Abstract
Environmental cues profoundly affect cellular plasticity in multicellular organisms. For instance, exercise promotes a glycolytic-to-oxidative fibre-type switch in skeletal muscle, and cold acclimation induces beige adipocyte biogenesis in adipose tissue. However, the molecular mechanisms by which physiological or pathological cues evoke developmental plasticity remain incompletely understood. Here we report a type of beige adipocyte that has a critical role in chronic cold adaptation in the absence of β-adrenergic receptor signalling. This beige fat is distinct from conventional beige fat with respect to developmental origin and regulation, and displays enhanced glucose oxidation. We therefore refer to it as glycolytic beige fat. Mechanistically, we identify GA-binding protein α as a regulator of glycolytic beige adipocyte differentiation through a myogenic intermediate. Our study reveals a non-canonical adaptive mechanism by which thermal stress induces progenitor cell plasticity and recruits a distinct form of thermogenic cell that is required for energy homeostasis and survival.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
RNA-seq datasets generated in this study are available at ArrayExpress under the following accession codes: E-MTAB-4526 (adipose tissues in β-less mice), E-MTAB-4528 (skeletal muscle in β-less mice), E-MTAB-6392 (Myod1+-derived beige fat), E-MTAB-7175 (BAT, WAT and skeletal muscle), E-MTAB-6441 (Myod1+-derived progenitors), and E-MTAB-7164 (myoblasts). The datasets in the present study are available from the corresponding author upon request.
References
Collins, S. β-adrenoceptor signaling networks in adipocytes for recruiting stored fat and energy expenditure. Front. Endocrinol. 2, 102 (2011).
Seale, P. et al. Prdm16 determines the thermogenic program of subcutaneous white adipose tissue in mice. J. Clin. Invest. 121, 96–105 (2011).
Ikeda, K. et al. UCP1-independent signaling involving SERCA2b-mediated calcium cycling regulates beige fat thermogenesis and systemic glucose homeostasis. Nat. Med. 23, 1454–1465 (2017).
Cohen, P. et al. Ablation of PRDM16 and beige adipose causes metabolic dysfunction and a subcutaneous to visceral fat switch. Cell 156, 304–316 (2014).
Sharp, L. Z. et al. Human BAT possesses molecular signatures that resemble beige/brite cells. PLoS ONE 7, e49452 (2012).
Wu, J. et al. Beige adipocytes are a distinct type of thermogenic fat cell in mouse and human. Cell 150, 366–376 (2012).
Lidell, M. E. et al. Evidence for two types of brown adipose tissue in humans. Nat. Med. 19, 631–634 (2013).
Shinoda, K. et al. Genetic and functional characterization of clonally derived adult human brown adipocytes. Nat. Med. 21, 389–394 (2015).
Yoneshiro, T. et al. Recruited brown adipose tissue as an antiobesity agent in humans. J. Clin. Invest. 123, 3404–3408 (2013).
van der Lans, A. A. et al. Cold acclimation recruits human brown fat and increases nonshivering thermogenesis. J. Clin. Invest. 123, 3395–3403 (2013).
Hanssen, M. J. et al. Short-term cold acclimation improves insulin sensitivity in patients with type 2 diabetes mellitus. Nat. Med. 21, 863–865 (2015).
Arch, J. R. Challenges in β3-adrenoceptor agonist drug development. Ther. Adv. Endocrinol. Metab. 2, 59–64 (2011).
Bachman, E. S. et al. βAR signaling required for diet-induced thermogenesis and obesity resistance. Science 297, 843–845 (2002).
Ye, L. et al. Fat cells directly sense temperature to activate thermogenesis. Proc. Natl Acad. Sci. USA 110, 12480–12485 (2013).
Razzoli, M. et al. Stress-induced activation of brown adipose tissue prevents obesity in conditions of low adaptive thermogenesis. Mol. Metab. 5, 19–33 (2016).
Sanchez-Gurmaches, J. & Guertin, D. A. Adipocytes arise from multiple lineages that are heterogeneously and dynamically distributed. Nat. Commun. 5, 4099 (2014).
Berry, D. C., Jiang, Y. & Graff, J. M. Mouse strains to study cold-inducible beige progenitors and beige adipocyte formation and function. Nat. Commun. 7, 10184 (2016).
