Abstract
Most patients with cancer either do not respond to immune checkpoint blockade or develop resistance to it, often because of acquired mutations that impair antigen presentation. Here we show that loss of function of the RNA-editing enzyme ADAR1 in tumour cells profoundly sensitizes tumours to immunotherapy and overcomes resistance to checkpoint blockade. In the absence of ADAR1, A-to-I editing of interferon-inducible RNA species is reduced, leading to double-stranded RNA ligand sensing by PKR and MDA5; this results in growth inhibition and tumour inflammation, respectively. Loss of ADAR1 overcomes resistance to PD-1 checkpoint blockade caused by inactivation of antigen presentation by tumour cells. Thus, effective anti-tumour immunity is constrained by inhibitory checkpoints such as ADAR1 that limit the sensing of innate ligands. The induction of sufficient inflammation in tumours that are sensitized to interferon can bypass the therapeutic requirement for CD8+ T cell recognition of cancer cells and may provide a general strategy to overcome immunotherapy resistance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data presented in this manuscript are available from the corresponding author upon reasonable request. Bulk tumour cell RNA sequencing has been deposited at the Gene Expression Omnibus (GEO) under accession number GSE110708. Single-cell RNA sequencing of tumour cells were also deposited at the GEO under accession number GSE110746.
References
Zaretsky, J. M. et al. Mutations associated with acquired resistance to PD-1 blockade in melanoma. N. Engl. J. Med. 375, 819–829 (2016).
Gao, J. et al. Loss of IFN-γ pathway genes in tumor cells as a mechanism of resistance to anti-CTLA-4 therapy. Cell 167, 397–404 (2016).
Yoshihama, S. et al. NLRC5/MHC class I transactivator is a target for immune evasion in cancer. Proc. Natl Acad. Sci. USA 113, 5999–6004 (2016).
Sade-Feldman, M. et al. Resistance to checkpoint blockade therapy through inactivation of antigen presentation. Nat. Commun. 8, 1136 (2017).
Manguso, R. T. et al. In vivo CRISPR screening identifies Ptpn2 as a cancer immunotherapy target. Nature 547, 413–418 (2017).
Mannion, N. M. et al. The RNA-editing enzyme ADAR1 controls innate immune responses to RNA. Cell Reports 9, 1482–1494 (2014).
Liddicoat, B. J., Chalk, A. M. & Walkley, C. R. ADAR1, inosine and the immune sensing system: distinguishing self from non-self. Wiley Interdiscip. Rev. RNA 7, 157–172 (2016).
Ahmad, S. et al. Breaching self-tolerance to Alu duplex RNA underlies MDA5-mediated inflammation. Cell 172, 797–810.e13 (2018).
Chung, H. et al. Human ADAR1 prevents endogenous RNA from triggering translational shutdown. Cell 172, P811–P824 (2018).
Deng, L. et al. STING-dependent cytosolic DNA sensing promotes radiation-induced type I interferon-dependent antitumor immunity in immunogenic tumors. Immunity 41, 843–852 (2014).
Lim, J. Y. H., Gerber, S. A., Murphy, S. P. & Lord, E. M. Type I interferons induced by radiation therapy mediate recruitment and effector function of CD8+ T cells. Cancer Immunol. Immunother. 63, 259–271 (2014).
Twyman-Saint Victor, C. et al. Radiation and dual checkpoint blockade activate non-redundant immune mechanisms in cancer. Nature 520, 373–377 (2015).
Wenzel, J., Uerlich, M., Haller, O., Bieber, T. & Tueting, T. Enhanced type I interferon signaling and recruitment of chemokine receptor CXCR3-expressing lymphocytes into the skin following treatment with the TLR7-agonist imiquimod. J. Cutan. Pathol. 32, 257–262 (2005).
Sainathan, S. K. et al. Toll-like receptor-7 ligand Imiquimod induces type I interferon and antimicrobial peptides to ameliorate dextran sodium sulfate-induced acute colitis. Inflamm. Bowel Dis. 18, 955–967 (2012).
