Abstract
The immune system can suppress tumour development both by eliminating malignant cells and by preventing the outgrowth and spread of cancer cells that resist eradication1. Clinical and experimental data suggest that the latter mode of control—termed cancer–immune equilibrium1—can be maintained for prolonged periods of time, possibly up to several decades2,3,4. Although cancers most frequently originate in epithelial layers, the nature and spatiotemporal dynamics of immune responses that maintain cancer–immune equilibrium in these tissue compartments remain unclear. Here, using a mouse model of transplantable cutaneous melanoma5, we show that tissue-resident memory CD8+ T cells (TRM cells) promote a durable melanoma–immune equilibrium that is confined to the epidermal layer of the skin. A proportion of mice (~40%) transplanted with melanoma cells remained free of macroscopic skin lesions long after epicutaneous inoculation, and generation of tumour-specific epidermal CD69+ CD103+ TRM cells correlated with this spontaneous disease control. By contrast, mice deficient in TRM formation were more susceptible to tumour development. Despite being tumour-free at the macroscopic level, mice frequently harboured melanoma cells in the epidermal layer of the skin long after inoculation, and intravital imaging revealed that these cells were dynamically surveyed by TRM cells. Consistent with their role in melanoma surveillance, tumour-specific TRM cells that were generated before melanoma inoculation conferred profound protection from tumour development independently of recirculating T cells. Finally, depletion of TRM cells triggered tumour outgrowth in a proportion (~20%) of mice with occult melanomas, demonstrating that TRM cells can actively suppress cancer progression. Our results show that TRM cells have a fundamental role in the surveillance of subclinical melanomas in the skin by maintaining cancer–immune equilibrium. As such, they provide strong impetus for exploring these cells as targets of future anticancer immunotherapies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding authors upon reasonable request.
Change history
11 February 2019
Panel j was inadvertently labelled as panel k in the caption to Fig. 4. Similarly, ‘Fig. 4k’ should have been ‘Fig. 4j’ in the sentence beginning ‘TNF-α-deficient gBT-I cells were…’. In addition, the surname of author Umaimainthan Palendira was misspelled ‘Palendria’. These errors have been corrected online.
References
Schreiber, R. D., Old, L. J. & Smyth, M. J. Cancer immunoediting: integrating immunity’s roles in cancer suppression and promotion. Science 331, 1565–1570 (2011).
MacKie, R. M., Reid, R. & Junor, B. Fatal melanoma transferred in a donated kidney 16 years after melanoma surgery. N. Engl. J. Med. 348, 567–568 (2003).
Koebel, C. M. et al. Adaptive immunity maintains occult cancer in an equilibrium state. Nature 450, 903–907 (2007).
Teng, M. W. et al. Opposing roles for IL-23 and IL-12 in maintaining occult cancer in an equilibrium state. Cancer Res. 72, 3987–3996 (2012).
Wylie, B. et al. Cross-presentation of cutaneous melanoma antigen by migratory XCR1+CD103− and XCR1+CD103+ dendritic cells. OncoImmunology 4, e1019198 (2015).
Gebhardt, T. et al. Different patterns of peripheral migration by memory CD4+ and CD8+ T cells. Nature 477, 216–219 (2011).
Jiang, X. et al. Skin infection generates non-migratory memory CD8+ TRM cells providing global skin immunity. Nature 483, 227–231 (2012).
Gebhardt, T. et al. Memory T cells in nonlymphoid tissue that provide enhanced local immunity during infection with herpes simplex virus. Nature Immunol. 10, 524–530 (2009).
Ariotti, S. et al. Tissue-resident memory CD8+ T cells continuously patrol skin epithelia to quickly recognize local antigen. Proc. Natl Acad. Sci. USA 109, 19739–19744 (2012).
Mackay, L. K. et al. Long-lived epithelial immunity by tissue-resident memory T (TRM) cells in the absence of persisting local antigen presentation. Proc. Natl Acad. Sci. USA 109, 7037–7042 (2012).
Gebhardt, T., Palendira, U., Tscharke, D. C. & Bedoui, S. Tissue-resident memory T cells in tissue homeostasis, persistent infection, and cancer surveillance. Immunol. Rev. 283, 54–76 (2018).
