Abstract
Genetic regulators and environmental stimuli modulate T cell activation in autoimmunity and cancer. The enzyme co-factor tetrahydrobiopterin (BH4) is involved in the production of monoamine neurotransmitters, the generation of nitric oxide, and pain1,2. Here we uncover a link between these processes, identifying a fundamental role for BH4 in T cell biology. We find that genetic inactivation of GTP cyclohydrolase 1 (GCH1, the rate-limiting enzyme in the synthesis of BH4) and inhibition of sepiapterin reductase (the terminal enzyme in the synthetic pathway for BH4) severely impair the proliferation of mature mouse and human T cells. BH4 production in activated T cells is linked to alterations in iron metabolism and mitochondrial bioenergetics. In vivo blockade of BH4 synthesis abrogates T-cell-mediated autoimmunity and allergic inflammation, and enhancing BH4 levels through GCH1 overexpression augments responses by CD4- and CD8-expressing T cells, increasing their antitumour activity in vivo. Administration of BH4 to mice markedly reduces tumour growth and expands the population of intratumoral effector T cells. Kynurenine—a tryptophan metabolite that blocks antitumour immunity—inhibits T cell proliferation in a manner that can be rescued by BH4. Finally, we report the development of a potent SPR antagonist for possible clinical use. Our data uncover GCH1, SPR and their downstream metabolite BH4 as critical regulators of T cell biology that can be readily manipulated to either block autoimmunity or enhance anticancer immunity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The microarray dataset is accessible through GEO accession number GSE108101. All other datasets generated and/or analysed during this study are available from the corresponding authors upon reasonable request.
Change history
31 July 2019
An Amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Latremoliere, A. et al. Reduction of neuropathic and inflammatory pain through inhibition of the tetrahydrobiopterin pathway. Neuron 86, 1393–1406 (2015).
Werner, E. R., Blau, N. & Thöny, B. Tetrahydrobiopterin: biochemistry and pathophysiology. Biochem. J. 438, 397–414 (2011).
Chen, W. et al. Role of increased guanosine triphosphate cyclohydrolase-1 expression and tetrahydrobiopterin levels upon T cell activation. J. Biol. Chem. 286, 13846–13851 (2011).
Ziegler, I. et al. Control of tetrahydrobiopterin synthesis in T lymphocytes by synergistic action of interferon-γ and interleukin-2. J. Biol. Chem. 265, 17026–17030 (1990).
Chuaiphichai, S. et al. Cell-autonomous role of endothelial GTP cyclohydrolase 1 and tetrahydrobiopterin in blood pressure regulation. Hypertension 64, 530–540 (2014).
Eberl, G. & Littman, D. R. Thymic origin of intestinal αβ T cells revealed by fate mapping of RORγt+ cells. Science 305, 248–251 (2004).
Hobeika, E. et al. Testing gene function early in the B cell lineage in mb1-cre mice. Proc. Natl Acad. Sci. USA 103, 13789–13794 (2006).
Śledzińska, A. et al. TGF-β signalling is required for CD4+ T cell homeostasis but dispensable for regulatory T cell function. PLoS Biol. 11, e1001674 (2013).
Talbot, S. et al. Silencing nociceptor neurons reduces allergic airway inflammation. Neuron 87, 341–354 (2015).
Haworth, O., Cernadas, M., Yang, R., Serhan, C. N. & Levy, B. D. Resolvin E1 regulates interleukin 23, interferon-γ and lipoxin A4 to promote the resolution of allergic airway inflammation. Nat. Immunol. 9, 873–879 (2008).
Martin, S. F. et al. Toll-like receptor and IL-12 signaling control susceptibility to contact hypersensitivity. J. Exp. Med. 205, 2151–2162 (2008).
Rangachari, M. & Kuchroo, V. K. Using EAE to better understand principles of immune function and autoimmune pathology. J. Autoimmun. 45, 31–39 (2013).
Rangachari, M. et al. Bat3 promotes T cell responses and autoimmunity by repressing Tim-3-mediated cell death and exhaustion. Nat. Med. 18, 1394–1400 (2012).
Nar, H. et al. Active site topology and reaction mechanism of GTP cyclohydrolase I. Proc. Natl Acad. Sci. USA 92, 12120–12125 (1995).
Nar, H. et al. Atomic structure of GTP cyclohydrolase I. Structure 3, 459–466 (1995).
Volani, C. et al. Dietary iron loading negatively affects liver mitochondrial function. Metallomics 9, 1634–1644 (2017).
Archer, M. C., Vonderschmitt, D. J. & Scrimgeour, K. G. Mechanism of oxidation of tetrahydropterins. Can. J. Biochem. 50, 1174–1182 (1972).
Eberlein, G., Bruice, T. C., Lazarus, R. A., Henrie, R. & Benkovic, S. J. The interconversion of the 5,6,7,8-tetrahydro-, 7,8-dihydro-, and radical forms of 6,6,7,7-tetramethyldihydropterin. A model for the biopterin center of aromatic amino acid Mixed function oxidases. J. Am. Chem. Soc. 106, 7916–7924 (1984).