Cho, J. H. et al. The TLR7 agonist imiquimod induces anti-cancer effects via autophagic cell death and enhances anti-tumoral and systemic immunity during radiotherapy for melanoma. Oncotarget 8, 24932–24948 (2017).
Patterson, J. B. & Samuel, C. E. Expression and regulation by interferon of a double-stranded-RNA-specific adenosine deaminase from human cells: evidence for two forms of the deaminase. Mol. Cell. Biol. 15, 5376–5388 (1995).
George, C. X. & Samuel, C. E. Human RNA-specific adenosine deaminase ADAR1 transcripts possess alternative exon 1 structures that initiate from different promoters, one constitutively active and the other interferon inducible. Proc. Natl Acad. Sci. USA 96, 4621–4626 (1999).
Heidegger, S. et al. The RIG-I agonist 3pRNA synergizes with checkpoint blockade in cancer immunotherapy. Blood 126, 3436 (2015).
Goel, S. et al. CDK4/6 inhibition triggers anti-tumour immunity. Nature 548, 471–475 (2017).
Chiappinelli, K. B. et al. Inhibiting DNA methylation causes an interferon response in cancer via dsRNA including endogenous retroviruses. Cell 162, 974–986 (2015).
Li, H. et al. Immune regulation by low doses of the DNA methyltransferase inhibitor 5-azacitidine in common human epithelial cancers. Oncotarget 5, 587–598 (2014).
Ribas, A. et al. Oncolytic virotherapy promotes intratumoral T cell infiltration and improves anti-PD-1 immunotherapy. Cell 170, 1109–1119 (2017).
Juneja, V. R. et al. PD-L1 on tumor cells is sufficient for immune evasion in immunogenic tumors and inhibits CD8 T cell cytotoxicity. J. Exp. Med. 214, 895–904 (2017).
McGranahan, N. et al. Allele-specific HLA loss and immune escape in lung cancer evolution. Cell 171, 1259–1271 (2017).
Tumeh, P. C. et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature 515, 568–571 (2014).
Ngwa, W., Tsiamas, P., Zygmanski, P., Makrigiorgos, G. M. & Berbeco, R. I. A multipurpose quality assurance phantom for the small animal radiation research platform (SARRP). Phys. Med. Biol. 57, 2575–2586 (2012).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Waltman, L. & van Eck, N. J. A smart local moving algorithm for large-scale modularity-based community detection. Eur. Phys. J. B 86, 471 (2013).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
DeTomaso, D. & Yosef, N. FastProject: a tool for low-dimensional analysis of single-cell RNA-Seq data. BMC Bioinformatics 17, 315 (2016).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Schmieder, R. & Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 27, 863–864 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Bazak, L. et al. A-to-I RNA editing occurs at over a hundred million genomic sites, located in a majority of human genes. Genome Res. 24, 365–376 (2014).
Porath, H. T., Carmi, S. & Levanon, E. Y. A genome-wide map of hyper-edited RNA reveals numerous new sites. Nat. Commun. 5, 4726 (2014).
Yu, G., Wang, L.-G. & He, Q.-Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Paz-Yaacov, N. et al. Elevated RNA Editing activity is a major contributor to transcriptomic diversity in tumors. Cell Reports 13, 267–276 (2015).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453–457 (2015).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013).
Acknowledgements
We thank all members of the Haining, Hur, Levanon and Meyerson laboratories for their input and discussions regarding this project.
