Abstract
During early human pregnancy the uterine mucosa transforms into the decidua, into which the fetal placenta implants and where placental trophoblast cells intermingle and communicate with maternal cells. Trophoblast–decidual interactions underlie common diseases of pregnancy, including pre-eclampsia and stillbirth. Here we profile the transcriptomes of about 70,000 single cells from first-trimester placentas with matched maternal blood and decidual cells. The cellular composition of human decidua reveals subsets of perivascular and stromal cells that are located in distinct decidual layers. There are three major subsets of decidual natural killer cells that have distinctive immunomodulatory and chemokine profiles. We develop a repository of ligand–receptor complexes and a statistical tool to predict the cell-type specificity of cell–cell communication via these molecular interactions. Our data identify many regulatory interactions that prevent harmful innate or adaptive immune responses in this environment. Our single-cell atlas of the maternal–fetal interface reveals the cellular organization of the decidua and placenta, and the interactions that are critical for placentation and reproductive success.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Our expression data for different tissues are also available for user-friendly interactive browsing online at http://data.teichlab.org (maternal–fetal interface). The raw sequencing data, expression-count data with cell classifications and the whole-genome sequencing data are deposited at ArrayExpress, with experiment codes E-MTAB-6701 (for droplet-based data), E-MTAB-6678 (for Smart-seq2 data) and E-MTAB-7304 (for the whole-genome sequencing data). Our CellPhoneDB repository is available at www.CellPhoneDB.org.
References
Ramathal, C. Y., Bagchi, I. C., Taylor, R. N. & Bagchi, M. K. Endometrial decidualization: of mice and men. Semin. Reprod. Med. 28, 17–26 (2010).
Koopman, L. A. et al. Human decidual natural killer cells are a unique NK cell subset with immunomodulatory potential. J. Exp. Med. 198, 1201–1212 (2003).
Burton, G. J., Watson, A. L., Hempstock, J., Skepper, J. N. & Jauniaux, E. Uterine glands provide histiotrophic nutrition for the human fetus during the first trimester of pregnancy. J. Clin. Endocrinol. Metab. 87, 2954–2959 (2002).
Hempstock, J., Cindrova-Davies, T., Jauniaux, E. & Burton, G. J. Endometrial glands as a source of nutrients, growth factors and cytokines during the first trimester of human pregnancy: a morphological and immunohistochemical study. Reprod. Biol. Endocrinol. 2, 58 (2004).
Burton, G. J., Woods, A. W., Jauniaux, E. & Kingdom, J. C. P. Rheological and physiological consequences of conversion of the maternal spiral arteries for uteroplacental blood flow during human pregnancy. Placenta 30, 473–482 (2009).
Fisher, S. J. Why is placentation abnormal in preeclampsia? Am. J. Obstet. Gynecol. 213, S115–S122 (2015).
Jauniaux, E. & Burton, G. J. Placenta accreta spectrum: a need for more research on its aetiopathogenesis. BJOG 125, 1449–1450 (2018).
Apps, R., Gardner, L. & Moffett, A. A critical look at HLA-G. Trends Immunol. 29, 313–321 (2008).
Apps, R. et al. Human leucocyte antigen (HLA) expression of primary trophoblast cells and placental cell lines, determined using single antigen beads to characterize allotype specificities of anti-HLA antibodies. Immunology 127, 26–39 (2009).
Sharkey, A. M. et al. Killer Ig-like receptor expression in uterine NK cells is biased toward recognition of HLA-C and alters with gestational age. J. Immunol. 181, 39–46 (2008).
Parham, P. & Moffett, A. Variable NK cell receptors and their MHC class I ligands in immunity, reproduction and human evolution. Nat. Rev. Immunol. 13, 133–144 (2013).
Moffett, A. & Colucci, F. Co-evolution of NK receptors and HLA ligands in humans is driven by reproduction. Immunol. Rev. 267, 283–297 (2015).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protocols 9, 171–181 (2014).
Burton, G. J. & Watson, A. L. The structure of the human placenta: implications for initiating and defending against virus infections. Rev. Med. Virol. 7, 219–228 (1997).
Tsang, J. C. H. et al. Integrative single-cell and cell-free plasma RNA transcriptomics elucidates placental cellular dynamics. Proc. Natl Acad. Sci. USA 114, E7786–E7795 (2017).
Liu, Y. et al. Single-cell RNA-seq reveals the diversity of trophoblast subtypes and patterns of differentiation in the human placenta. Cell Res. 28, 819–832 (2018).