Capeillere-Blandin, C., Mathieu, D. & Mansuy, D. Reduction of ferric haemoproteins by tetrahydropterins: a kinetic study. Biochem. J. 392, 583–587 (2005).
Hondowicz, B. D. et al. Interleukin-2-dependent allergen-specific tissue-resident memory cells drive asthma. Immunity 44, 155–166 (2016).
Ewens, A., Mihich, E. & Ehrke, M. J. Distant metastasis from subcutaneously grown E0771 medullary breast adenocarcinoma. Anticancer Res. 25, 3905–3915 (2005).
Curti, A. et al. Indoleamine 2,3-dioxygenase-expressing leukemic dendritic cells impair a leukemia-specific immune response by inducing potent T regulatory cells. Haematologica 95, 2022–2030 (2010).
Haruki, H., Hovius, R., Pedersen, M. G. & Johnsson, K. Tetrahydrobiopterin biosynthesis as a potential target of the kynurenine pathway metabolite xanthurenic acid. J. Biol. Chem. 291, 652–657 (2016).
Oppenheimer, S. J. Iron and its relation to immunity and infectious disease. J. Nutr. 131, 616S–633S (2001).
Cassat, J. E. & Skaar, E. P. Iron in infection and immunity. Cell Host Microbe 13, 509–519 (2013).
Liu, C.-J., Chen, K.-W. Hu, Y.-W. Hong, Y.-C., Huang, Y.-C., Chiou, T.-J. & Tzeng, C.-H. Chronic iron deficiency anemia and cancer risk. Blood 120, 5172 (2012).
Hung, N. et al. Risk of cancer in patients with iron deficiency anemia: a nationwide population-based study. PLoS ONE 10, e0119647 (2015); correction https://doi.org/10.1371/journal.pone.0125951 (2015).
Hennet, T., Hagen, F. K., Tabak, L. A. & Marth, J. D. T-cell-specific deletion of a polypeptide N-acetylgalactosaminyl-transferase gene by site-directed recombination. Proc. Natl Acad. Sci. USA 92, 12070–12074 (1995).
Sawada, S., Scarborough, J. D., Killeen, N. & Littman, D. R. A lineage-specific transcriptional silencer regulates CD4 gene expression during T lymphocyte development. Cell 77, 917–929 (1994).
Ventura, A. et al. Restoration of p53 function leads to tumour regression in vivo. Nature 445, 661–665 (2007).
Crabtree, M. J. et al. Quantitative regulation of intracellular endothelial nitric-oxide synthase (eNOS) coupling by both tetrahydrobiopterin-eNOS stoichiometry and biopterin redox status: insights from cells with tet-regulated GTP cyclohydrolase I expression. J. Biol. Chem. 284, 1136–1144 (2009).
Banerjee, A. et al. Cellular and site-specific mitochondrial characterization of vital human amniotic membrane. Cell Transplant. 27, 3–11 (2018).
Schmitt, T. M. & Zúñiga-Pflücker, J. C. Induction of T cell development from hematopoietic progenitor cells by delta-like-1 in vitro. Immunity 17, 749–756 (2002).
Becker, C. et al. In vivo imaging of colitis and colon cancer development in mice using high resolution chromoendoscopy. Gut 54, 950–954 (2005).
Collison, L. W. & Vignali, D. A. A. In vitro Treg suppression assays. Methods Mol. Biol. 707, 21–37 (2011).
Boivin, N., Baillargeon, J., Doss, P. M. I. A., Roy, A. P. & Rangachari, M. Interferon-β suppresses murine Th1 cell function in the absence of antigen-presenting cells. PLoS ONE 10, e0124802 (2015).
Lin, K. Y. et al. Treatment of established tumors with a novel vaccine that enhances major histocompatibility class II presentation of tumor antigen. Cancer Res. 56, 21–26 (1996).
Smyth, G. K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, https://doi.org/10.2202/1544-6115.1027 (2004).
Edgar, R., Domrachev, M. & Lash, A. E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Arai, N., Narisawa, K., Hayakawa, H. & Tada, K. Hyperphenylalaninemia due to dihydropteridine reductase deficiency: diagnosis by enzyme assays on dried blood spots. Pediatrics 70, 426–430 (1982).
Theurl, I. et al. On-demand erythrocyte disposal and iron recycling requires transient macrophages in the liver. Nat. Med. 22, 945–951 (2016).
