Abstract
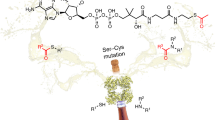
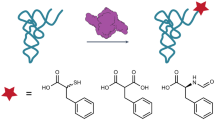
Many enzymes catalyse reactions that proceed through covalent acyl-enzyme (ester or thioester) intermediates1. These enzymes include serine hydrolases2,3 (encoded by one per cent of human genes, and including serine proteases and thioesterases), cysteine proteases (including caspases), and many components of the ubiquitination machinery4,5. Their important acyl-enzyme intermediates are unstable, commonly having half-lives of minutes to hours6. In some cases, acyl-enzyme complexes can be stabilized using substrate analogues or active-site mutations but, although these approaches can provide valuable insight7,8,9,10, they often result in complexes that are substantially non-native. Here we develop a strategy for incorporating 2,3-diaminopropionic acid (DAP) into recombinant proteins, via expansion of the genetic code11. We show that replacing catalytic cysteine or serine residues of enzymes with DAP permits their first-step reaction with native substrates, allowing the efficient capture of acyl-enzyme complexes that are linked through a stable amide bond. For one of these enzymes, the thioesterase domain of valinomycin synthetase12, we elucidate the biosynthetic pathway by which it progressively oligomerizes tetradepsipeptidyl substrates to a dodecadepsipeptidyl intermediate, which it then cyclizes to produce valinomycin. By trapping the first and last acyl-thioesterase intermediates in the catalytic cycle as DAP conjugates, we provide structural insight into how conformational changes in thioesterase domains of such nonribosomal peptide synthetases control the oligomerization and cyclization of linear substrates. The encoding of DAP will facilitate the characterization of diverse acyl-enzyme complexes, and may be extended to capturing the native substrates of transiently acylated proteins of unknown function.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Source data for all figures are available from the corresponding authors upon reasonable request. The models and structure factors for the crystal structures are deposited in the Protein Data Bank with accession numbers 6ECB, 6ECC, 6ECD, 6ECE and 6ECF. Detailed methods, including chemical syntheses, are available in the Supplementary Information.
References
Holliday, G. L., Mitchell, J. B. O. & Thornton, J. M. Understanding the functional roles of amino acid residues in enzyme catalysis. J. Mol. Biol. 390, 560–577 (2009).
Hedstrom, L. Serine protease mechanism and specificity. Chem. Rev. 102, 4501–4524 (2002).
Long, J. Z. & Cravatt, B. F. The metabolic serine hydrolases and their functions in mammalian physiology and disease. Chem. Rev. 111, 6022–6063 (2011).
Otto, H. H. & Schirmeister, T. Cysteine proteases and their inhibitors. Chem. Rev. 97, 133–172 (1997).
Swatek, K. N. & Komander, D. Ubiquitin modifications. Cell Res. 26, 399–422 (2016).
Yang, W. & Drueckhammer, D. G. Understanding the relative acyl-transfer reactivity of oxoesters and thioesters: computational analysis of transition state delocalization effects. J. Am. Chem. Soc. 123, 11004–11009 (2001).
Liu, B., Schofield, C. J. & Wilmouth, R. C. Structural analyses on intermediates in serine protease catalysis. J. Biol. Chem. 281, 24024–24035 (2006).
Scaglione, J. B. et al. Biochemical and structural characterization of the tautomycetin thioesterase: analysis of a stereoselective polyketide hydrolase. Angew. Chem. Int. Ed. 49, 5726–5730 (2010).
Cappadocia, L. & Lima, C. D. Ubiquitin-like protein conjugation: structures, chemistry, and mechanism. Chem. Rev. 118, 889–918 (2018).
Plechanovová, A., Jaffray, E. G., Tatham, M. H., Naismith, J. H. & Hay, R. T. Structure of a RING E3 ligase and ubiquitin-loaded E2 primed for catalysis. Nature 489, 115–120 (2012).
Chin, J. W. Expanding and reprogramming the genetic code of cells and animals. Annu. Rev. Biochem. 83, 379–408 (2014).
Magarvey, N. A., Ehling-Schulz, M. & Walsh, C. T. Characterization of the cereulide NRPS alpha-hydroxy acid specifying modules: activation of alpha-keto acids and chiral reduction on the assembly line. J. Am. Chem. Soc. 128, 10698–10699 (2006).
