Abstract
Development and routine tissue homeostasis require a high turnover of apoptotic cells. These cells are removed by professional and non-professional phagocytes via efferocytosis1. How a phagocyte maintains its homeostasis while coordinating corpse uptake, processing ingested materials and secreting anti-inflammatory mediators is incompletely understood1,2. Here, using RNA sequencing to characterize the transcriptional program of phagocytes actively engulfing apoptotic cells, we identify a genetic signature involving 33 members of the solute carrier (SLC) family of membrane transport proteins, in which expression is specifically modulated during efferocytosis, but not during antibody-mediated phagocytosis. We assessed the functional relevance of these SLCs in efferocytic phagocytes and observed a robust induction of an aerobic glycolysis program, initiated by SLC2A1-mediated glucose uptake, with concurrent suppression of the oxidative phosphorylation program. The different steps of phagocytosis2—that is, ‘smell’ (‘find-me’ signals or sensing factors released by apoptotic cells), ‘taste’ (phagocyte–apoptotic cell contact) and ‘ingestion’ (corpse internalization)—activated distinct and overlapping sets of genes, including several SLC genes, to promote glycolysis. SLC16A1 was upregulated after corpse uptake, increasing the release of lactate, a natural by-product of aerobic glycolysis3. Whereas glycolysis within phagocytes contributed to actin polymerization and the continued uptake of corpses, lactate released via SLC16A1 promoted the establishment of an anti-inflammatory tissue environment. Collectively, these data reveal a SLC program that is activated during efferocytosis, identify a previously unknown reliance on aerobic glycolysis during apoptotic cell uptake and show that glycolytic by-products of efferocytosis can influence surrounding cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
RNA sequencing data presented in this study have been deposited in the NCBI GEO repository under the accession number GSE119273.
References
Arandjelovic, S. & Ravichandran, K. S. Phagocytosis of apoptotic cells in homeostasis. Nat. Immunol. 16, 907–917 (2015).
Elliott, M. R. & Ravichandran, K. S. The dynamics of apoptotic cell clearance. Dev. Cell 38, 147–160 (2016).
Colegio, O. R. et al. Functional polarization of tumour-associated macrophages by tumour-derived lactic acid. Nature 513, 559–563 (2014).
A-Gonzalez, N. et al. Phagocytosis imprints heterogeneity in tissue-resident macrophages. J. Exp. Med. 214, 1281–1296 (2017).
Cummings, R. J. et al. Different tissue phagocytes sample apoptotic cells to direct distinct homeostasis programs. Nature 539, 565–569 (2016).
Cesar-Razquin, A. et al. A call for systematic research on solute carriers. Cell 162, 478–487 (2015).
Lin, L., Yee, S. W., Kim, R. B. & Giacomini, K. M. SLC transporters as therapeutic targets: emerging opportunities. Nat. Rev. Drug Discov. 14, 543–560 (2015).
Mueckler, M. & Thorens, B. The SLC2 (GLUT) family of membrane transporters. Mol. Aspects Med. 34, 121–138 (2013).
Verdone, J. E., Zarif, J. C. & Pienta, K. J. Aerobic glycolysis, motility, and cytoskeletal remodeling. Cell Cycle 14, 169–170 (2015).
Macintyre, A. N. et al. The glucose transporter Glut1 is selectively essential for CD4 T cell activation and effector function. Cell Metab. 20, 61–72 (2014).
Chan, D. A. et al. Targeting GLUT1 and the Warburg effect in renal cell carcinoma by chemical synthetic lethality. Sci. Transl. Med. 3, 94ra70 (2011).
Park, D. et al. Continued clearance of apoptotic cells critically depends on the phagocyte Ucp2 protein. Nature 477, 220–224 (2011).
Jensen, P. J., Gitlin, J. D. & Carayannopoulos, M. O. GLUT1 deficiency links nutrient availability and apoptosis during embryonic development. J. Biol. Chem. 281, 13382–13387 (2006).