Acknowledgements
We thank all members of our laboratories for helpful discussions and Life Science Editors for editorial support. We thank Shanghai ChemPartners for running the drug metabolism and pharmacokinetic assays associated with QM385. J.M.P. is supported by grants from IMBA, the Austrian Ministry of Sciences and the Austrian Academy of Sciences, and the T. Von Zastrow Foundation as well as a European Research Council (ERC) Advanced Grant and an Era of Hope Innovator award. C.J.W. is supported by a National Institutes of Health (NIH) R35 grant (NS105076). We also acknowledge the Christian Doppler Laboratory for Iron Metabolism and Anemia Research as a funding body for our research (M.S. and G.W.). M.R. is supported by EMD Serono, Canada, and a MS Network Transitional Career Development Award.
Reviewer information
Nature thanks R.S. Johnson, L. O’Neill and N. Restifo for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
S.J.F.C., together with C.J.W. and J.M.P., conceived and designed the study. All experiments were performed by S.J.F.C. with the following exceptions: A.W. and A.J. performed mitochondrial respiration analyses; S. Reissig performed colonoscopy grading; S.T., C.S. and B.L.T. carried out the asthma model; C.S. and B.L.T. carried out the HDM model; Y.P. performed the iron-reduction experiment; M.S.L., G.L. and G.W. carried out assays for human T cell proliferation; M.S. performed the iron measurements; T.K. performed in vitro thymocyte differentiation experiments; M.N. performed microarray analysis; E.M., B.L.T. and D.d.L.S. performed biopterin and sepiapterin measurements; M.R., M.K., D.H., M.T., L.T., D.C., S. Rao, M.P. and M.A. helped with the cancer studies; L.B., N.A., A. Latremoliere and M.C. helped with compound dosing and discussions of BH4 biology; M.T. and S.Z. performed QM385 pharmacokinetic analysis. S.J.F.C., C.J.W. and J.M.P. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Upregulation of Gch1 and BH4 in activated T cells.
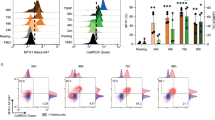
a, Percentage of CD62Llo GFP+ cells from purified Gch1-Gfp CD4+ T cells stimulated for 24 h with phorbol myristate acetate and ionomycin (50 ng ml−1 each). Data are shown as means ± s.e.m., from n = 3 samples. The experiment was repeated two independent times. ***P < 0.001 (two-tailed Student’s t-test). b, c, Representative Gch1-Gfp expression in 16-h-activated (CD62Llow) CD4+ T cells after anti-CD3/CD28 stimulation (b) and representative dose–response of anti-(α)CD3/CD28 stimulation of purified CD4+ Gch1-Gfp T cells for 24 h (c). The experiment was repeated two independent times with similar results. d, Cell numbers of various immune populations in the thymus (left) and spleen (right) from control (n = 3) and Gch1;Lck (n = 3) 8-week-old mice. Data from individual mice are shown as means ± s.e.m. NS, not significant (two-tailed Student’s t-test). e, f, CD4+ (e) and CD8+ (f) T cell proliferation after three days of anti-CD3/28 stimulation, from control and Gch1;Lck mice. g, Representative histogram depicting the proliferation of DN3a thymocytes from control and Gch1;Lck mice cultured on OP9-Dl1 stromal cells for five days. The experiment was repeated two independent times with similar results. h, i, Representative FACS blot depicting the differentiation into CD4+ and CD8+ T cells of DN3a thymocytes from control and Gch1;Lck mice cultured on OP9-Dl1 stromal cells for five days (h), and quantification of the differentiated cell types from n = 3 animals (i). Data from individual mice are shown as means ± s.e.m. NS, not significant (two-tailed Student’s t-test).
Extended Data Fig. 2 Normal T cell development and B cell biology in the absence of Gch1.
a, Thymocyte cell death induced over 24 h by various stimuli: anti-CD3 (0.5 μg ml−1 and 5 μg ml−1), Fas ligand (0.2 μg ml−1 and 2 μg ml−1), dexamethasone (Dex, 0.1 μg ml−1 and 0.5 μg ml−1) and γ-irradiation (1 Gray (Gy)). Data are shown as means ± s.e.m. n = 3 for each genotype. NS, not significant (two-tailed Student’s t-test). b, Death by neglect of purified CD4+ T cells cultured without stimulation for up to 56 h. Data are shown as means ± s.e.m. n = 3 for each genotype. NS, not significant (two-tailed Student’s t-test). c, d, Proliferation of CD4+ T cells from control and Gch1;RORc mice after three days of anti-CD3/28 stimulation. Panels show representative FACS proliferation traces (c) and representative dose response (d). Experiments were repeated independently more than six times with similar results. e, Representative FACS plots from spleens of control and Gch1;MB1 mice. MB1-Cre is an early B cell deleter Cre line using endogenous CD79a B cell specific expression. The experiment was repeated two independent times with similar results. f, g, Representative FACS histogram depicting the proliferation of wild-type B cells treated with vehicle (DMSO) or SPRi3 (50 μM) (f), and of B cells from control and Gch1;MB1 mice in response to LPS (1 μg ml−1) after three days (f). Shaded grey peaks represent unstimulated cells. FACS plots are representative of two independent experiments showing similar results. n = 3 mice per group. h, Class-switch recombination. FACS analysis of splenic CD43− B cells from control and Gch1;MB1 mice stimulated with LPS (20 μg ml−1) for five days to induce class-switch recombination to IgG3. FACS plots are representative of two independent experiments showing similar results.