Reviewer information
Nature thanks C. Walkley and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
W.N.H., J.J.I., R.T.M. and C.K.C. contributed to the study design. J.J.I., C.K.C., D.E.C., R.T.M., A.A., I.C.K., H.W.P., M.D.Z., S.K.L.-R., E.J.R. and A.H.L. performed validation experiments in cell lines and with live animals. C.K.C., A.I.-V. and R.T.M. carried out ADAR1 re-expression/rescue experiments. C.K.C., J.J.I., R.T.M. and M.D.Z. conducted epistasis experiments. A.P. and F.D.B. performed cell death experiments. J.J.I. and D.K. designed and performed radiation experiments. J.J.I., B.C.M. and G.K.G. conducted and analysed immunohistochemical experiments. J.J.I., J.D., A.I.-V. and B.C.M. conducted flow cytometry experiments, which B.C.M. and J.D. analysed. J.J.I. and B.C.M. prepared samples for single-cell RNA-seq, which K.B. analysed. J.J.I. and K.B.Y. prepared samples for RNA-seq analysis. K.B.Y., I.B., E.Y.L., A.P., P.P.D., K.B. and D.R.S. performed editing, RNA-seq and ATAC-seq analysis. J.G.D. performed screening processing and analysis. J.J.I., N.B.C., K.B. and A.P. undertook analysis of human tumours. J.J.I., R.T.M. and W.N.H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
This work was supported in part by funding from Calico Life Sciences, LLC. J.J.I., R.T.M. and W.N.H. are authors of a patent application related to ADAR. W.N.H. consults for and has equity in Tango Therapeutics.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Supporting evidence that ADAR1 loss enhances the response to immunotherapy.
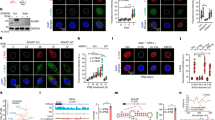
a, Expression of ADAR1 protein in control (grey), Adar1 p150-null (orange) and Adar1 p150/p110-null (red) B16 cells. Results are representative of three independent experiments. b, Tumour volume (left) and survival analysis (right) of control (grey), Adar1 p150-null (orange) or Adar1 p110/p150-null (red) B16 tumours in GVAX- and anti-PD-1-treated wild-type C57BL/6 mice. n = 5 animals per guide with two separate guides for the control group and at least two separate guides for each Adar1-null group. Data are representative of two independent experiments. c, Tumour volume and survival analysis of control (grey), Adar1 p150-null (orange) or Adar1 p110/p150-null (red) CT26 and Braf/Pten tumours in NSG, wild-type and wild-type anti-PD-1-treated mice. n = 5 mice per group; data are representative of two independent experiments. d, Survival analysis of control and Adar1-null MC38 tumours in wild-type and wild-type anti-PD-1-treated C57BL/6 mice. n = 5 animals per guide with two separate guides for the control group and three separate guides for the Adar1-null group. Data are representative of two independent experiments. e, Tumour volume and survival analysis of Adar1-null and control B16 tumours size matched at the time of PD-1 treatment initiation. b–e, Tumour volume curves are mean ± s.e.m and assessed with Student’s t-test; survival curves assessed with log-rank test, *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 2 Flow cytometry gating strategies and representative plots.
a, Gating strategy and representative flow cytometry plots for the assessment of CD4+, CD8+ and γδ T cells in Adar1-null and control B16 tumours. b, Gating strategy and representative flow cytometry plots for the assessment of NK cells in Adar1-null and control B16 tumours. c, Gating strategy and representative flow cytometry plots for the assessment of CD11b+Ly6c+ and CD11b+Ly6cloCD24+ cells in Adar1-null and control B16 tumours.
Extended Data Fig. 3 Further flow cytometry gating strategies and representative plots.
a, Gating strategy and representative flow cytometry plots for the assessment of granzyme B+CD4+ T cells in Adar1-null and control B16 tumours. b, Gating strategy and representative flow cytometry plots for the assessment of TAM1 and TAM2 populations in Adar1-null and control B16 tumours.
Extended Data Fig. 4 Single-cell RNA-seq extended data.
a, Gene expression matrix from single-cell RNA-seq experiment characterizing expression of lineage-defining genes in cell clusters. b, Key differentially expressed transcripts that distinguish cell clusters in Fig. 2. c, Paired quantile–quantile (Q–Q) plots comparing the expression of a curated set of genes in immune cells from Adar1-null and control tumours and matched t-SNEs depicting the distribution of gene expression for proinflammatory, suppressive and T cell activation/effector genes. P values calculated using Wilcoxon rank-sum test. d, Single-cell gene set enrichment scores of an IFNγ response signature score within individual immune subpopulations from Adar1-null and control tumours (P values calculated using Kolmogorov–Smirnov test). a, c, d, n = 7,406 cells. *P < 0.05; **P < 0.01; ***P < 0.001.