Rodeheffer, M. S., Birsoy, K. & Friedman, J. M. Identification of white adipocyte progenitor cells in vivo. Cell 135, 240–249 (2008).
Tseng, Y. H. et al. New role of bone morphogenetic protein 7 in brown adipogenesis and energy expenditure. Nature 454, 1000–1004 (2008).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Yang, Z. F., Drumea, K., Mott, S., Wang, J. & Rosmarin, A. G. GABP transcription factor (nuclear respiratory factor 2) is required for mitochondrial biogenesis. Mol. Cell. Biol. 34, 3194–3201 (2014).
Mootha, V. K. et al. Errα and Gabpa/b specify PGC-1α-dependent oxidative phosphorylation gene expression that is altered in diabetic muscle. Proc. Natl Acad. Sci. USA 101, 6570–6575 (2004).
Gantner, M. L., Hazen, B. C., Eury, E., Brown, E. L. & Kralli, A. Complementary roles of estrogen-related receptors in brown adipocyte thermogenic function. Endocrinology 157, 4770–4781 (2016).
Jaworski, A., Smith, C. L. & Burden, S. J. GA-binding protein is dispensable for neuromuscular synapse formation and synapse-specific gene expression. Mol. Cell. Biol. 27, 5040–5046 (2007).
Lee, Y. H., Petkova, A. P., Mottillo, E. P. & Granneman, J. G. In vivo identification of bipotential adipocyte progenitors recruited by β3-adrenoceptor activation and high-fat feeding. Cell Metab. 15, 480–491 (2012).
Vishvanath, L. et al. Pdgfrβ+ mural preadipocytes contribute to adipocyte hyperplasia induced by high-fat-diet feeding and prolonged cold exposure in adult mice. Cell Metab. 23, 350–359 (2016).
Long, J. Z. et al. A smooth muscle-like origin for beige adipocytes. Cell Metab. 19, 810–820 (2014).
Kajimura, S., Spiegelman, B. M. & Seale, P. Brown and beige fat: physiological roles beyond heat generation. Cell Metab. 22, 546–559 (2015).
Nedergaard, J. & Cannon, B. UCP1 mRNA does not produce heat. Biochim. Biophys. Acta 1831, 943–949 (2013).
Kazak, L. et al. A creatine-driven substrate cycle enhances energy expenditure and thermogenesis in beige fat. Cell 163, 643–655 (2015).
Hasegawa, Y. et al. Repression of adipose tissue fibrosis through a PRDM16–GTF2IRD1 complex improves systemic glucose homeostasis. Cell Metab. 27, 180–194 (2018).
Nishijo, K. et al. Biomarker system for studying muscle, stem cells, and cancer in vivo. FASEB J. 23, 2681–2690 (2009).
Jung, S. et al. In vivo depletion of CD11c+ dendritic cells abrogates priming of CD8+ T cells by exogenous cell-associated antigens. Immunity 17, 211–220 (2002).
Buch, T. et al. A Cre-inducible diphtheria toxin receptor mediates cell lineage ablation after toxin administration. Nat. Methods 2, 419–426 (2005).
Liisberg Aune, U., Ruiz, L. & Kajimura, S. Isolation and differentiation of stromal vascular cells to beige/brite cells. J. Vis. Exp. (2013).
Ohno, H., Shinoda, K., Spiegelman, B. M. & Kajimura, S. PPARγ agonists induce a white-to-brown fat conversion through stabilization of PRDM16 protein. Cell Metab. 15, 395–404 (2012).
Trapnell, C. et al. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nat. Biotechnol. 31, 46–53 (2013).
Tripathi, S. et al. Meta- and orthogonal integration of influenza “OMICs” data defines a role for UBR4 in virus budding. Cell Host Microbe 18, 723–735 (2015).
Acknowledgements
We thank Y. Seo, S. Haldar, S. Burden and C. Keller for sharing their reagents and technical advice. This work was supported by the NIH (DK97441, DK112268, and DK108822) and the Edward Mallinckrodt Jr. Foundation to S.K., and AR060868 and AR061002 to A.B.
Reviewer information
Nature thanks A. Pfeifer and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Y.C. and S.K. conceived the study and designed experiments. Y.C., K.I., T.Y., A.S., K.T., Q.W., K.K., C.H.S. and Z.B. performed experiments. K.S. performed bioinformatics analyses. Y.C., K.I., T.Y., A.S., K.T., Q.W., K.K., C.H.S., K.S. and S.K. analysed and interpreted the data. Y.C. and S.K. wrote the manuscript. Y.C., Z.B., A.B. and S.K. edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Mouse models with defects in β-AR signalling.