Kojima, Y., Weissman, I. L. & Leeper, N. J. The role of efferocytosis in atherosclerosis. Circulation 135, 476–489 (2017).
Zhang, S., Hulver, M. W., McMillan, R. P., Cline, M. A. & Gilbert, E. R. The pivotal role of pyruvate dehydrogenase kinases in metabolic flexibility. Nutr. Metab. 11, 10 (2014).
Palmada, M. et al. SGK1 kinase upregulates GLUT1 activity and plasma membrane expression. Diabetes 55, 421–427 (2006).
Elliott, M. R. et al. Nucleotides released by apoptotic cells act as a find-me signal to promote phagocytic clearance. Nature 461, 282–286 (2009).
Sprowl-Tanio, S. et al. Lactate/pyruvate transporter MCT-1 is a direct Wnt target that confers sensitivity to 3-bromopyruvate in colon cancer. Cancer Metab. 4, 20 (2016).
Biswas, S. K. & Mantovani, A. Macrophage plasticity and interaction with lymphocyte subsets: cancer as a paradigm. Nat. Immunol. 11, 889–896 (2010).
Kelly, B. & O’Neill, L. A. Metabolic reprogramming in macrophages and dendritic cells in innate immunity. Cell Res. 25, 771–784 (2015).
Gaber, T., Strehl, C. & Buttgereit, F. Metabolic regulation of inflammation. Nat. Rev. Rheumatol. 13, 267–279 (2017).
Han, C. Z. et al. Macrophages redirect phagocytosis by non-professional phagocytes and influence inflammation. Nature 539, 570–574 (2016).
Pollard, J.W. & Stanners, C.P. Characterization of cell lines showing growth control isolated from both the wild type and a leucyl-tRNA synthetase mutant of Chinese hamster ovary cells. J. Cell. Physiol. 98, 571–585 (1979).
Fond, A. M., Lee, C. S., Schulman, I. G., Kiss, R. S. & Ravichandran, K. S. Apoptotic cells trigger a membrane-initiated pathway to increase ABCA1. J. Clin. Invest. 125, 2748–2758 (2015).
Ahn, Y. Y., Bagrow, J. P. & Lehmann, S. Link communities reveal multiscale complexity in networks. Nature. 466, 761–764 (2010).
Chen, S. et al. Genome-wide CRISPR screen in a mouse model of tumor growth and metastasis. Cell. 160, 124–1260 (2015).
Takanaga, H., Chaudhuri, B. & Frommer, W. B. GLUT1 and GLUT9 as major contributors to glucose influx in HepG2 cells identified by a high sensitivity intramolecular FRET glucose sensor. Biochim. Biophys. Acta. 1778, 1091–1099 (2008).
Macintyre, A. N. et al. The glucose transporter Glut1 is selectively essential for CD4 T cell activation and effector function. Cell Metab. 20, 61–72 (2014).
Michalek, R. D. et al. Cutting edge: distinct glycolytic and lipid oxidative metabolic programs are essential for effector and regulatory CD4+ T cell subsets. J. Immunol. 186, 3299–3303 (2011).
Acknowledgements
This work is supported by R35GM122542, P01HL120840, and UVA Center for Cell Clearance (K.S.R.), American Heart Association 13BGIA17070106 and UTHSC funds (L.M.), Mishima Kaiun Memorial Foundation and Kanae Foundation (S.M.), CRI–Mark Foundation Fellowship, 5T32CA009109-39 (J.S.A.P.), Neuroscience Training Program (M.H.R.), and Erik and Mabel Johansson Scholarship (L.Z.).
Reviewer information
Nature thanks L. O’Neill and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
S.M. and J.S.A.P. designed and performed most experiments, with input from K.S.R. M.H.R., C.B.M., V.S., N.L., S.O.-G., J.C.R., Y.Z., S.K., L.Z. and L.M. performed and/or assisted with specific experiments. S.M., J.S.A.P. and K.S.R. wrote the manuscript with input from co-authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 RNA preparation for RNA-seq experiments.
