramming the fate of stem cell niche in cotton lateral meristem
Genome Biology (2023)
-
Transformation and expressional studies of GaZnF gene to improve drought tolerance in Gossypium hirsutum
Scientific Reports (2023)
-
An integrase toolbox to record gene-expression during plant development
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.