Madigan, J. et al. Chemokine scavenger D6 is expressed by trophoblasts and aids the survival of mouse embryos transferred into allogeneic recipients. J. Immunol. 184, 3202–3212 (2010).
Mariathasan, S. et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548 (2018).
Maltepe, E. & Fisher, S. J. Placenta: the forgotten organ. Annu. Rev. Cell Dev. Biol. 31, 523–552 (2015).
Bolnick, J. M. et al. Altered biomarkers in trophoblast cells obtained noninvasively prior to clinical manifestation of perinatal disease. Sci. Rep. 6, 32382 (2016).
Lucas, E. S. et al. Reconstruction of the decidual pathways in human endometrial cells using single-cell RNA-seq. Preprint at https://www.biorxiv.org/content/early/2018/07/13/368829 (2018).
Muñoz-Fernández, R. et al. Human predecidual stromal cells have distinctive characteristics of pericytes: cell contractility, chemotactic activity, and expression of pericyte markers and angiogenic factors. Placenta 61, 39–47 (2018).
Gamliel, M. et al. Trained memory of human uterine NK cells enhances their function in subsequent pregnancies. Immunity 48, 951–962 (2018).
Kozuki, N. et al. The associations of parity and maternal age with small-for-gestational-age, preterm, and neonatal and infant mortality: a meta-analysis. BMC Public Health 13, S2 (2013).
Cichocki, F. et al. ARID5B regulates metabolic programming in human adaptive NK cells. J. Exp. Med. 215, 2379–2395 (2018).
Hanna, J. et al. Decidual NK cells regulate key developmental processes at the human fetal–maternal interface. Nat. Med. 12, 1065–1074 (2006).
Jokhi, P. P., King, A., Boocock, C. & Loke, Y. W. Secretion of colony stimulating factor-1 by human first trimester placental and decidual cell populations and the effect of this cytokine on trophoblast thymidine uptake in vitro. Hum. Reprod. 10, 2800–2807 (1995).
Hamilton, G. S., Lysiak, J. J., Watson, A. J. & Lala, P. K. Effects of colony stimulating factor-1 on human extravillous trophoblast growth and invasion. J. Endocrinol. 159, 69–77 (1998).
Sato, Y. et al. Trophoblasts acquire a chemokine receptor, CCR1, as they differentiate towards invasive phenotype. Development 130, 5519–5532 (2003).
Böttcher, J. P. et al. NK cells stimulate recruitment of cDC1 into the tumor microenvironment promoting cancer immune control. Cell 172, 1022–1037 (2018).
Sotiropoulou, G. & Pampalakis, G. Kallikrein-related peptidases: bridges between immune functions and extracellular matrix degradation. Biol. Chem. 391, 321–331 (2010).
Takenaka, M. C., Robson, S. & Quintana, F. J. Regulation of the T cell response by CD39. Trends Immunol. 37, 427–439 (2016).
Vijayan, D., Young, A., Teng, M. W. L. & Smyth, M. J. Targeting immunosuppressive adenosine in cancer. Nat. Rev. Cancer 17, 709–724 (2017).
Smith, G. C. S. First-trimester determination of complications of late pregnancy. J. Am. Med. Assoc. 303, 561–562 (2010).
Pavličev, M. et al. Single-cell transcriptomics of the human placenta: inferring the cell communication network of the maternal–fetal interface. Genome Res. 27, 349–361 (2017).
Camp, J. G. et al. Multilineage communication regulates human liver bud development from pluripotency. Nature 546, 533–538 (2017).
Puram, S. V. et al. Single-cell transcriptomic analysis of primary and metastatic tumor ecosystems in head and neck cancer. Cell 171, 1611–1624 (2017).
Skelly, D. A. et al. Single-cell transcriptional profiling reveals cellular diversity and intercommunication in the mouse heart. Cell Reports 22, 600–610 (2018).
Pardoll, D. M. The blockade of immune checkpoints in cancer immunotherapy. Nat. Rev. Cancer 12, 252–264 (2012).
Bulmer, J. N., Morrison, L., Longfellow, M., Ritson, A. & Pace, D. Granulated lymphocytes in human endometrium: histochemical and immunohistochemical studies. Hum. Reprod. 6, 791–798 (1991).
King, A., Wellings, V., Gardner, L. & Loke, Y. W. Immunocytochemical characterization of the unusual large granular lymphocytes in human endometrium throughout the menstrual cycle. Hum. Immunol. 24, 195–205 (1989).