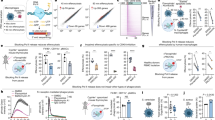
a, Representative fluorescence-activated cell sorting plots of engulfment assays with LR73 hamster phagocytes (left) and annexin V/7-AAD staining of apoptotic human Jurkat cells (right) in conditions matching experiments performed for RNA-seq (2 h with apoptotic cells followed by 2 h rest in the absence of apoptotic cells). b, Principal component analysis on hamster-genome-aligned RNA-seq data as a quality control statistic.
Extended Data Fig. 2 Regulation of SLC expression during efferocytosis.
a, SLC genes are differentially regulated during efferocytosis. Left, plot of the 165 SLC genes detected by RNA-seq of efferoytic LR73 cells, highlighting the 19 significantly upregulated (red) and 14 downregulated (blue) SLC genes that were altered during efferocytosis. The 132 SLC genes that were not altered are located on the midline (black). Right, the current genetic classifications of these 33 SLC genes that are altered during engulfment are shown. b, Efferocytosis-associated SLCs and their properties. Current genetic classification and/or functional linkages of the 33 SLCs modulated during apoptotic cell engulfment. The significantly upregulated and downregulated SLCs and the substrates they are known to transport grouped by predicted general function are shown, as are the known monogenic diseases and single nucleotide polymorphism (SNP) or disease phenotype to which the specific SLCs have been linked.
Extended Data Fig. 3 qRT–PCR confirmation of the RNA-seq data.
a, qRT–PCR of mRNA of specific SLCs during efferocytosis. Indicated SLC genes were tested for mRNA expression levels during engulfment assays performed similarly to those in Fig. 1a. Data are representative of at least two independent experiments with 3–4 replicates per condition. b, The table presents the cycle numbers for each species-specific qRT–PCR primer. None of these primers produced signals when tested against human Jurkat cell mRNA (target) alone.
Extended Data Fig. 4 Dynamic expression of SLCs during efferocytosis.
a, Schematic of the experiment and time points when RNA from phagocytes was assessed for specific SLC gene expression. Apoptotic Jurkat cells were added to LR73 cells and co-cultured for 2 h. Unbound and floating apoptotic cells were then washed away, and the LR73 cells were cultured in fresh medium for the indicated times. The time scale bar reflects total time of experiment, such that the 4-h time point reflects 2 h with apoptotic cells plus 2 h subsequent incubation (to match the timeframe used in our RNA-seq experiment). Total RNA was subsequently isolated and qRT–PCR was performed for specific SLC genes. Flow cytometry plots indicate that fluorescent signals from the internalized corpses are significantly degraded by the 8 h time point. b, Expression of SLC genes is regulated over the time course of efferocytosis. Relative expression of mRNAs for specific SLC genes belonging to different functional classes over the time course of engulfment is shown. Data are representative of three biological replicates. c, Immunoblotting for some of the SLCs modified during efferocytosis. Indicated SLCs were probed at various time points after addition of apoptotic cells. Relative intensities of specific bands, normalized to ERK2, are shown below representative blots. d, Immunoblotting for the some of the SLCs in LR73 phagocytes and apoptotic Jurkat cells.
Extended Data Fig. 5 The role of SLC2A1 in efferocytosis.