Enamorado, M. et al. Enhanced anti-tumour immunity requires the interplay between resident and circulating memory CD8+ T cells. Nat. Commun. 8, 16073 (2017).
Malik, B. T. et al. Resident memory T cells in the skin mediate durable immunity to melanoma. Sci. Immunol. 2, eaam6346 (2017).
Murray, T. et al. Very late antigen-1 marks functional tumor-resident CD8 T cells and correlates with survival of melanoma patients. Front. Immunol. 7, 573 (2016).
Boddupalli, C. S. et al. Interlesional diversity of T cell receptors in melanoma with immune checkpoints enriched in tissue-resident memory T cells. JCI Insight 1, e88955 (2016).
Salerno, E. P., Olson, W. C., McSkimming, C., Shea, S. & Slingluff, C. L. Jr T cells in the human metastatic melanoma microenvironment express site-specific homing receptors and retention integrins. Int. J. Cancer 134, 563–574 (2014).
Edwards, J. et al. CD103+ tumor-resident CD8+ T cells are associated with improved survival in immunotherapy naive melanoma patients and expand significantly during anti-PD1 treatment. Clin. Cancer Res. 24, 3036–3045 (2018).
Mackay, L. K. et al. The developmental pathway for CD103+ CD8+ tissue-resident memory T cells of skin. Nat. Immunol. 14, 1294–1301 (2013).
Schenkel, J. M., Fraser, K. A., Vezys, V. & Masopust, D. Sensing and alarm function of resident memory CD8+ T cells. Nat. Immunol. 14, 509–513 (2013).
Landsberg, J. et al. Melanomas resist T-cell therapy through inflammation-induced reversible dedifferentiation. Nature 490, 412–416 (2012).
Braumüller, H. et al. T-helper-1-cell cytokines drive cancer into senescence. Nature 494, 361–365 (2013).
Webb, J. R., Milne, K., Watson, P., Deleeuw, R. J. & Nelson, B. H. Tumor-infiltrating lymphocytes expressing the tissue resident memory marker CD103 are associated with increased survival in high-grade serous ovarian cancer. Clin. Cancer Res. 20, 434–444 (2014).
Djenidi, F. et al. CD8+ CD103+ tumor-infiltrating lymphocytes are tumor-specific tissue-resident memory T cells and a prognostic factor for survival in lung cancer patients. J. Immunol. 194, 3475–3486 (2015).
Wang, B. et al. CD103+ tumor infiltrating lymphocytes predict a favorable prognosis in urothelial cell carcinoma of the bladder. J. Urol. 194, 556–562 (2015).
Komdeur, F. L. et al. CD103+ tumor-infiltrating lymphocytes are tumor-reactive intraepithelial CD8+ T cells associated with prognostic benefit and therapy response in cervical cancer. OncoImmunology 6, e1338230 (2017).
Nizard, M. et al. Induction of resident memory T cells enhances the efficacy of cancer vaccine. Nat. Commun. 8, 15221 (2017).
Cockburn, I. A. et al. In vivo imaging of CD8+ T cell-mediated elimination of malaria liver stages. Proc. Natl Acad. Sci. USA 110, 9090–9095 (2013).
Halle, S. et al. In vivo killing capacity of cytotoxic T cells is limited and involves dynamic interactions and T cell cooperativity. Immunity 44, 233–245 (2016).
Zhu, J. et al. Immune surveillance by CD8αα+ skin-resident T cells in human herpes virus infection. Nature 497, 494–497 (2013); corrigendum 500, 242 (2013).
Woon, H. G. et al. Compartmentalization of total and virus-specific tissue-resident memory CD8+ T cells in human lymphoid organs. PLoS Pathog. 12, e1005799 (2016).
Mueller, S. N., Heath, W., McLain, J. D., Carbone, F. R. & Jones, C. M. Characterization of two TCR transgenic mouse lines specific for herpes simplex virus. Immunol. Cell Biol. 80, 156–163 (2002).
van Lint, A. et al. Herpes simplex virus-specific CD8+ T cells can clear established lytic infections from skin and nerves and can partially limit the early spread of virus after cutaneous inoculation. J. Immunol. 172, 392–397 (2004).