Extended Data Fig. 3 Development of regulatory T cells and their function in Gch1-ablated mice.
a, b, Representative FACS plot depicting CD4+FoxP3+ regulatory T cells (T regs; a) and quantification of T-reg proportions as well as absolute numbers in the spleen (b) of control and Gch1;RORc mice (n = 6 each). Data are shown as means ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001; NS, not significant (two-tailed Student’s t-test). c, d, In vitro T-reg suppression assay, in which naive, wild-type CD4+ T cells were activated in the presence of varying ratios of T-reg cells from control and Gch1;RORc mice for four days. Representative histogram showing the suppressive capacity of control and Gch1;RORc T-reg cells (c) and quantification of proliferation with various ratios of T-reg cells (d). n = 4 samples. Data are shown as means ± s.e.m. *P < 0.05; **P < 0.01; NS, not significant (two-tailed Student’s t-test with multiple comparisons). Tconv, conventional CD4+ T cells (CD4+, CD25− CD45RBhigh). e, Naive CD4+ transfer colitis model, with co-transfer of FACS-purified T-reg cells from control (n = 4) and Gch1;RORc (n = 4) mice. As a control, Tconv cells (from n = 16 mice) with no co-transfer of T-reg cells were used. Changes to initial body weight (BW) were scored over five weeks. Data are shown as means ± s.e.m. *P < 0.05; ***P < 0.001; NS, not significant (two-way ANOVA with Tukey’s multiple comparison test). f, Total numbers of CD4+ splenic T cells at two weeks post-transfer in mice (n = 3) transferred with naive CD4+ cells only (‘no T regs’) and mice transferred with T regs from control or Gch1-ablated (Gch1;RORc) mice. Data are shown as means ± s.e.m. ***P < 0.001; ****P < 0.0001; NS, not significant (one-way ANOVA with Dunnett’s multiple comparison test). g, Transfer colitis model of intestinal autoimmunity. Body-weight changes are plotted relative to initial weight in mice transferred with naive CD4+ T cells from control or Gch1;Lck mice (n = 10 each). Data are shown as means ± s.e.m. NS, not significant (two-way ANOVA with Sidak’s multiple comparisons). h, i, Proportion of CD4+ T cells in the draining mesenteric lymph nodes in week 4 (h), and profiles of ntracellular cytokines (IFN-γ and IL-17) from transferred control and Gch1;Lck cells (i). Data are shown as means ± s.e.m. n = 10 for each genotype for h and n = 5 for each genotype for i. ***P < 0.001; NS, not significant (two-tailed Student’s t-test).
Extended Data Fig. 4 Blockage of GCH1–BH4 abrogates T-cell-mediated autoimmunity.
a, OVA immunization of control and Gch1;Lck mice. T-cell-dependent IgG responses and T-cell-independent IgM responses are shown two weeks after OVA immunization (left panels, 100 μg OVA in 200 μg alum) as well as two weeks after re-challenge (right panels). n = 5 for control mice; n = 6 for Gch1;Lck mice. Data are shown as means ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001; NS, not significant (two-tailed Student’s t-test with multiple comparisons). b, c, EAE model of autoimmunity towards the central nervous system. Data are shown as means ± s.e.m. b, EAE scores of control and Gch1;Lck mice. n = 6 for each genotype. ****P < 0.0001 (linear regression analysis was performed on the slope of each curve). c, Mean maximal EAE severity in control and littermate Gch1;Lck mice. *P < 0.05 (Mann–Whitney test). d, Schematic of the de novo, salvage and recycling arms of the BH4 pathway. The dotted arrow indicates non-enzymatic reactions; solid arrows indicate enzymatic reactions. DHFR, dihydrofolate reductase; GTP, guanosine triphosphate; PCDB, pterin-4α-carbinolamine dehydratase; PTPS, 6-pyruvoyl tetrahydropterin synthase; QDPR, quinoid dihydropteridine reductase; SPR, sepiapterin reductase. e, Representative FACS plots depicting activation marker profiles of purified wild-type control CD4+ T cells left unstimulated or stimulated with anti-CD3/28 antibodies for 16 h and then treated with vehicle (DMSO), SPRi3 (50 μM) or sepiapterin (5 μM). The experiment was repeated two independent times with similar results. f, Cell survival as defined by the percentage of DAPI−annexinV− cells from purified CD4+ T cells stimulated for 24 h or 48 h with anti-CD3/28 antibodies and then treated with vehicle (DMSO), SPRi3 (50 μM) or sepiapterin (5 μM). The experiment was repeated two independent times with similar results. g, h, Representative FACS blots depicting EdU cell-cycle analysis after 28 hours anti-CD3/CD28 stimulation of control, Gch1;RORc and SPRi3-treated control CD4+ T cells. EdU was pulsed for the last 4 hours (g) and quantification of S-phase entry (h). Data from individual mice are shown ± s.e.m. ***P < 0.001 (one-way ANOVA with Dunnett’s multiple comparisons test). i, Quantification of subG1 (dead cells) populations after 24- and 48-h stimulation. EdU was pulsed for the last 4 hours of each time point. Data from individual mice are shown ± s.e.m.). **P < 0.01; NS, not significant (multiple t-test comparisons). j, k, Amino acid profiles in the supernatants (j) and cell pellets (k) from 24-h anti-CD3/CD28-stimulated CD4+ T cells from control and Gch1;Lck mice. n = 3 for each genotype. Data are shown as means ± s.e.m. NS, not significant (two-tailed Student’s t-test).