a, Slc2a1fl/fl BMDMs were treated with or without TAT-Cre to delete Slc2a1. The cells were then incubated with IgG-coated Jurkat cells and engulfment was assessed by CypHer5E signal within BMDMs. The uptake by control BMDMs (not treated with TAT-Cre, and denoted wild type (WT)) was set to 1. b, siRNA targeting of Slc2a1 downregulates SLC2A1 protein expression. Representative western blots from siRNA knockdown of Slc2a1 versus scrambled siRNA in LR73 cells are shown. LR73 cells expressing siRNA-resistant SLC2A1 are also shown. c, Slc2a1 deletion efficiency in Cas9 LR73 cells. Slc2a1 guide was introduced into Cas9–EGFP+ LR73 cell clones. The efficiency of Slc2a1 deletion was quantified using qRT–PCR. d, Introduction of TAT-Cre into Slc2a1fl/fl BMDMs efficiently knocks down SLC2A1 protein expression. Slc2a1fl/fl bone marrow cells were treated with recombinant TAT-Cre during macrophage differentiation after isolation from the bone marrow. e, STF-31 did not affect antibody-mediated phagocytosis by peritoneal macrophages. C57BL/6 mice were intraperitoneally injected with 10 mg kg−1 of either STF-31 in X-VIVO medium 1 h before injection of IgG-coated Jurkat cells. CypHer5E-labelled Jurkat cells were injected intraperitoneally along with the drug. Mice were euthanized 1 h later, peritoneal cells were collected, and apoptotic cell engulfment by CD11b+F4/80high macrophages was analysed by fluorescence-activated cell sorting. f, Slc2a1-deficient LR73 cells or BMDMs were treated with STF-31, and the engulfment assay was conducted using CypHer5E-labelled apoptotic Jurkat cells. CypHer5E+ phagocytic cells after 2 h of incubation were identified by flow cytometry. n.s., not significant. Data are representative of at least two independent experiments with 3–4 replicates per condition. g, The SLC2A1 inhibitor STF-31 does not increase the number of thymocytes stained with 7-aminoactinomycin D (7AAD+) in vitro. Isolated thymocytes were incubated with dexamethasone (10 μM) with or without STF-31 (2 mM). Four hours later, the cell death of the thymocytes was indicated by annexin7+7AAD+. Data are representative of two independent experiments.
Extended Data Fig. 6 The role of glycolytic genes in efferocytosis.
a, The effect of physiological (1 mg ml−1) or high (5 mg ml−1) glucose on apoptotic cell engulfment (2 h) in control and Slc2a1-siRNA-treated LR73 cells. Note that the enhanced engulfment due to higher glucose concentration is lost in siRNA-treated conditions. Data are representative of at least three independent experiments with 3–4 replicates per condition. b, Apoptotic cell engulfment by LR73 cells in the presence of the glucose analogue 2-DG (10 mM). Data are representative of two independent experiments with 2–3 replicates per condition. c, BMDMs undergo glycolytic flux during apoptotic cell clearance. Glycolytic flux and OXPHOS were measured during engulfment assays using Seahorse XF to assess ECAR (left) and OCR (right), respectively. Data are mean ± s.d. for ECAR and OCR over the course of standard glycolytic flux and cellular respiration tests. Data are representative of four replicates per condition. d, Genes within the glycolytic pathway that are significantly upregulated during apoptotic cell clearance. A schematic of the glycolytic pathway and subsequent steps is shown, with enzymes that are significantly upregulated (determined via RNA-seq) indicated in red.
Extended Data Fig. 7 Testing SGK1 and glycolysis in efferocytosis.
a, Differential metabolic requirements of macrophages for efferocytosis versus antibody-mediated phagocytosis. BMDMs were co-cultured with apoptotic or antibody-coated Jurkat cells. Mitochondrial respiration was inhibited by addition of the mitochondrial complex I inhibitor rotenone (200 nM), the mitochondrial complex III inhibitor antimycin A1 (1 μM), or both (R + A). Aerobic glycolysis was inhibited by the addition of the pan-PDK inhibitor dichloroacetate (1 mM). Data are representative of three independent experiments. b, SGK1 inhibition blocks efferocytosis in vitro. LR73 cells were treated with SGK1 inhibitor and uptake of CypHer5E-labelled apoptotic Jurkat cells was assessed. c, BMDMs from GLUT1–Myc knock-in mice were co-cultured with apoptotic thymocytes with or without the SGK1 inhibitor GSK650394 (5 μM) for 2 h, unbound apoptotic cells were washed away and the cell-surface expression of SLC2A1 was measured by flow cytometry after staining for surface Myc tag. Data are representative of at least two independent experiments. d, Continued uptake of apoptotic thymocytes was determined by the MFI (indicative of corpse-derived signal per phagocyte) of LR73 phagocytes over a time course of engulfment. SLC2A1 or SGK1 inhibitors were added at the beginning of engulfment (left of each pair of graphs) or 1 h post-apoptotic cell addition (right of each pair of graphs). Data are representative of at least three independent experiments with 3–4 replicates per condition.