Acknowledgements
We thank F. Carbone and A. Kallies for critical reading of our manuscript. This work was supported by the University of Melbourne (Elizabeth and Vernon Puzey Scholarship to S.L.P); the Sylvia and Charles Viertel Charitable Foundation (fellowship to T.G.); the Australian National Health and Medical Research Council (fellowships to L.K.M., R.A.S., J.S.W, S.N.M. and N.D.H; grants to R.A.S., J.S.W (1093017) and N.D.H. (1124907, 1124784)); the Cancer Councils of Victoria (grant to N.D.H. (1145730)) and Western Australia (fellowship to J.W.); BHP (grant to J.W.); and the German Research Foundation (GRK2168 Bo&MeRanG Faculty Support Scholarship to M.E., Excellence Cluster ImmunoSensation to M.H. and Program Grant SFB 854/TP27 to T.T.). K.H. is a Rhian and Paul Brazis Fellow in Translational Melanoma Immunology. N.D.H. was supported by the Harry J. Lloyd Charitable Trust, Melanoma Research Alliance, Ian Potter Foundation, Tour de Cure and Cancer Research Institute (USA) and the Victorian State Government Operational Infrastructure Support Scheme.
Reviewer information
Nature thanks A. Goldrath, D. Masopust and D. Speiser for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
S.L.P., J.R., J.L.H, K.H., N.M., J.E. and R.M. performed experiments. A.B., M.E., T.W., T.T., M.H. and J.W. generated cell lines. S.L.P., U.P., L.K.M., J.W. and T.G. designed experiments. S.L.P., N.M., J.E., U.P. and T.G. analysed data. R.A.S., D.G., S.N.M., N.D.H., J.S.W. and S.B. contributed intellectual input and helped to interpret data. T.G. led the research program. S.L.P. and T.G. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Growth kinetics and persistence of B16.gB cells after e.c. inoculation.
a, Rate of tumour incidence across individual experiments after s.c. (Subcut.) and e.c. (Epicut.) inoculation with either B16.gB or B16.gB.Luc cells in WT mice. Data pooled from n = 3 (s.c.) and n = 46 (e.c.) biologically independent experiments with n = 15 (s.c.) or n = 452 (e.c.) mice. b, Tumour growth kinetics after s.c. or e.c. inoculation with B16.gB or B16.gB.Luc cells. Data pooled from n = 3 biologically independent experiments with n = 15 mice (s.c.) or n = 22 independent experiments with n = 154 mice (e.c.). c, Photographs of metastases in tumour-draining brachial lymph nodes (LNs) of mice with progressive melanoma following e.c. B16.gB inoculation. Representative of n = 46 biologically independent experiments. d, B6 albino mice were inoculated epicutaneously with B16.gB.Luc cells and subjected to longitudinal bioluminescence imaging. Bioluminescence signals (arrows) emitted from progressing melanomas or from skin of non-developer mice. Data representative of n = 2 biologically independent experiments with n = 18 mice. e, Two-photon microscopy image of a macroscopic melanoma 2 or more weeks after e.c. inoculation with B16.Tyr–/–.mCherry cells. f, Two-photon microscopy images of macroscopically tumour-free skin with persistent B16.Tyr–/–.mCherry cells (arrows) at indicated times post-inoculation. e, f, B16.Tyr–/–.mCherry (B16) cells, red; SHG, blue. Note autofluorescent hair (green). Data representative of n = 3 biologically independent experiments with n = 22 mice.
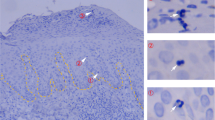
Extended Data Fig. 2 Responses of gBT-I cells to e.c. inoculation with B16.gB cells.