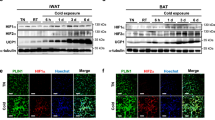
a, Schematic of the experiment. The inguinal WAT from wild-type and β-less mice at 23 °C or 15 °C was collected and analysed by RNA-seq. b, mRNA expression of indicated genes in interscapular BAT (iBAT) of mice in Fig. 1a. n.s., not significant. n = 5. c, GO analysis of gene set I in Fig. 1a. P values (−log10) by delta-method–based test. d, mRNA expression of gene set I: Ucp1, n = 10; Elovl3, n = 5; Cidea, n = 5; Pgc1a, n = 5; Cox8b, n = 5. e, mRNA expression of skeletal muscle-related genes in inguinal WAT. Myh1, n = 9; Myl1, n = 8; Mylpf, n = 8; Mybpc1, n = 9; Acta1, n = 8. f, mRNA expression of myogenesis-related genes in the iBAT of mice in a. n = 5. g, mRNA expression of indicated genes in the skeletal muscle of mice in a. n = 5. h, Schematic of the experiment. Wild-type mice were treated with β-blocker or vehicle (saline) at 23 °C for 5 days. i, mRNA expression of indicated genes in inguinal WAT (left), iBAT (middle) and skeletal muscle (right) of mice treated with β-blocker or vehicle. *P < 0.05. n = 4. Data are mean ± s.e.m. of biologically independent samples; unpaired two-sided Student’s t-test. j, Immunofluorescent staining of MHC in differentiated SVFs from the iBAT of mice in h. k, Immunofluorescent staining of MHC in differentiated SVF cells from the epididymal WAT of mice in h. l, Schematic of the experiment. SVFs from β-blocker-treated Myod1-CreERT2 reporter mice were cultured. b, d–g, Data are mean ± s.e.m. of biologically independent samples; one-way ANOVA followed by Tukey’s test. j, k, DAPI for counter staining. Images represent three independent replicates. Scale bar, 50 µm.
Extended Data Fig. 2 Myod1+-derived beige adipocytes in adipose tissue.
a, Immunofluorescent staining of GFP and UCP1 in the anterior, middle and posterior regions of inguinal WAT of Myod1-CreERT2 reporter mice. Mice were pre-treated with β-blocker and acclimated to 15 °C. Scale bar, 100 µm. Note that GFP+ adipocytes were enriched in the middle part of inguinal WAT near the lymph node (LN). The ratio (%) of UCP1+ cells among total GFP+ cells is shown on the top. N.D., not detected. b, Immunofluorescent staining of GFP and CD31 in the middle part of inguinal WAT. Myod1-CreERT2 reporter mice treated with β-blocker. Scale bar, 100 µm. Note that GFP+ adipocytes are localized near the CD31+ microvasculature. c, Immunofluorescent staining of GFP and UCP1 in the iBAT of Myod1-CreERT2 reporter mice treated with β-blocker or vehicle. Scale bar, 100 µm. d, Immunofluorescent staining of GFP and UCP1 in the epididymal WAT of mice in c. Scale bar, 100 µm. e, Immunofluorescent staining of GFP and UCP1 in the inguinal WAT of Myod1-CreERT2 reporter mice that were acclimated to 6 °C for 2 weeks without β-blocker treatment. Scale bar, 100 µm. f, Immunofluorescent staining of GFP and UCP1 in the inguinal WAT of Myod1-CreERT2 reporter mice treated with CL316,243 (1 mg per kg (body weight) per day) for 2 weeks. Scale bar, 100 µm. g, Quantification of GFP+ beige adipocytes among total UCP1+ beige adipocytes in e and f. n = 8. Data are mean ± s.e.m. of biologically independent experiments. h, Schematic of the experiment. Myog-Cre;Rosa26-mTmG reporter mice were treated with β-blocker at room temperature for 5 days and subsequently acclimated at 15 ° C for 5 days. i, Immunofluorescent staining of GFP and UCP1 in the inguinal WAT of mice in h. Scale bar, 100 µm. a–f, i, tdTomato or DAPI for counter stain. The images represent three independent replicates.