Lan, Y. et al. Incorporation of 2,3-diaminopropionic acid into linear cationic amphipathic peptides produces pH-sensitive vectors. ChemBioChem 11, 1266–1272 (2010).
Radzicka, A. & Wolfenden, R. Rates of uncatalyzed peptide bond hydrolysis in neutral solution and the transition state affinities of proteases. J. Am. Chem. Soc. 118, 6105–6109 (1996).
Reimer, J. M., Haque, A. S., Tarry, M. J. & Schmeing, T. M. Piecing together nonribosomal peptide synthesis. Curr. Opin. Struct. Biol. 49, 104–113 (2018).
Horsman, M. E., Hari, T. P. A. & Boddy, C. N. Polyketide synthase and non-ribosomal peptide synthetase thioesterase selectivity: logic gate or a victim of fate? Nat. Prod. Rep. 33, 183–202 (2016).
Hoyer, K. M., Mahlert, C. & Marahiel, M. A. The iterative gramicidin s thioesterase catalyzes peptide ligation and cyclization. Chem. Biol. 14, 13–22 (2007).
Jaitzig, J., Li, J., Süssmuth, R. D. & Neubauer, P. Reconstituted biosynthesis of the nonribosomal macrolactone antibiotic valinomycin in Escherichia coli. ACS Synth. Biol. 3, 432–438 (2014).
Akey, D. L. et al. Structural basis for macrolactonization by the pikromycin thioesterase. Nat. Chem. Biol. 2, 537–542 (2006).
Bruner, S. D. et al. Structural basis for the cyclization of the lipopeptide antibiotic surfactin by the thioesterase domain SrfTE. Structure 10, 301–310 (2002).
Samel, S. A., Wagner, B., Marahiel, M. A. & Essen, L. O. The thioesterase domain of the fengycin biosynthesis cluster: a structural base for the macrocyclization of a non-ribosomal lipopeptide. J. Mol. Biol. 359, 876–889 (2006).
Li, J. et al. Palladium-triggered deprotection chemistry for protein activation in living cells. Nat. Chem. 6, 352–361 (2014).
Baker, A. S. & Deiters, A. Optical control of protein function through unnatural amino acid mutagenesis and other optogenetic approaches. ACS Chem. Biol. 9, 1398–1407 (2014).
Virdee, S., Ye, Y., Nguyen, D. P., Komander, D. & Chin, J. W. Engineered diubiquitin synthesis reveals Lys29-isopeptide specificity of an OTU deubiquitinase. Nat. Chem. Biol. 6, 750–757 (2010).
Tseng, C. C. et al. Characterization of the surfactin synthetase C-terminal thioesterase domain as a cyclic depsipeptide synthase. Biochemistry 41, 13350–13359 (2002).
Trauger, J. W., Kohli, R. M., Mootz, H. D., Marahiel, M. A. & Walsh, C. T. Peptide cyclization catalysed by the thioesterase domain of tyrocidine synthetase. Nature 407, 215–218 (2000).
Zhou, Y., Prediger, P., Dias, L. C., Murphy, A. C. & Leadlay, P. F. Macrodiolide formation by the thioesterase of a modular polyketide synthase. Angew. Chem. Int. Ed. 54, 5232–5235 (2015).
Frueh, D. P. et al. Dynamic thiolation-thioesterase structure of a non-ribosomal peptide synthetase. Nature 454, 903–906 (2008).
Whicher, J. R. et al. Structure and function of the RedJ protein, a thioesterase from the prodiginine biosynthetic pathway in Streptomyces coelicolor. J. Biol. Chem. 286, 22558–22569 (2011).
Ekici, O. D., Paetzel, M. & Dalbey, R. E. Unconventional serine proteases: variations on the catalytic Ser/His/Asp triad configuration. Protein Sci. 17, 2023–2037 (2008).
Kavran, J. M. et al. Structure of pyrrolysyl-tRNA synthetase, an archaeal enzyme for genetic code innovation. Proc. Natl Acad. Sci. USA 104, 11268–11273 (2007).