Extended Data Fig. 5 Mitochondrial dysfunction in BH4-depleted T cells after activation.
a, b, ATP measurements in control (n = 3) and Gch1;Lck (n = 3) CD4+ T cells (a) and in wild-type CD4+ T cells treated with DMSO vehicle (n = 3) or SPRi3 (50 μM; n = 3) (b), either left unstimulated or assayed at the indicated time points after T cell activation with anti-CD3/28 antibodies. Data are shown as means ± s.e.m. n = 3 for each genotype. *P < 0.05; **P < 0.01 (two-tailed Student’s t-test with multiple comparisons). c, Metabolomic measurements of lactate and pyruvate levels in cell pellets of 16-h anti-CD3/28-activated CD4+ T cells from control and Gch1;Lck mice. Data are shown as means ± s.e.m. n = 4 for each genotype. *P < 0.05 (two-tailed Student’s t-test). d, Routine and total capacitance oxygen respiration in intact, 16-h anti-CD3/CD28-stimulated CD4+ T cells from control and Gch1;Lck mice. Data from individual mice are indicated ± s.e.m. n = 4 for each genotype. **P < 0.01 (two-tailed Student’s t-test). e, f, Oxygen uptake rate in permeabilized, 16-h anti-CD3/CD28-stimulated CD4+ T cells from control (n = 4) and Gch1;RORc (n = 4) mice (e) and wild-type CD4+ T cells treated with DMSO or SPRi3 (50 μM (n = 5 each) (f). Data from individual mice are indicated ± s.e.m. *P < 0.05; **P < 0.01 (two-tailed Student’s t-test). g, Left, representative oxygen consumption traces of complex-I-linked and complex-II-linked ETC activity from 16-h-activated wild-type CD4+ T cells treated with vehicle or SPRi3 (50 μM). Right, relative complex-I- and complex-II-linked activities in activated control cells treated with vehicle (n = 4) or SPRi3 (50 μM; n = 4). Data are shown as means ± s.e.m. NS, not significant; *P < 0.05 (two-tailed Student’s t-test).
Extended Data Fig. 6 Enhanced superoxide levels independent of iNOS coupling observed in BH4-deficient activated T cells.
a, b, Representative FACS histogram (a) and quantification of the mean fluorescent intensity (MFI; b) showing levels of DHE (dihydroethidium, a superoxide ROS indicator) in unstimulated and 20-h anti-CD3/28-activated CD4+ T cells from control and GCH1;RORc mice as well as control cells treated with SPRi3 (50 μM). n = 3 samples per group. The experiment was repeated three independent times with similar results. c, d, Proliferation of control (n = 6) and Gch1;Lck (n = 9) CD4+ T cells and treatment with the superoxide scavenger NAC (500 μM; n = 4 each). Representative three-day proliferation histograms are shown in c; quantification is shown in d. Data are given as means ± s.e.m. Individual mice for each genotype are shown. ****P < 0.0001 (one-way ANOVA with Tukey’s multiple comparison test). e, Total iron content from unstimulated or 24-h anti-CD3/28-stimulated CD4+ T cells (untreated or treated with 500 μM NAC) from control (n = 17, 4, respectively) and Gch1;RORc (n = 22, 6, respectively) mice. Data are shown as means ± s.e.m. Individual mice for each genotype are shown. **P < 0.01 (two-tailed Student’s t-test with Tukey’s multiple comparisons). f, ATP measurements from stimulated wild-type CD4+ T cells treated with DMSO, sepiapterin and NAC for 24 h. Data are shown as means ± s.e.m. n = 5 for each genotype. *P < 0.05; **P < 0.01 (two-tailed Student’s t-test with multiple comparisons). g, Intracellular iNOS expression in purified CD4+ control T cells left untreated or anti-CD3/CD28-stimulated for 12 h, 24 h or 72 h. The experiment was repeated two independent times with similar results. h, i, Representative histogram showing iNOS expression in control and Gch1-ablated CD4+ T cells stimulated with anti-CD3/CD28 antibodies for 72 h (h) and the percentage of iNOS+ cells was quantified over time (i). n = 4 for each genotype. Data are shown as means ± s.e.m. NS, not significant (two-tailed Student’s t-test). j, Nitrite measurements in the supernatant of stimulated cells from i. Peritoneal, thioglycollate-elicited macrophages stimulated with LPS (100 ng ml−1) for 24 h were used as a positive control. Data are shown as means ± s.e.m. n = 4 for each genotype. NS, not significant (two-tailed Student’s t-test).