Gerrelli, D., Lisgo, S., Copp, A. J. & Lindsay, S. Enabling research with human embryonic and fetal tissue resources. Development 142, 3073–3076 (2015).
O’Rahilla, R. & Muller, F. Human Embryology and Teratology (Wiley-Liss, New York, 1992).
Hern, W. M. Correlation of fetal age and measurements between 10 and 26 weeks of gestation. Obstet. Gynecol. 63, 26–32 (1984).
Bulmer, J. N., Williams, P. J. & Lash, G. E. Immune cells in the placental bed. Int. J. Dev. Biol. 54, 281–294 (2010).
Trombetta, J. J. et al. Preparation of single-cell RNA-seq libraries for next generation sequencing. Curr. Protoc. Mol. Biol. 107, 4.22.1–4.22.17 (2014).
Villani, A.-C. et al. Single-cell RNA-seq reveals new types of human blood dendritic cells, monocytes, and progenitors. Science 356, eaah4573 (2017).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Butler, A. & Satija, R. Integrated analysis of single cell transcriptomic data across conditions, technologies, and species. Preprint at https://www.biorxiv.org/content/early/2017/07/18/164889 (2017).
van den Brink, S. C. et al. Single-cell sequencing reveals dissociation-induced gene expression in tissue subpopulations. Nat. Methods 14, 935–936 (2017).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Hiby, S. E. et al. Combinations of maternal KIR and fetal HLA-C genes influence the risk of preeclampsia and reproductive success. J. Exp. Med. 200, 957–965 (2004).
Robinson, J. et al. The IPD and IMGT/HLA database: allele variant databases. Nucleic Acids Res. 43, D423–D431 (2015).
Kang, H. M. et al. Multiplexed droplet single-cell RNA-sequencing using natural genetic variation. Nat. Biotechnol. 36, 89–94 (2018).
Stubbington, M. J. T. et al. T cell fate and clonality inference from single-cell transcriptomes. Nat. Methods 13, 329–332 (2016).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Mills, R. E. et al. Natural genetic variation caused by small insertions and deletions in the human genome. Genome Res. 21, 830–839 (2011).
1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Van der Auwera, G. A. et al. From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr. Protoc. Bioinformatics 43, 11.10.1–11.10.33 (2013).
Broad Institute. Picard tools https://broadinstitute.github.io/picard/ (Broad Institute, 2018).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
Robinson, J., Mistry, K., McWilliam, H., Lopez, R. & Marsh, S. G. E. IPD—the Immuno Polymorphism Database. Nucleic Acids Res. 38, D863–D869 (2010).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Turro, E. et al. Haplotype and isoform specific expression estimation using multi-mapping RNA-seq reads. Genome Biol. 12, R13 (2011).
Bachelerie, F. et al. International Union of Basic and Clinical Pharmacology. LXXXIX. Update on the extended family of chemokine receptors and introducing a new nomenclature for atypical chemokine receptors. Pharmacol. Rev. 66, 1–79 (2013).
Orchard, S. et al. Protein interaction data curation: the International Molecular Exchange (IMEx) consortium. Nat. Methods 9, 345–350 (2012).
Acknowledgements
We thank G. Graham, J. Shilts, A. Lopez, N. Reuter, S. Orchard and P. Porras for discussions on CellPhoneDB; D. Dixon, D. Popescu, J. Fletcher, O. Chazara, L. Mamanova, A. Jinat, C. I. Mazzeo, D. McDonald and D. Bulmer for experimental help; A. Hupalowska for help with the illustrations; S. Lindsay, A. Farnworth, the HDBR, P. Ayuk and the Newcastle Uteroplacental Tissue Bank for providing samples; R. Rostom, D. McCarthy, V. Svensson, M. Hemberg and T. Gomes for computational discussions. We are indebted to the donors for participating in this research. This project was supported by ERC grants (ThDEFINE, ThSWITCH) and an EU FET-OPEN grant (MRG-GRAMMAR no. 664918) and Wellcome Sanger core funding (no. WT206194). R.V.-T. is supported by an EMBO and HFSP Long-Term Fellowship and J.-E.P. by an EMBO Long-Term Fellowship; M.Y.T. holds a Royal Society Dorothy Hodgkin Fellowship and A.M. has a Wellcome Trust Investigator award. The human embryonic and fetal material was provided by the Joint MRC/Wellcome Trust (MR/R006237/1) HDBR.