Liu, Y., Zheng, T. & Bruner, S. D. Structural basis for phosphopantetheinyl carrier domain interactions in the terminal module of nonribosomal peptide synthetases. Chem. Biol. 18, 1482–1488 (2011).
Acknowledgements
We thank P. Emsley and G. Murshudov for their help in modelling depsipeptides; A. Wahba, J. Reimer, G. Bridon, K. Heesom and T. Elliott for help with mass spectrometry; M. Tarry and N. Rogerson for editing; C. Alonso for early crystal trials; R. Hay for early discussions on DAP; and S. Zhang for help with cloning. We also thank the staff of beamlines CLS 08ID-1 (S. Labiuk, J. Gorin, M. Fodje, K. Janzen, D. Spasyuk and P. Grochulski) and APS 24-ID-C (grants GM124165, RR029205, DE-AC02-06CH11357; F. Murphy). This work is supported by grants to J.W.C. from the Medical Research Council, UK (grants MC_U105181009 and MC_UP_A024_1008), to T.M.S. from the Canada Research Chair and the Natural Sciences and Engineering Research Council of Canada (NSERC; Discovery Grant 418420) and to C.N.B. from the NSERC (Discovery Grant 06167).
Author information
Authors and Affiliations
Contributions
N.H.-D. and D.A.A. contributed equally. M.M and G.W.H. contributed equally. N.H.-D. and D.P.N. selected the synthetases for 2–6. N.H.-D. characterized the incorporation of 6 and its deprotection to produce DAP, and performed the TEV conjugation. D.P.N and M.M. designed, and M.M. synthesized 2–6. T.M.S. and D.A.A. designed, and D.A.A. performed, the biochemistry and crystallography with Vlm TEwt. D.A.A. and N.H.-D. performed biochemistry and crystallography with Vlm TEDAP. G.W.H. and M.H.D. synthesized Vlm TE substrates. J.W.C. supervised N.H.-D., D.P.N. and M.M. T.M.S. supervised D.A.A. C.N.B. supervised G.W.H. and M.H.D. T.M.S., D.A.A., N.H.-D. and J.W.C. wrote the paper with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Schematic representation and reaction cycle of a canonical NRPS.
a, Schematic representation of a generic type I NRPS. The square brackets denote a single module. b, i–vii, Synthetic cycle of a canonical elongation module. NRPSs assemble peptides from amino acyl and other small acyl building blocks using a modular and thio-templated logic. A canonical NRPS is composed of one module for every residue in the peptide product. The initiation module contains an adenylation (A) domain, which binds cognate acyl substrate and performs adenylation and transfer of that substrate as a thioester on the phosphopantetheine arm (PPE, shown as a wavy line) of a peptidyl carrier protein (PCP) domain, for transport between active sites. Each elongation module contains an A and a PCP domain, and also a condensation (C) domain, which condenses aminoacyl and peptidyl substrates bound to PCP domains, thus progressively elongating the nascent chain. Termination modules contain C, A and PCP domains, and a specialized terminating/offloading domain responsible for the release of the peptide in its final form. The most common and most versatile terminating domain in NRPSs is the TE domain. Similar TE domains terminate synthesis in polyketide and fatty acid synthases. PPi, diphosphate; aa, amino acid.
Extended Data Fig. 2 Genetically directing DAP incorporation in recombinant proteins.