WT mice were transferred with naive gBT-I.CD45.1 cells before e.c. inoculation with B16.gB cells. a, Detection of gBT-I cells (blue ovals; based on expression of CD45.1 and the T-cell-receptor chain Vα2) in spleen, tumour-draining brachial lymph nodes (brLN) and tumour-challenged skin at 7 days (d7) post-inoculation. b, c, Number (b) and phenotype (c) of gBT-I cells shown in panel a. Data representative of (a, c) or pooled from (b) n = 2 biologically independent experiments with n = 10 mice. d, Detection of gBT-I.CD45.1 cells in the indicated organs from developer (‘Melanoma’) and non-developer (‘Tumour-free’) mice at 14 days post-inoculation. Data representative of n = 2 biologically independent experiments with n = 10 mice. e, Number of gBT-I.CD45.1 cells isolated from the indicated organs of developer (‘Melanoma’) and non-developer (‘Tumour-free’) mice 14 days post-inoculation. Data pooled from n = 2 biologically independent experiments with n = 10 (Melanoma) or n = 5 (Tumour-free) mice. f–h, Number (f) and phenotype (g, h) of gBT-I.CD45.1 cells isolated from the indicated organs of developer (‘Melanoma’) and non-developer (‘Tumour-free’) mice more than 21 days post-inoculation. Data pooled from n = 5 biologically independent experiments with n = 83 mice. ***P = 0.0006, ****P < 0.0001, Mann–Whitney test.
Extended Data Fig. 3 Generation of melanoma-associated TRM cells does not require local antigen.
a, Experimental protocol. WT mice were inoculated epicutaneously with B16.gB cells on the upper left flank and monitored for tumour progression. Mice developing macroscopic melanoma were co-transferred intravenously (i.v.) with equal numbers of naive gBT-I.CD45.1 cells and OT-I.CD45.1.CD45.2 cells, and challenged with HSV.Ova on the lower right flank. Tissues were harvested for fluorescence-activated cell sorting (FACS) analysis 4 weeks after HSV infection. b, Surface phenotype of gBT-I or OT-I cells isolated from peritumoural skin 4 weeks after HSV infection. c, d, Proportion of gBT-I or OT-I cells expressing CD69 and CD103 (c) or CD69 only (d) in the indicated organs (Spl, spleen; Peritum, peritumoural). Data are representative of (b) or pooled (c) from one experiment with n = 6 mice.
Extended Data Fig. 4 gBT-I.WT cells can rescue the melanoma-protection defect in Cd69–/– mice.
a, WT or Cd69–/– mice were transferred naive gBT-I.WT or gBT-I.Cd69–/– (1 × 105) cells, then inoculated epicutaneously with B16.gB cells and monitored for tumour development. b, Proportion of tumour-free non-developer WT (grey and black) or Cd69–/– (coloured) mice following tumour inoculation over time. Data pooled from n = 2 biologically independent experiments with n = 12 (gBT-I.WT→WT, gBT-I.Cd69–/–→WT), n = 14 (gBT-I.Cd69–/–→Cd69–/–) or n = 15 (gBT-I.WT→Cd69–/–) mice per group. *P = 0.0217, log-rank Mantel Cox test.
Extended Data Fig. 5 MHC I expression by progressing or controlled melanoma.
a, WT mice were transferred naive gBT-I.CD45.1 cells and inoculated epicutaneously with B16.gB.Tyr–/–.mCherry cells. Tumours isolated from mice with progressively growing melanomas (defined by an increase in tumour volume over at least four consecutive days; ‘escaped’) or tumours that remained small and did not progress (defined by a stable tumour volume over a period of 1.5–3 weeks; ‘controlled’) were harvested and analysed by flow cytometry. a, Proportion of B16.gB.Tyr–/–.mCherry cells isolated from escaped or controlled tumours that expressed MHC class I (H2-Kb) proteins at the time of harvest. ***P = 0.0003. b, Geometric mean fluorescence intensity (gMFI) of MHC I expression by B16.gB.Tyr–/–.mCherry cells isolated from escaped or controlled tumours, normalized to the gMFI of fluorescence minus one (FMO) controls. ***P = 0.0005 Data in a, b pooled from n = 3 biologically independent experiments with n = 19 (escaped) or n = 8 (controlled) mice. Mann–Whitney test.
Extended Data Fig. 6 Tumour-primed gBT-I.uGFP cells survey developing and persistent melanoma in skin.