Reviewer information
Nature thanks B. Treutlein and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
R.V.-T. and S.A.T. conceived the study. Sample and library preparation was performed by R.V.-T. with contributions from M.Y.T., J.-E.P., E.S. and S.L.; FACS experiments were performed by R.V.-T., R.A.B., A.F., A.M.S., R.P.P. and M.A.I.; histology staining was performed by J.N.B., L.G., R.V.-T., M.Y.T., B.M., B.I., S.H., D.H.R. and A.W.-C.; M.E. and R.V.-T. analysed and interpreted the data with contributions from M.V.-T., M.J.T.S., L.W., G.J.W., A.G., A.Z., J.H., K.B.M., K.P., M.H., A.M. and S.A.T.; R.V.-T., A.M. and S.A.T. wrote the manuscript with contributions from M.H., M.E., K.B.M. and M.Y.T.; M.H., A.M. and S.A.T. co-directed the study. All authors read and accepted the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Gating strategy for Smart-seq2 data.
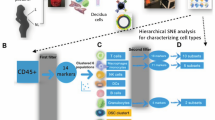
a, Gating strategy for a panel of 14 antibodies to analyse immune cells in decidual samples by Smart-seq2 (CD3, CD4, CD8, CD9, CD14, CD16, CD19, CD20, CD34, CD45, CD56, CD94, DAPI, HLA-DR and HLA-G). Cells isolated for Smart-seq2 data were gated on live; CD19- and CD20-negative, singlets and the following cell types were sorted: (i) CD45+CD14highHLA-DRhigh; (ii) CD45+HLA-DR+; (iii) CD45+HLA-DR−CD56−CD3+CD4+CD8−; (iv) CD45+HLA-DR−CD56−CD3+CD8+; (v) CD45+HLA-DR−CD56−CD3+CD4−CD8−; (vi) CD45+HLA-DR−CD3−CD56−CD94− (labelled ‘all -’ on the figure); (vii) CD45+HLA-DR−CD3−CD56+CD94−; (viii) autofluorescence; (ix) CD45+HLA-DR−CD3−CD56+CD94+CD9−; (x) CD45+HLA-DR−CD3−CD56+CD94+CD9+; (xi) CD45−HLA-G+; (xii) CD45−HLA-G−. Sample F9 is shown as an example. Cells from different gates were sorted in different plates: myeloid cells (gates (i) and (ii)); T cells (gates (iii), (iv) and (v)); natural killer cells (gates (vi), (vii), (viii), (ix) and (x)); CD45− (gates (xi) and (xii)). Antibody information is provided in Supplementary Table 10.
Extended Data Fig. 2 Quality control of droplet and Smart-seq2 datasets.
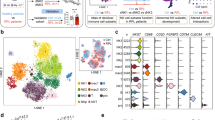
a, Histograms show the distribution of the cells from the Smart-seq2 dataset ordered by number of detected genes and mitochondrial gene expression content. b, Histograms show the distribution of the cells from the droplet-based dataset ordered by number of detected genes and mitochondrial gene expression content. c, Total numbers of cells that passed the quality control, processed by Smart-seq2 and droplet scRNA-seq. Each row is a separate donor. d, Canonical correlation vectors (CC1 and CC2) of integrated analysis of decidual and placental cells from the Smart-seq2 (n = 5 deciduas, n = 2 peripheral blood samples) and droplet-based datasets (n = 5 placentas, n = 6 deciduas and n = 4 blood samples), coloured on the basis of their assignment to clusters and the technology that was used for scRNA-seq.
Extended Data Fig. 3 Overview of droplet and Smart-seq2 datasets.
a, UMAP plot showing the integration of the Smart-seq2 and droplet-based dataset and the log-transformed expression of MKI67 (which marks proliferating cells). b, UMAP plots showing the separate and more-detailed integration analysis of the cells from cluster 14 (perivascular cells), cluster 19 (endothelial cells) and cluster 25 (epithelial cells). Clusters are labelled as in Fig. 1c. c, UMAP visualization of T cell clusters obtained by integrating Smart-seq2 and droplet-based T cells subpopulations (clusters 4, 8, 10 and 15) from Fig. 1c. Cells are coloured by the tissue of origin (top) and the identified clusters (bottom). d, Heat map showing the z-score of the mean log-transformed, normalized counts for each cluster of selected marker genes used to annotate clusters. For a more extensive set of genes, see Supplementary Table 2. Adjusted P value < 0.1; Wilcoxon rank-sum test with Bonferroni correction. NK, natural killer cells; NKp, proliferating natural killer cells; MO, monocytes; Granulo, granulocytes; Treg, regulatory T cells; GD, γδ T cells; CD8c, cytotoxic CD8+ T cells; Plasma, plasma cells. e, log-likelihood differences between assignment to fetal versus assignment to maternal origin of cells, on the basis of single nucleotide polymorphism calling from the droplet RNA-seq data. Cells are coloured by their assignment as determined by demuxlet. For this figure, we used n = 5 placentas, n = 6 deciduas and n = 4 blood individuals. f, UMAP visualization of the log-transformed, normalized expression of selected marker genes of the M3 subpopulation.