a, Structure of DAP and the protected versions investigated herein. 1, 2,3-diaminopropionic acid (DAP); 2, (S)-3-(((allyloxy)carbonyl)amino)-2-aminopropanoic acid; 3, (S)-2-amino-3-((2-nitrobenzyl)amino)propanoic acid; 4, (2S)-2-amino-3-((1-(6-nitrobenzo[d][1,3]dioxol-5-yl)ethyl)amino)propanoic acid; 5, (2S)-2-amino-3-(((1-(6-nitrobenzo[d][1,3]dioxol-5-yl)ethoxy)carbonyl)amino)propanoic acid; 6, (2S)-2-amino-3-(((2-((1-(6-nitrobenzo[d][1,3]dioxol-5-yl)ethyl)thio)ethoxy)carbonyl)amino)propanoic acid. Calculated logP values are indicated (calculated using the Molinspiration molecular property calculation services at www.molinspiration.com/cgi-bin/properties). b–f, Determining the intracellular concentration of compounds 2–6 by an LC–MS assay, performed on extracts. The dark-blue trace represents a 100 µM standard for each compound. The light-blue trace represents a 10 µM standard for each compound. The red trace results from cells grown in the absence of the compound. The brown trace results from cells grown in the absence of the compound, but spiked with the compound to 100 µM. The green trace results from cells grown in the presence of 1 mM compound. The experiments were repeated in two biological replicates with similar results. g, Phenotyping of the DAPRS/tRNACUA pair. Cells containing the DAPRS/tRNACUA pair and cat(112TAG) (encoding a chloramphenicol-resistance gene containing an amber stop codon (TAG) at codon 112) were plated in the presence or absence of 6 on the indicated concentrations of chloramphenicol. The experiment was performed in two biological replicates with similar results. h, The side chain of 6 (grey sticks) was modelled into the active site of PylRS using a co-crystal structure of PylRS and adenylated pyrrolysine (PDB accession number 2ZIM31). PylRS is displayed in pale yellow and amino-acid positions randomized in DAPRSlib are shown in marine blue.
Extended Data Fig. 3 Stably trapping the acyl-enzyme intermediate of a cysteine protease.
a, Different variants of TEV protease (shown at the top) were reacted with Ub–tev–His. The use of TEV(wt) results in cleavage of the TEV cleavage sequence. The use of TEV(C151A) results in minimal cleavage. The presence of DAP in the active site of TEV results in the presence of an extra band in the Coomassie gel, representing the isopeptide-linked TEV(C151DAP)–Ub complex. b, c, Anti-streptavidin (α-strep; b) and anti-Ub (α-Ub antibody P4D1; c) western blots of the reactions confirm the identity of the complex. For a–c, the experiment was repeated in two biological replicates with similar results. d, Tandem mass spectrometry following tryptic digest of the TEV(C151DAP)–Ub conjugate confirms amide-bond formation at the expected position. Top, the sequence of the branched peptide subject to fragmentation. Fragmentation of the substrate chain is predicted to lead to a series of y ions (yellow) and a series of b ions (green); the ions from this chain are labelled as ‘β’. Fragmentation of the TEV(C151DAP)-derived chain is predicted to lead to a series of y ions (blue) and a series of b ions (red); the ions from this chain are labelled as ‘α’. Bottom, MS/MS spectra with peak assignments. Ions in the α-chain were assigned by treating DAP and the β-chain as a modification of known mass. Ions in the β-chain were manually assigned. The mass-spectrometry analysis was performed once.
Extended Data Fig. 4 Chemical structures of key Vlm TE substrates and products.
The chemical structures and the numbers used to refer to them are shown.
Extended Data Fig. 5 The mechanism by which by Vlm TE catalyses oligomerization.
Oligomerization could conceivably take place in two ways. a, In the first scenario, ‘forward transfer’, the distal hydroxyl group of the tetradepsipeptidyl–O-TE complex attacks the thioester group in the tetradepsipeptidyl–S-PCP enzyme intermediate, directly forming octadepsipeptidyl–O-TE as a product. b, In the second scenario, ‘reverse transfer’, the distal hydroxyl group of the tetradepsipeptidyl–S-PCP complex attacks the ester group in the tetradepsipeptidyl–O-TE enzyme intermediate, forming octadepsipeptidyl–S-PCP as a product, which would then need to be transferred onto the TE-domain serine (here labelled as ‘re-capture’). c, d, Analogous scenarios involving tetradepsipeptidyl–SNAC (7) as the substrate instead of tetradepsipeptidyl–S-PCP. e, f, EICs (HR LC–ESI-MS) of a mix of 7 (1.7 mM) and buffer (e), or the products of a reaction between 7 (1.7 mM) and Vlm TEDAP (6.5 μM) (f). g–i, EICs (low-resolution (LR) LC–ESI-MS) of reactions using a higher-volume injection into an ion-trap MS instrument. g, The higher-volume injection of a reaction of 7 (1.7 mM) and Vlm TEwt (6.5 μM) enabled detection of a peak consistent with the 20-mer depsipeptidyl–SNAC (24). h, LC ion-trap MS of the reaction of 7 (1.7 mM) and Vlm TEDAP (6.5 μM). i, Small amounts of the cyclic 16-mer depsipeptide 29 elute during post-run column clean-up of experiment shown in g. j, EICs (HR LC–ESI-MS) of products of reactions between Vlm TEwt (6.5 μM) and a mix of 7 and deoxy-tetradepsipeptidyl–SNAC (8; 1.7 mM of each). TEwt produces the intermediates deoxy-octadepsipeptidyl–SNAC (12), deoxy-dodecadepsipeptidyl–SNAC (16) and deoxy 16-mer depsipeptidyl–SNAC (20), confirming the reaction pathway shown in b. See ‘Supplementary Methods for Statistics and Reproducibility’ for accurate mass analysis and deviations from calculated m/z values of each compound. The experiments in e–i were repeated independently twice with similar results. Mass-spectrometry analysis of the experiment in j was performed once.