Extended Data Fig. 7 Functional evaluation of the SPR blocker QM385.
a, The BH4 pathway, indicating how QM385 acts on SPR, limiting BH4 production and correspondingly increasing sepiapterin levels, which can be used as a biomarker for QM385-mediated SPR inhibition. b, c, A representative concentration–response curve showing the binding affinity of QM385 to human SPR, tested in vitro by TR-FRET (b); and reduction of BH4 levels upon QM385 treatment in anti-CD3/28-stimulated mouse splenocytes (left panel, two independent experiments) and human PBMCs (right panel, two independent experiments) (c). The calculated half maximal inhibitory concentration (IC50) values for each assay are indicated in red. The binding-effect assay was repeated 162 independent times with similar results. d, The oxygen-uptake rate in permeabilized, 16-h anti-CD3/CD28-stimulated wild-type CD4+ T cells treated with DMSO or QM385 (2.5 μM). Data from individual mice (n = 3) are indicated ± s.e.m. ***P < 0.001 (two-tailed Student’s t-test). e, ATP measurements of unstimulated (n = 8) and 24-h-activated wild-type CD4+ T cells treated with DMSO vehicle (n = 4) or varying doses of QM385 (n = 4 for each dose). Data are shown as means ± s.e.m. NS, not significant; **P < 0.01 (one-way ANOVA with Dunnett’s multiple comparisons). f, Fold changes in DHE levels between CD4+ T cells treated with DMSO or QM385 (2.5 μM) and activated for 20 h. Data from individual mice (n = 4) are indicated ± s.e.m. **P < 0.01 (two-tailed Student’s t-test). g, Allergic airway inflammatory disease model and quantification of inflammatory cells in bronchoalveolar lavage fluids (BALFs). Data are shown as box-and-whisker plots (running from minimal to maximal values); individual data points are shown. n = 15 for vehicle-treated mice; n = 17 for QM385-treated mice. QM385 (1 mg kg−1) was administered orally (peritoneally) twice a day for three consecutive days as depicted in the diagram. *P < 0.05; **P < 0.01 (two-tailed Student’s t-test). h, Proliferation of human CD4+ T cells from two donors performed in triplicate samples. Anti-CD3/28 T cells were stimulated with varying doses of QM385 and total counts were measured. Data are shown as means ± s.e.m. **P < 0.01; P < 0.05 (one-way ANOVA with Dunnett’s multiple comparisons).
Extended Data Fig. 8 Increased numbers of effector T cells in naive mice overexpressing Gch1, and enhanced T cell proliferation after stimulation.
a, Representative immunoblot to detect GCH1 and the HA tag in naive CD4+ T cells from control and GOE;Lck-overexpressing mice. The experiment was repeated three times with similar findings. b, c, The proportion of splenic T and B cells (b) and the proportion of CD4+ and CD8+ T cells among the splenic T cell (TCRβ+) population (c), from control (n = 4) and GOE;Lck (n = 4) mice. Data for individual mice aged eight weeks are shown as means ± s.e.m. NS, not significant (two-tailed Student’s t-test). d, Quantification of naive (CD44lowCD62Lhigh), memory (CD44highCD62Lhigh) and effector (CD44highCD62Llow) T cell subtypes from the spleen of control (n = 6) and GOE;Lck (n = 10) mice. Data for individual mice are shown as means ± s.e.m. **P < 0.01; ***P < 0.001; NS, not significant (two-tailed Student’s t-test). e, Cell numbers for B cells, T cells, and CD4+ and CD8+ T cells in the spleens of control and GOE;Cd4 mice. Data from individual mice (n = 4 for each genotype) are shown as means ± s.e.m. NS, not significant (two-tailed Student’s t-test). f, Proportion of CD4+ and CD8+ naive, memory and effector T cells in the spleens of naive control and GOE;Cd4 mice. Data for individual mice (n = 4 for each genotype) are shown as means ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001; NS, not significant (two-tailed Student’s t-test). g, Representative histograms depicting dose-dependent proliferation of control and GOE;Cd4 CD4+ T cells stimulated for three days with anti-CD3/28 antibodies. Experiments were repeated more than three times with comparable results. h, IL-2 and IFN-γ secretion after three days of stimulation (with anti-CD3/28 antibodies) of control and GOE;Cd4 CD4+ T cells. Data are shown as means ± s.e.m. n = 3 for each genotype. *P < 0.05; ***P < 0.0001 (two-tailed Student’s t-test). i, j, Cells from control (n = 3) and GOE;ERT (n = 3) mice were stimulated with anti-CD3/28 antibodies for three days and treated with 4-hydroxytamoxifen (4-OHT; 0.5 μM) to induce Gch1 overexpression in vitro. i, Quantification of proliferation of CD4+ T cells; j, cytokine secretion. Data from individual mice are shown as means ± s.e.m. **P < 0.01; ***P < 0.001 (two-tailed Student’s t-test).