Extended Data Fig. 8 Testing SLC16A1 in efferocytosis.
a, qRT–PCR determination of Sgk1 expression in phagocytes treated with ATP. LR73 cells were treated with indicated amounts of ATP for 4 h. Expression of Sgk1 was determined by qRT–PCR using hamster-specific primers. Data are representative of at least two independent experiments with 3–4 replicates per condition. b, qRT–PCR determination of Slc2a1, Slc16a1 and Sgk1 expression in phagocytes after addition of the PtdSer-masking peptide (GST–TSR) during efferocytosis. Apoptotic cells (AC) were added with or without TSR peptide (10 ng μl−1) for 4 h. Expression of indicated genes was determined by qRT–PCR using hamster-specific primers. Data are representative of at least two independent experiments with 3–4 replicates per condition. c, SLC16A1 inhibition blocks efferocytosis in vitro. LR73 cells were treated with Slc16a1 siRNA and uptake of CypHer5E-labelled apoptotic Jurkat cells was assessed. d, SLC16A1 inhibitor SR13800 dampens efferocytosis by peritoneal macrophages. C57BL/6 mice were injected intraperitoneally with SR13800 (10 mg kg−1) in X-VIVO medium 1 h before injection of apoptotic cells. CypHer5E-labelled apoptotic Jurkat cells were injected intraperitoneally. After 1 h, apoptotic cell engulfment by CD11b+F4/80high peritoneal macrophages was analysed by flow cytometry. Data are representative of two independent experiments with at least six mice in each group per experiment. e, f, Supernatants were prepared from LR73 cells, treated with control or Slc16a1 siRNA, that were engulfing apoptotic cells. The supernatants were added to BMDMs and incubated for 12 h. e, Expression of inflammatory markers was determined by qRT–PCR. f, After 24 h of incubation, expression of CD206 and F4/80 was determined by flow cytometry. Data are representative of two independent experiments with 2–3 replicates per condition.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables 1-4. Supplementary Table 1: Gene List Associated with Fig. 1a. Supplementary Table 2: Categorization and disease associations of SLCs modified during efferocytosis. Supplementary Table 3: List of Hamster-Specific and Mouse-Specific Taqman Primers. Supplementary Table 4: List of Statistical Tests and Sample Sizes Used From Each Figure.
Rights and permissions
About this article
Cite this article
Morioka, S., Perry, J.S.A., Raymond, M.H. et al. Efferocytosis induces a novel SLC program to promote glucose uptake and lactate release. Nature 563, 714–718 (2018). https://doi.org/10.1038/s41586-018-0735-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0735-5
Keywords
This article is cited by
-
TREM2 macrophage promotes cardiac repair in myocardial infarction by reprogramming metabolism via SLC25A53
Cell Death & Differentiation (2024)
-
Dysregulated cellular metabolism in atherosclerosis: mediators and therapeutic opportunities
Nature Metabolism (2024)
-
Differential impact of high-salt levels in vitro and in vivo on macrophage core functions
Molecular Biology Reports (2024)
-
Mechanism of Efferocytosis in Determining Ischaemic Stroke Resolution—Diving into Microglia/Macrophage Functions and Therapeutic Modality
Molecular Neurobiology (2024)
-
Ghost messages: cell death signals spread
Cell Communication and Signaling (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.