Extended Data Fig. 6 Structures of Vlm TEwt and tetradepsipeptidyl–TEDAP, and top-down LC–ESI-MS of Vlm TEwt.
a, Secondary-structure elements of Vlm TE; the naming is based on the convention for α/β-hydrolase proteins. b, Comparison of two TEwt structures (PDB accession numbers 6ECB and 6ECC). The active-site lid of the first structure (light grey) is nearly completely ordered, whereas the lid of second structure (dark grey) shows density for Lα3, Lα4 and Lα5 only. In the second structure, Lα3 is rotated 10° towards the active site. c, Deconvoluted mass spectra of TEwt incubated with different substrates. Solid line, buffer control: expected molecular mass 31,028.22 Da; observed 31,028.75 Da. Dashed line, TEwt incubated with tetradepsipeptidyl–SNAC: expected 31,028.22 Da (unmodified) and 31,399.44 Da (modified); observed 31,026.29 Da. Dotted line, TEwt incubated with valinomycin: expected 31,028.22 Da (unmodified) and 32,139.86 Da (modified); observed 31,027.01 Da. Experiments were repeated independently twice with similar results. d, Comparison of near-identical conformations of TEwt (light grey; 6ECB) and tetradepsipeptidyl–TEDAP (tan and dark grey; 6ECD).
Extended Data Fig. 7 Expression and substrate conjugation to Vlm TE containing DAP at position 2463.
a, Following expression and purification of Vlm TEDAP, the protein was loaded on an SDS–PAGE gel and Coomassie stained; the experiment was repeated in two biological replicates with similar results. b, The deprotection of 6 in TEDAP–strep was followed by ESI-MS analysis. Green trace, purified TEDAP–strep containing 6 at position 2463: expected mass 32,364.6 Da, observed 32,365.78 Da. Red trace, TEDAP–strep containing 6 at position 2463 following illumination to convert 6 to the intermediate: expected 32,171.56 Da, observed 32,168.48 Da; and further incubation (1 h, 4 °C) to convert the intermediate to product: expected 32,067.62 Da, observed 32,068 Da). Blue trace, TEDAP–strep containing 6 at position 2,463 following illumination (to convert 6 to the intermediate) and further incubation (10 h, 4 °C) to convert the intermediate to DAP (1): expected 32,067.62 Da; observed, 32,067.84 Da. The experiment was repeated in two biological replicates with similar results. c, Purified TEDAP after illumination and intermediate fragmentation: expected 31,027.24 Da, observed 31,026.95 Da and 31,131.82 Da. d, TEDAP incubated with tetradepsipeptidyl–SNAC 7: expected 31,027.24 Da (unmodified) and 31,398.69 Da (modified); observed 31,025.92 Da and 31,396.55 Da. The experiments in c, d were repeated independently twice with similar results.
Extended Data Fig. 8 Electron density of the active site of covalent depsipeptidyl–TEDAP complexes.
Unbiased mFo – DFc maps (green mesh, contoured at 2.5σ), calculated before depsipeptide residues were placed in the model. DAP (brown) and depsipeptide residues (cyan) are depicted as sticks. a, Tetradepsipeptidyl–TEDAP (PDB accession number 6ECD). b–g, Dodecadepsipeptidyl–TEDAP P1 space-group structure (6ECF), with crystallographically independent molecules A to F shown in sequential order. h, i, Dodecadepsipeptidyl–TEDAP H3 space group (6ECE), for crystallographically independent molecules A and B. j–l, Electron density of the active site of covalent depsipeptidyl–TEDAP complexes extends beyond modelled depsipeptides. Unbiased mFo – DFc maps (green mesh, contoured at 2.5σ), calculated before depsipeptide residues were placed in the model, for dodecadepsipeptidyl–TEDAP P1 space-group structure, with crystallographically independent molecules A, B and D in sequential order. The observed electron density that extends beyond the modelled depsipeptides (cyan sticks) could accommodate extra depsipeptide residues in different orientations. However, unambiguous modelling into this density could not be achieved.