Extended Data Fig. 3 Molecular analyses of Myod1+-derived beige adipocytes.
a, mRNA expression (in fragments per kilobase of transcript per million mapped reads (FPKM)) of Myh1, Myog, Fabp4, Adipoq, Ucp1 and Kcnk3 in indicated tissues. n = 3 biologically independent experiments. Data are mean ± s.e.m.; ANOVA followed by Tukey’s test. b, mRNA expression of beige fat markers in GFP+ and GFP− beige fat from Myod1-CreERT2 reporter mice. n = 3 biologically independent experiments. Data are mean ± s.e.m.; unpaired Student’s t-test. c, mRNA expression of glucose metabolism genes in indicated tissues. n = 3 biologically independent samples. Data are mean ± s.e.m., and analysed by ANOVA followed by Tukey’s test. d, mRNA expression of adrenergic receptors in GFP+ and GFP− beige fat. n = 3 biologically independent samples. Data are mean ± s.e.m.; unpaired two-sided Student’s t-test. e, Fatty acid oxidation in GFP+ and GFP− beige fat. n = 6 biologically independent samples. Data are mean ± s.e.m.; unpaired two-sided Student’s t-test.
Extended Data Fig. 4 MyoD+ progenitors in inguinal WAT.
a, Schematic of the experiment. BrdU was administered in Myod1-CreERT2 reporter mice during β-blocker treatment. b, Immunofluorescent staining of BrdU and GFP in the inguinal WAT-derived SVFs from Myod1-CreERT2 reporter mice. Scale bar, 50 µm. Enlarged image, scale bar, 10 µm. c, Schematic of the experiment. GFP+ and GFP− progenitors were isolated from lineage-negative (Lin−) stromal cells in the inguinal WAT of Myod1-CreERT2 reporter mice by fluorescence-activated cell sorting. d, Transcriptome analysis in c. Each type of progenitor was pooled from 3 mice. Cut-off values was P < 0.05 by the delta-method-based hypothesis test. e, PCA of transcriptome dataset from indicated cells. Transcriptome of FAPs (GSE86073) is included in the analysis. f, Immunofluorescent staining of MyoD in isolated GFP+ progenitors in c. Note that all the GFP+ cells express MyoD protein. Scale bar, 100 µm. g, Immunofluorescent staining of MyoD and PAX7 in the SVFs from the inguinal WAT of Pax7-CreERT2;Rosa26-tdTomato reporter mice. Pax7 lineage cells were marked with tdTomato. Scale bar, 20 µm. h, Immunofluorescent staining of MyoD and PAX7 in the SVFs from iBAT of Pax7-CreERT2;Rosa26-tdTomato reporter mice in g. Scale bar, 10 µm. b, f–h, DAPI was used as counter stain. Images represent three independent experiments.
Extended Data Fig. 5 Isolation of MyoD+ progenitors from inguinal WAT.
a, Expression of CD29 and CD34 in the SVFs from Pdgfra-CreERT reporter mice. b, Sequential gating to isolate GFP+(PDGFRα+)CD34+CD29+ cells in the SVFs from the inguinal WAT of Pdgfra-CreERT reporter mice treated with β-blocker. c, Immunofluorescent staining of MyoD in GFP+CD34+CD29+ cells isolated from the SVFs in b. DAPI was used as counter-stain. Scale bar, 25 µm. Data represent three independent experiments.
Extended Data Fig. 6 Developmental regulation of MyoD+ progenitors in the inguinal WAT.
a, mRNA expression of indicated genes in isolated GFP+ and GFP− progenitors from the inguinal WAT of Myod1-CreERT2 reporter mice treated with β-blocker. Smad5 (GFP−, n = 6; GFP+, n = 8), Myod1, Bmpr1a, Bmpr1b and Bmpr2 (n = 4). Data are mean ± s.e.m. of biologically independent samples; unpaired two-sided Student’s t-test. b, Schematic of the experiment. The inguinal WAT-derived SVF cells from Myod1-CreERT2 reporter mice were pre-treated with rBMP7 or vehicle for 2 days. Cells were subsequently differentiated under pro-adipogenic conditions for 8 days. c, Immunofluorescent staining of GFP and lipid droplets stained with Nile red in differentiated MyoD+ cells prepared as in b. Cells were treated with indicated compounds, including Br-cAMP (200 µM), 3-isobutyl-1-methylxanthine (IBMX, 0.5 µM), adenosine (100 nM), agonists for α1-AR (phenylephrine, 10 µM), α2-AR (clonidine, 10 µM), β1-AR (denopamine, 10 µM), β2-AR agonist (formoterol, 2.5 µM), β3-AR (CL316,243, 0.1 µM), noradrenaline (1 µM), and recombinant human BMP7 (rhBMP7, 3.3 nM), or vehicle control. DAPI was used as counter stain. The images represent three independent experiments. Scale bar, 100 µm.