B6 albino mice were transferred naive gBT-I.uGFP cells and inoculated epicutaneously with B16.gB.Tyr–/–.mCherry cells. a, Two-photon microscopy images depicting localization of gBT-I.uGFP cells around a B16.gB.Tyr–/–.mCherry tumour (red) in the skin of mice developing macroscopic melanoma. b, Representative two-photon microscopy image and time-lapse series (minutes, ′) of gBT-I.uGFP cells in proximity to persistent B16.gB.Tyr–/–.mCherry cells in the skin of a non-developer mouse. a, b, Red, B16.gB.Tyr–/–.mCherry cells (B16.gB; arrows); green, gBT-I.uGFP cells (gBT-I); blue, SHG. Note autofluorescent hair (green). Data in a and b are representative of n = 3 biologically independent experiments with n = 17 mice.
Extended Data Fig. 7 Detection of B16-derived genomic DNA in skin of HSV-immune non-developer mice.
Frequency of luciferase (Luc) gDNA detection by ddPCR in the skin of previously naive mice developing macroscopic melanoma (a positive control), HSV-immune non-developer mice, or naive mouse skin (negative control) more than 4 weeks after e.c. inoculation with B16.gB.Luc cells. Data pooled from n = 2 biologically independent experiments with n = 17 (HSV), n = 6 (naive skin) or n = 2 (melanoma) mice.
Extended Data Fig. 8 Generation of TRM cells by localized deposition, and antibody strategies for selective TCIRC or TRM depletion.
WT mice received activated gBT-I.CD45.1 or gBT-I.Thy1.1 cells (1 × 106 or 4 × 106) by e.c. transfer. a, Frequency and phenotype of transferred Vα2+ gBT-I.CD45.1 cells in the spleen and skin at 8 days and more than 4 weeks after transfer. Data representative of n = 2 biologically independent experiments with n = 10 mice (day 8) and n = 8 mice (more than 4 weeks). b, Number of gBT-I.CD45.1 cells isolated from spleen and skin more than 4 weeks after transfer, determined by flow cytometry. Data pooled from n = 5 biologically independent experiments with n = 21 mice. c, d, Frequency (c) and number (d) of e.c. transferred gBT-I.Thy1.1 cells isolated from the indicated organs more than 7 days after intraperitoneal (i.p.) treatment with PBS (‘Control’) or anti-Thy1.1 antibody (‘Depletion’). Data representative of (c) or pooled from (d) n = 2 biologically independent experiments with n = 9 mice per group. ***P = 0.0002, Mann–Whitney test. e, f, Frequency (e) and number (f) of e.c. transferred gBT-I.Thy1.1 cells in the spleen and skin more than 50 days after i.p. treatment with low-dose anti-Thy1.1 antibody and intradermal (i.d.) treatment with high-dose anti-Thy1.1 antibody or PBS. Data representative of (e) or pooled from (f) n = 4 biologically independent experiments with n = 20 (i.p. only) or n = 22 (i.p. + i.d.) mice. Only values greater than zero are plotted on the logarithmic scale. *P = 0.0261, Mann–Whitney test.
Extended Data Fig. 9 Tumour incidence in mice genetically deficient in perforin or IFN-γ production.
a, Proportion of tumour-free non-developer WT or perforin-deficient Perf1–/– mice after e.c. inoculation with B16.gB cells. b, Proportion of tumour-free non-developer WT or IFN-γ-deficient (Ifng–/–) mice following e.c. inoculation with B16.gB cells. Data pooled from n = 4 (a) or n = 3 (b) biologically independent experiments with n = 36 (Perf1–/–) or n = 41 (WT) mice (a) or n = 32 mice per group (b).
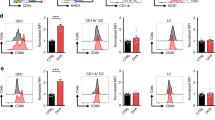
Extended Data Fig. 10 Cytokine production and expression of inhibitory receptors by melanoma-associated TRM cells.