Extended Data Fig. 9 Modelling of interaction between the PCP domain and TE domain and putative pathway.
a, Superimposition of dodecadepsipeptidyl–TEDAP with the structure of the EntF PCP–TE didomain32 (PDB accession number 3TEJ) shows the path of the PPE moiety to the active site. b, Hypothetical pathway for oligomerization and cyclization, starting from octadepsipeptidyl–TE. i, The position of Lα1 in the observed apo/tetradepsipeptide conformation promotes an extended peptide conformation. ii, The tetradepsipeptidyl–PCP accepts the octadepsipeptide onto its terminal hydroxyl, perhaps using a dodecadepsipeptide-like lid conformation which could accommodate the roughly 30-Å tetradepsipeptidyl–PPE bound to the PCP domain and guide it towards the active site. iii, The PCP domain presents the thioester for transfer back to serine 2463. iv, Finally, the lid conformation observed in the dodecadepsipeptide–TEDAP structures could help to curl the dodecadepsipeptide back towards serine 2463 for cyclization.
Supplementary information
Supplementary Figure 1
Full gels for figures are shown. The Figure number that the gels correspond to is indicated, the molecular size standards are annotated and the area shown in the relevant Figure is indicated by a colour coded outline.
Supplementary Information
This file contains Supplementary Methods & Supplementary Discussion 1 and 2. All experimental methods, including methods for chemical synthesis. Additional discussion of the direction of the oligomerization pathway catalyzed by Vlm TEwt, and additional discussion of a putative model of oligomerization and cyclization catalyzed by Vlm TE.
Supplementary Data 1
This file contains Supplementary Data 1. Published thioesterase domain structures. TE domain structures deposited in the PDB are listed and annotated as NRPS, PKS or hybrid systems. The release mechanism catalyzed by the domain is also annotated. Each structure was analyzed for the presence of ligands, the topology and movement of the lid and whether the released product or the biosynthetic pathway product is commercially available.
Supplementary Video 1
Animation of a feasible transition between observed conformations of the Vlm TE lid. Note that the apo/tetradepsipeptidyl-bound and dodecadepsipeptidyl-bound conformations observed in this study are each in a cluster of similar conformations, rather than on single conformation. For simplicity, one representative apo/tetradepsipeptidyl-bound and one representative dodecadepsipeptidyl-bound conformation was used in the making of this animation.
Supplementary Video 2
Animation of a second feasible transition between observed conformations of the Vlm TE lid. Note that the apo/tetradepsipeptidyl-bound and dodecadepsipeptidyl-bound conformations observed in this study are each in a cluster of similar conformations, rather than on single conformation. For simplicity, one representative apo/tetradepsipeptidyl-bound and one representative dodecadepsipeptidyl-bound conformation was used in the making of this animation.
Rights and permissions
About this article
Cite this article
Huguenin-Dezot, N., Alonzo, D.A., Heberlig, G.W. et al. Trapping biosynthetic acyl-enzyme intermediates with encoded 2,3-diaminopropionic acid. Nature 565, 112–117 (2019). https://doi.org/10.1038/s41586-018-0781-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0781-z
This article is cited by
-
Genetically encoded bioorthogonal tryptophan decaging in living cells
Nature Chemistry (2024)
-
Site-specific encoding of photoactivity and photoreactivity into antibody fragments
Nature Chemical Biology (2023)
-
Mechanism of d-alanine transfer to teichoic acids shows how bacteria acylate cell envelope polymers
Nature Microbiology (2023)
-
Genetically programmed cell-based synthesis of non-natural peptide and depsipeptide macrocycles
Nature Chemistry (2023)
-
Mechanism-based traps enable protease and hydrolase substrate discovery
Nature (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.