Extended Data Fig. 7 Transcriptional regulation of g-beige adipocyte differentiation.
a, mRNA expression of Gabpa, Erra, and Errg in GFP+ and GFP− progenitors in the inguinal WAT of Myod1-CreERT2 reporter mice treated with β-blocker. n = 4. b, mRNA expression of indicated genes in GFP+ and GFP− beige fat from Myod1-CreERT2 reporter mice. n = 3. c, Protein expression of PRDM16, EHMT1 and CKIIα in GFP+ and GFP− progenitors in the inguinal WAT of Myod1-CreERT2 reporter mice. β-actin was used as a loading control. Molecular mass (kDa) is shown on the right. d, mRNA expression of indicated genes in C2C12 myoblasts expressing empty vector, GABPα, ERRα or ERRγ using lentivirus. n = 3. e, mRNA expression of Myod1 in C2C12 myoblasts expressing an empty vector, GABPα, ERRα or ERRγ by lentivirus. n = 4. f, Protein expression of PPARγ and UCP1 in differentiated C2C12 cells expressing an empty vector or GABPα in pro-adipogenic conditions. β-actin was used as a loading control. Blots represent five independent samples. g, mRNA expression of Prdm16 in C2C12 myoblasts expressing empty vector, GABPα, ERRα or ERRγ using lentivirus. Differentiated immortalized beige adipocytes are included as a reference. n = 4. a, b, Data are mean ± s.e.m. of biologically independent replicates; unpaired two-sided Student’s t-test. d, e, g, Data are mean ± s.e.m. of biologically independent replicates; ANOVA followed by Tukey’s test.
Extended Data Fig. 8 GABPα controls g-beige fat development.
a, Glucose uptake in differentiated cells expressing an empty vector or GABPα, and in immortalized beige adipocytes. The values were normalized to total protein concentration. n = 5. b, OCR of differentiated C2C12 cells expressing empty vector, GABPα or PRDM16. n = 3. c, mRNA expression of Gabpa in differentiated cells expressing a scramble control (sh-SCR) or shRNAs targeting Gabpa (shGabpa#1 and shGabpa#2). n = 4. d, Immunofluorescent staining of lipid droplets, MHC and GFP in differentiated C2C12 cells expressing sh-SCR or shGabpa. Scale bar, 50 µm. DAPI was used as counter stain. Images represent three independent experiments. e, mRNA expression of indicated genes in differentiated cells expressing sh-SCR or shGabpa. Ucp1, n = 5; Adipoq, Fabp4, Myh1, Myod1, n = 6. f, OCR in differentiated cells in c. n = 14. g, ECAR in differentiated cells in c. n = 14. a–c, e, Data are mean ± s.e.m. of biologically independent samples; one-way ANOVA followed by Tukey’s test. f, g, Data are mean ± s.e.m. of biologically independent samples; two-way ANOVA followed by Bonferroni’s test.
Extended Data Fig. 9 A mouse model of g-beige fat depletion.