a, WT mice were transferred naive gBT-I.CD45.1 (gBT-I) cells and e.c. inoculated with B16.gB cells. A separate group of mice received naive gBT-I.CD45.1 cells followed by skin infection with HSV (not shown). Three to four weeks after tumour or viral challenge, mice were i.v. injected with either gB or Ova peptide and brefeldin A, and organs were harvested for flow cytometric analysis 4–5 hours later. b, Proportion of gBT-I cells expressing the indicated cytokines (IFN-γ and TNF-α) isolated from HSV-challenged skin (green), peritumoural skin from developer mice (pink) or tumour-free skin of non-developer mice (orange) administered either gB or Ova peptide. c, In vitro activated gBT-I.CD45.1 cells were transferred to WT mice bearing macroscopic B16.gB tumours or to a separate group of previously naive mice that were then treated with DNFB on skin to facilitate TRM generation. Two weeks later, populations of CD69+ CD103+, CD69+ CD103− and CD69− CD103− gBT-I cells were sorted from DNFB-treated skin (‘Control skin’) or tumour samples, respectively, and expression of the indicated genes was analysed using qPCR. Gene expression is normalized to expression of the housekeeping genes Hprt, Gapdh and Tbp and shown as fold change relative to expression in CD69− populations of gBT-I cells in the spleen. d, WT mice were transferred naive gBT-I.CD45.1 cells and e.c. inoculated with B16.gB cells. Shown is surface expression of PD-1 by gBT-I cells isolated from the indicated organs compared with PD-1 expression in bulk CD8+ Vα2+ T cells from the same tissue at more than 3 weeks post-inoculation. e, Mice were transferred naive gBT-I.CD45.1 cells and e.c. inoculated with B16.gB.Tyr–/–.mCherry cells. Shown is the proportion of gBT-I TRM cells in peritumoural or tumour-free (non-developer) skin expressing PD-1 at more than 2–3 weeks post-inoculation. Data pooled from n = 4 biologically independent experiments with n = 9 (HSV, Ova), n = 17 (melanoma, Ova), n = 5 (non-developer, Ova), n = 11 (HSV, gB), n = 19 (melanoma, gB) or n = 9 (non-developer, gB) mice (a, b); n = 2 biologically independent experiments with n = 20 (control) and n = 43 (melanoma) mice per group (c); are representative of n = 2 biologically independent experiments with n = 10 mice (d); or pooled from n = 2 biologically independent experiments with n = 27 (melanoma, peritumoural) or n = 20 (non-developer) mice (e).
Supplementary information
Video 1 gBT-I TRM survey progressing melanoma in skin.
WT mice were transferred naive gBT-I.uGFP (green) prior to e.c. inoculation with B16.gB.Tyr–/–.mCherry (red) and imaged more than 2 wks later after developing macroscopic melanoma. Data representative of n = 3 biologically independent experiments with n = 17 mice. Second harmonic generation signal (SHG), blue
Video 2 gBT-I TRM cells survey persistent melanoma cells in skin.
WT mice were transferred naive gBT-I.uGFP (green) prior to e.c. inoculation with B16.gB.Tyr–/–.mCherry (red). Non-developers remaining macroscopically tumour-free were imaged more than 4 wks p.i. Data representative of n = 3 biologically independent experiments with n = 17 mice. Second harmonic generation (SHG), blue
Video 3 gBT-I TRM cells survey persistent melanoma cluster in skin.
WT mice were transferred naive gBT-I.uGFP (green) prior to e.c. inoculation with B16.gB.Tyr–/–.mCherry (red). Non-developers remaining macroscopically tumour-free were imaged more than 4 wks p.i. Data representative of n = 3 biologically independent experiments with n = 17 mice. Second harmonic generation (SHG), blue
Source data
Rights and permissions
About this article
Cite this article
Park, S.L., Buzzai, A., Rautela, J. et al. Tissue-resident memory CD8+ T cells promote melanoma–immune equilibrium in skin. Nature 565, 366–371 (2019). https://doi.org/10.1038/s41586-018-0812-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0812-9
This article is cited by
-
Dormancy of cutaneous melanoma
Cancer Cell International (2024)
-
Decoding the transcriptional heterogeneity, differentiation lineage, clinical significance in tissue-resident memory CD8 T cell of the small intestine by single-cell analysis
Journal of Translational Medicine (2024)
-
Novel strategies for cancer immunotherapy: counter-immunoediting therapy
Journal of Hematology & Oncology (2023)
-
Flubendazole inhibits PD-1 and suppresses melanoma growth in immunocompetent mice
Journal of Translational Medicine (2023)
-
Prognostic values of tissue-resident CD8+T cells in human hepatocellular carcinoma and intrahepatic cholangiocarcinoma
World Journal of Surgical Oncology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.