Extended Data Fig. 9 T cells overproducing BH4 display enhanced ATP production, proliferation and autoimmunity.
a, Allergic airway inflammatory disease model and fold change of inflammatory cells in BALFs, comparing control and GOE;Lck mice. Data are shown as means ± s.e.m. n = 18 for control mice; n = 17 for GOE;Lck mice. *P < 0.05; **P < 0.01 (two-tailed Student’s t-test). b, Transfer colitis model. Changes in body weight of Rag2−/− mice transferred with control (n = 6 animals) or GOE;Cd4 (n = 5) naive CD4+ T cells. Data are shown as means ± s.e.m. *P < 0.05; ***P < 0.001; NS, not significant (two-way ANOVA with Tukey’s multiple comparison test). c, Total numbers of activated (CD62LlowCD44high) CD4+ splenic T cells at three weeks post-transfer in mice transferred with control or GOE;Cd4 naive CD4+ T cells. Data for two mice from each group are shown. d, Transfer colitis model, involving transfer of naive CD4+ T cells (150,000 cells) and co-transfer of FACS-purified T-reg cells from control (n = 5) and GOE;Cd4 (n = 6) mice. Changes to initial body weights were scored over seven weeks. Data are shown as means ± s.e.m. ***P < 0.001; NS, not significant (two-way ANOVA with Tukey’s multiple comparison test). e, Representative histograms depicting the proliferation of purified unstimulated and anti-CD3/28-stimulated CD4+ and CD8+ wild-type and Gch1;RORc T cells treated for three days with sepiapterin (5 μM). The profile for the unstimulated T cells of each genotype is shown in grey. Experiments were repeated three independent times with comparable results. f, g, Representative FACS plots showing EdU-based cell-cycle analysis following 28-h anti-CD3/28 stimulation of control CD4+ T cells, GOE;Cd4 CD4+ T cells, control CD4+ T cells treated with sepiapterin (5 μM), and GCH1;RORc CD4+ T cells treated with sepiapterin (5 μM) (f); and quantification of the S-phase-entry population (g). EdU was pulsed for the last 4 h of stimulation. n = 4 mice for control; n = 3 mice for all other genotypes. ***P < 0.001 (one-way ANOVA with Dunnett’s multiple comparisons test). h, i, Effect of BH4 on the proliferation (3H-thymidine incorporation; h) and IL-2 secretion (i) of CD4+ wild-type T cells activated with anti-CD3/28 antibodies for 24 h and treated with vehicle (n = 3/4) or BH4 (10 μM; n = 3/4). Data are shown for individual mice as means ± s.e.m. **P < 0.01 (two-tailed Student’s t-test). j, Representative histograms depicting the proliferation of control and Gch1;RORc CD4+ T cells after three days of anti-CD3/28 stimulation supplemented with BH4 (10 μM). FACS blots are representative of two independent experiments with comparable results.
Extended Data Fig. 10 Overactivation of the GCH1–BH4 pathway leads to enhanced anti-tumour immunity.