a, Schematic of the experiment. Myod1-DTR+ and control mice were pre-treated with β-blocker and tamoxifen, and subsequently acclimated to 15 °C for 5 days. Diphtheria toxin (DT) was administered to deplete cold-induced Myod1+-derived g-beige fat. b,18F-FDG PET/CT images of vehicle- or β-blocker-treated mice at 15 °C. c, Schematic of tissue temperature recording in iBAT and skeletal muscle of mice. d, Changes in tissue temperature (ΔT) in iBAT and skeletal muscle. Mice were treated with saline (control) or β-blocker for 5 days (chronic β-blocker). A subset of the saline-treated mice was acutely treated with β-blocker (acute β-blocker). To stimulate thermogenesis, noradrenaline (NE) was administered at the indicated time point (black arrow). n = 4 for control, n = 6 for acute β-blocker treatment and n = 6 for chronic β-blocker treatment. Data are mean ± s.e.m. of biologically independent mice; two-way ANOVA followed by Bonferroni’s test. e, EMG measurement of skeletal muscle shivering in wild-type mice treated with β-blocker or vehicle (saline) at 30 °C or 15 °C. The shivering data were converted to the root mean square (RMS, µV). n = 4 biologically independent mice. Data are mean ± s.e.m.; ANOVA followed by Tukey’s test. f, H&E staining (left) and immunostaining of UCP1 (middle) or ENO1 (right) in the inguinal WAT of control and Myod1-DTR+ mice. Scale bar, 100 µm; enlarged image scale bar, 20 µm. Images represent five independent animals. g, EMG measurement of skeletal muscle shivering in control mice and Myod1-DTR+ mice at 30 °C or 15 °C. h, Quantification of data in g converted to RMS (µV). n = 6 biologically independent mice. Data are mean ± s.e.m.; ANOVA followed by Tukey’s test.
Extended Data Fig. 10 Requirement of g-beige fat for adaptive thermogenesis in the absence of β-AR signalling.
a, Schematic of the experiment. PpargMyod1-KO mice (Myod1-CreERT2;Ppargflox/flox) and littermate control mice (Ppargflox/flox) were pre-treated with β-blocker and tamoxifen. Subsequently, these mice were acclimated to 15 °C for 5 days. b, Immunofluorescent staining of UCP1 and ENO1 in the inguinal WAT of PpargMyod1-KO mice and controls. Scale bar, 100 µm. Images represent three independent experiments. c, Quantification of glycolytic beige fat in b. n = 10. d, OCR in the inguinal WAT of PpargMyod1-KO mice and littermate controls. n = 6. e, EMG measurement of skeletal muscle shivering in PpargMyod1-KO (n = 6) and littermate control mice (n = 5) at 30 °C or 15 °C. The shivering data were converted to RMS (µV). Data are mean ± s.e.m. of biologically independent mice; ANOVA followed by Tukey’s test. f, Whole-body oxygen consumption (VO2) in PpargMyod1-KO mice and littermate controls. Mice were treated with β-blocker at indicated time points at 15 °C. n = 5 for PpargMyod1-KO mice, n = 6 for control. Data are expressed as mean ± s.e.m. of biologically independent mice; ANOVA followed by Bonferroni’s test. g, Total food intake in f. h, Locomotor activity in f. i, Body weight of PpargMyod1-KO mice and littermate controls on a regular chow diet. Mice were treated with β-blocker and acclimated to 15 °C for 5 days. PpargMyod1-KO mice, n = 5; littermate control mice, n = 7. c, d, g–i, Data are mean ± s.e.m. of biologically independent samples; unpaired two-sided Student’s t-test.
Supplementary information
Supplementary Information
This file contains Supplementary Table 1, which contains a list of primer sequences used for qRT-PCR and Supplementary Figure 1, which contains the uncropped western blot films in Extended Fig.7c and 7f.
Video 1
SVF cells isolated from the inguinal WAT of Myod-CreERT2 reporter mice following β-blocker treatment. Myod-derived cells formed GFP+ (green) myotubes and actively twitched even under pro-adipogenic conditions. Scale bar=100µm.
Rights and permissions
About this article
Cite this article
Chen, Y., Ikeda, K., Yoneshiro, T. et al. Thermal stress induces glycolytic beige fat formation via a myogenic state. Nature 565, 180–185 (2019). https://doi.org/10.1038/s41586-018-0801-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0801-z
This article is cited by
-
Ubiquitin ligase RNF20 coordinates sequential adipose thermogenesis with brown and beige fat-specific substrates
Nature Communications (2024)
-
The Different Shades of Thermogenic Adipose Tissue
Current Obesity Reports (2024)
-
Fibroblastic reticular cells in lymph node potentiate white adipose tissue beiging through neuro-immune crosstalk in male mice
Nature Communications (2023)
-
Age-dependent Pdgfrβ signaling drives adipocyte progenitor dysfunction to alter the beige adipogenic niche in male mice
Nature Communications (2023)
-
Blockage of PPARγ T166 phosphorylation enhances the inducibility of beige adipocytes and improves metabolic dysfunctions
Cell Death & Differentiation (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.