Extended Data Fig. 5 Further studies corroborating the reported in vitro phenotype of Adar1-null tumour cells.
a, Western blot demonstrating expression of ovalbumin in modified Adar1 p150/p110-null (red), Adar1 p150-null (orange) and control (grey) B16 tumour cell lines. Data are representative of two independent experiments. b, Calcein cell viability and 7-AAD cell death staining of control or Adar1-null B16 tumour cells following stimulation with IFNβ, IFNγ or a combination of both. Data are representative of three independent experiments with n = 3 for each condition. c, Growth and viability of Adar1 p150/p110-null, Adar1 p150-null and control B16 tumour cells in response to increasing doses of IFNβ and IFNγ (n = 3 for each condition). Doses are relative to 1× standard of 1,000 U ml–1 IFNβ and 100 ng ml–1 IFNγ. Data are representative of two independent experiments. d, Growth and viability of Adar1 p150/p110-null and control CT26 tumour cells following stimulation with IFNβ or IFNγ relative to the unstimulated state (n = 3 for each condition). Data are representative of two independent experiments. e, Growth and viability of Adar1 p150/p110-null and control Braf/Pten tumour cells following stimulation with TNF, IFNβ or IFNγ relative to the unstimulated state (n = 3 for each condition). Data are representative of two independent experiments. f, GSEA of gene signatures in Adar1-null compared with control B16 tumours cells after in vitro culture without cytokine stimulation. n = 3 for each condition; FDR calculated using GSEA. g, Heat map showing differentially expressed genes from Adar1-null and control B16 tumour cells 36 h after IFNβ stimulation in vitro (n = 3 for each condition). Genes listed in adjacent text were manually curated as antiviral or relevant to anti-tumour immunity. h, IFNβ ELISA of control and Adar1 p150/p110 CT26 tumour cells following stimulation with IFNβ or IFNγ (n = 3 for each condition). b–e, h, Two-sided Student’s t-test, *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 6 ADAR1 re-expression and corroborating in vitro epistatic IFN-signalling experiments.
a, Western blot of B16 Adar1-null tumour cells following re-expression of wild-type ADAR1 or an irrelevant control (CD19) protein. Data are representative of two independent experiments. b, IFNβ secretion (left) and relative growth (right) of control (grey), Adar1 p150/p110-null (red), Adar1-null with full-length ADAR1 re-expression construct (red outline) and control with ADAR1 re-expression construct (grey outline) B16 tumour cells following cytokine stimulation as indicated (n = 3 for each condition). Data are representative of two independent experiments. c, qPCR and western blot validation of the loss of expression of Ifnar2, Ifngr1 and Stat1 from B16 tumour cells used to generate the control and Adar1-null tumour cell lines shown in Fig. 3. n = 3 for qPCR experiments and data are representative of two independent experiments. d, Growth inhibition (left two panels) and IFNβ ELISA (right panel) of control and Adar1-null B16 tumour cells modified to delete Ifnar2, Ifngr1 or Stat1 (n = 3 for each condition; data representative of three independent experiments). e, IFNβ secretion in vitro following irradiation with 4 Gy in Adar1-null and control B16 tumour cells with and without IFNAR-blocking antibodies (left). Growth and viability of Adar1-null and control B16 tumour cells in vitro following irradiation with 4 Gy with and without IFNAR-blocking antibodies (right). For both plots: n = 3 for each condition; data are representative of two independent experiments. f, Survival analysis corresponding to the tumour volume curves depicted in Fig. 3h of Adar1 and control tumours treated with therapeutic irradiation. n = 10 mice for each group. Data are representative of two independent experiments. g, Tumour volume and survival analysis of control and Adar1-null B16 tumours treated with topical imiquimod. Data are representative of two independent experiments with n = 10 mice per group. b, e and tumour volume curves, two-sided Student’s t-test; survival curves, log-rank test, *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 7 Corroborating data for dsRNA editing and epistasis studies of Adar1-null tumours.