a, Total iron content from 24-h anti-CD3/28 stimulated CD4+ T cells (untreated or treated with 5 μM sepiapterin) from control (n = 5/4) and Gch1;RORc (n = 5) mice. Data are shown as means ± s.e.m.; individual mice for each genotype are shown. *P < 0.05 (one-way ANOVA with Tukey’s multiple comparisons. b, Representative FACS histogram depicting DHE levels (left) and quantification of the mean fluorescent intensity (right) in unstimulated and 20-h anti-CD3/28-activated CD4+ T cells from control and GOE;Cd4 littermates as well as wild-type cells treated with sepiapterin (5 μM). n = 3 for each condition. Data are shown as means ± s.e.m. **P < 0.01; ***P < 0.001 (one-way ANOVA with Tukey’s multiple comparisons test). c, ATP measurements for stimulated wild-type CD4+ T cells treated with DMSO, sepiapterin (5 μM) or SPRi3 (50 μM) for 24 h. Data are shown as means ± s.e.m. n = 3 for each genotype. *P < 0.05; **P < 0.01 (two-tailed Student’s t-test with multiple comparisons). d, Quantification of intratumoral effector CD4+ T cells (CD44+CD62Llow) assayed from E0071 tumours on day 28 for vehicle- and BH4-treated mice. Data are shown as means ± s.e.m. n = 5 mice for each condition. *P < 0.05; **P < 0.01 (two-tailed Student’s t-test). e, Effect of BH4 supplementation on H-Ras-transformed TC-1 tumour growth. TC-1 tumour cells were orthotopically injected; once the tumours were palpable (day 7), BH4 (35 mg kg−1; n = 15) or vehicle (saline; n = 10) was therapeutically administered for seven days as indicated. Data are shown for individual mice as means ± s.e.m. ***P < 0.001; ****P < 0.0001 (two-way ANOVA with Sidak’s multiple comparisons). f, Quantification of intratumoral effector CD8+ T cells (CD44+CD62Llow) assayed from TC-1 tumours on day 21 in vehicle- or BH4-treated mice (n = 9 mice for each genotype). Data are shown as means ± s.e.m. **P < 0.01 (two-tailed Student’s t-test). g, Effect of BH4 supplementation on TC-1 tumour growth in Rag2−/− hosts. TC-1 tumour cells were orthotopically injected into Rag2−/− female mice; once the tumours were palpable (day 7), BH4 (35 mg kg−1; n = 5) or vehicle (saline; n = 5) was administered. BH4 and vehicle supplementation was carried out for seven days as indicated on the graph. Data are shown for individual mice as means ± s.e.m. NS, not significant (two-way ANOVA with Sidak’s multiple comparisons). h, Sepiapterin levels in the supernatant of wild-type CD4+ T cells stimulated with anti-CD3/28 antibodies for 20 h and treated with vehicle or kynurenine (KYN; 150 μM). Culture medium was also included for comparison. BQL, below quantifiable levels. Data are shown as means ± s.e.m. n = 4 independent samples for each condition. ***P < 0.001 (one-way ANOVA with Tukey’s multiple comparisons test). i, Representative histogram depicting proliferation of three-day anti-CD3/28-activated wild-type CD4+ T cells treated with vehicle or kynurenine (50 μM). j, Representative FACS histograms depicting DHE levels in anti-CD3/28-stimulated wild-type CD4+ T cells treated with vehicle (DMSO), kynurenine alone (50 μM) or kynurenine (50 μM) plus BH4 (10 μM) for 20 h. The experiment was repeated three independent times with comparable results.
Supplementary information
Supplementary Figure
This file contains Supplementary Figures.
Supplementary Table 1
Amino acid and neurotransmitter profiles in stimulated CD4+ T cells from control and Gch1;Lck mice. Amino acid profiles in the supernatants (upper) and the cell pellets (lower) from 24-hour anti-CD3/28 stimulated CD4+ T cells from control and Gch1;Lck mice. n=3 for each genotype. Data are shown as means ± s.e.m. c, Biogenic amine profiles in the cell pellets (upper panel) and supernatants (lower panel) from 24-hour anti-CD3/28 stimulated CD4+ T cells from control and Gch1;Lck mice. n=3 for each genotype.
Supplementary Table 2
Pharmacokinetic parameters of QM385. Pharmacokinetic (PK) parameters of QM385 after intravenous (IV) or oral (PO) administration in male C57BL/6 mice including maximum observed concentration (Cmax), time to Cmax (Tmax), half-life of elimination, (Terminal t1/2), area under the curve (AUC) parameters, mean residence time (MRTinf), apparent total clearance of the drug from plasma after oral administration (CL) and apparent volume of distribution during terminal phase (Vss). F (%) is the fraction of oral QM385 administration that reaches systemic circulation.
Supplementary Table 3
Evaluation of potential off-target liabilities of QM385. Biochemical assay results are presented as a percentage inhibition of specific binding or activity. Significant responses are typically those demonstrating ≥ 50% inhibition or stimulation for biochemical assays. No significant effects were observed in the above assays.Hum,human.
Supplementary Table 4
Information on the antibodies used in the study. A list of all the antibodies used for FACS or western blot experiments including the clone/catalog number, commercial provider and dilution used.
Supplementary Table 5
Endoscopic colitis grading. Table showing how the parameters were used and scoring method to assess colitis score.
Rights and permissions
About this article
Cite this article
Cronin, S.J.F., Seehus, C., Weidinger, A. et al. The metabolite BH4 controls T cell proliferation in autoimmunity and cancer. Nature 563, 564–568 (2018). https://doi.org/10.1038/s41586-018-0701-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0701-2
Keywords
This article is cited by
-
The roles and molecular mechanisms of non-coding RNA in cancer metabolic reprogramming
Cancer Cell International (2024)
-
The pleiotropic functions of reactive oxygen species in cancer
Nature Cancer (2024)
-
Enhanced T cell immune activity mediated by Drp1 promotes the efficacy of PD-1 inhibitors in treating lung cancer
Cancer Immunology, Immunotherapy (2024)
-
Amino acid metabolism in immune cells: essential regulators of the effector functions, and promising opportunities to enhance cancer immunotherapy
Journal of Hematology & Oncology (2023)
-
The diversified role of mitochondria in ferroptosis in cancer
Cell Death & Disease (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.