a, Genomic localization of SINEs (left) and detected editing sites within SINEs (right) in control B16 tumour cells. b, Representative tracks of ATAC-seq and RNA-seq mapped to SINEs and detected edits in IFN-inducible regions of accessible chromatin and transcription. c, Transcriptional upregulation of Adar1 and dsRNA sensors 36 h after stimulation with IFNβ or IFNγ in control B16 tumour cells as measured by RNA-seq (n = 3 for each condition). d, Volcano plot depicting the relative depletion and enrichment of sgRNAs targeting 20,146 genes in a Cas9+ Adar1-null B16 tumour cell line following stimulation with IFNγ in vitro. P values are derived using STARS v1.3. e, Western blots demonstrating the loss of expression of PKR, MDA5, RIG-I, MAVS and ADAR1 from double knockout and triple knockout B16 tumour cell lines. Data are representative of two independent experiments. f, IFNβ and IFNγ ELISAs from tumour lysate extracted from Adar1-null and control tumours that were epistatically deleted for dsRNA sensors including Eif2ak2 (PKR), Ifih1 (MDA5) or both (n = 5 for each condition). f, Two-sided Student’s t-test, *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 8 Correlations between RNA editing and signatures of immune infiltration in TCGA.
a, Pearson’s correlations between hyperediting index and Hallmark Inflammatory Response, CIBERSORT Absolute Immune Infiltrate, Hallmark Interferon Gamma Response, Hallmark Apoptosis and ESTIMATE immune infiltrate gene signatures from 356 tumours in TCGA for which hyperediting index information was available. b, Distribution of hyperediting index values of individual tumour types from the same samples from TCGA. Box plots represent the range, median, 25th and 75th percentile with n as indicated in the figure.
Extended Data Fig. 9 Corroborating data for models of immunotherapy resistance.
a, Western blot demonstrating the expression of ovalbumin in control and B2m-null B16 tumour cell lines depicted in Fig. 5c. Data are representative of two independent experiments. b, Quantitative PCR and western blots demonstrating loss of expression of B2m, Jak1, H2k1, Jak2, and Nlrc in B16 tumour cell lines used to make epistatically deleted Adar1-null or control tumour cells lines. n = 3 for qPCR experiments and data are representative of two independent experiments with P value calculated using two-sided Student’s t-test, *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 10 Loss of ADAR1 overcomes multiple common mechanisms of resistance to immunotherapy in vivo.
a, Survival analysis corresponding to tumour volume curves from Fig. 5c depicting the effect of Adar1 deletion in B16 tumours in which Jak1 or B2m have been epistatically deleted. n = 5 animals per group; data are representative of two independent experiments. b, Tumour volume curves in control (grey) and Adar1-null (red) B16 tumour cells epistatically deleted for H2k1, Jak2 and Nlrc5 and treated with GVAX and anti-PD-1 as indicated (n = 5 animals per group). c, Survival analysis corresponding to the tumour volume curves depicted in b. d, Additional TME characterization of control and Adar1-null B16 tumour cells epistatically deleted for Jak1 and B2m as indicated (n = 5 mice per group). Data in d and all tumour volume curves assessed with two-sided Student’s t-test; all survival curves assessed with log-rank test, *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Supplementary information
Supplementary Information
This file contains Supplementary Text, Supplementary References and Supplementary Figure 1, which contains the uncropped blots
Supplementary Tables
This file contains Supplementary Tables 1-6
Rights and permissions
About this article
Cite this article
Ishizuka, J.J., Manguso, R.T., Cheruiyot, C.K. et al. Loss of ADAR1 in tumours overcomes resistance to immune checkpoint blockade. Nature 565, 43–48 (2019). https://doi.org/10.1038/s41586-018-0768-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0768-9
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.