Extended Data Fig. 4 Cell–cell communication networks in the maternal–fetal interface using CellPhoneDB.
a, Information aggregated within www.CellPhoneDB.org. b, Statistical framework used to infer ligand–receptor complex specific to two cell types from single-cell transcriptomics data. Predicted P values for a ligand–receptor complex across two cell clusters are calculated using permutations, in which cells are randomly re-assigned to clusters (see Methods) c, Networks visualizing potential specific interactions in the decidua, in which nodes are clusters (cell types) and edges represent the number of significant ligand–receptor pairs. The network was created for edges with more than 30 interactions and the network layout was set to force-directed layout. Only droplet data were considered for the CellPhoneDB analysis (n = 6 deciduas). d, Networks visualizing potential specific interactions in the placenta, in which nodes are clusters and edges represent the number of significant ligand–receptor pairs. The network layout was set to force-directed layout. Only droplet data were considered for the analysis (n = 5 placentas). e, An example of significant interactions identified by CellPhoneDB. Violin plots show log-transformed, normalized expression levels of the components of the IL6–IL6R complex in placental cells. IL6 expression is enriched in the fibroblast 2 cluster (F2; dark brown in d) and the two subunits of the IL6 receptors (IL6R and IL6ST) are co-expressed in Hofbauer cells.
Extended Data Fig. 5 Trophoblast analysis.
a, UMAP visualization of the integrated analysis of the trophoblast subpopulations that were used for pseudotime analysis, including the enriched EPCAM+ and HLA-G+ cells (see Methods). Cells that were excluded from the pseudotime analysis are coloured in grey (n = 5 placentas, n = 11 deciduas). b, UMAP visualization of the log-transformed, normalized expression of selected canonical trophoblast marker genes (n = 5 placentas). c, Visualization of log-transformed, normalized expression of HLA-G, MKI67 and LGALS13 across trophoblast differentiation. d, Heat map showing genes that are involved in the epithelial–mesenchymal transition, identified as varying significantly as EVT differentiate (q value < 0.1, likelihood ratio test, P values were adjusted for the false discovery rate).
Extended Data Fig. 6 Steroid synthesis.
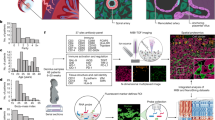
a, Heat map showing relative expression of enzymes involved in cholesterol and steroid synthesis in the three stromal subsets (n = 11 deciduas). b, Multiplexed smFISH in two decidua parietalis sections from two different individuals, showing an enrichment of CYP11A1 expression in the decidua compacta. Section stained by CYP11A1, LDLR and DAPI. Images are shown at 40× magnification. A high resolution is needed to detect differences between the sections (n = 2 individuals).
Extended Data Fig. 7 In situ staining for the different stromal cells.
a, Immunohistochemistry of decidual serial sections stained for cytokeratin (uterine glands), CD34 (endothelial cells), ACTA2 (perivascular populations and dS1) and IGFBP1 (stromal cells and glandular secretions) (n = 2 biological replicates). ACTA2+ stromal cells are confined to the stromal cells of the deeper decidua spongiosa, whereas stromal cells in the decidua compacta are ACTA2−. IGFBP1+ stromal cells are enriched in the decidua compacta, whereas stromal cells around the glands in the decidua spongiosa are IGFBP1−. Glandular secretions are IGFBP1+. b, Multiplexed smFISH for a decidua parietalis section showing the two decidual layers. ACTA2, dS1 population confined to decidua spongiosa; IGBP1 and PRL, dS2 and dS3 populations confined to decidua compacta. Samples shown are from a different individual than samples shown in Fig. 4d (n = 2 biological replicates). c, Multiplexed smFISH for a decidua parietalis section showing the two decidual layers. DKK1, decidual stromal marker; ACTA2, dS1 population confined to decidua spongiosa; PRL, dS3 population confined to decidua compacta (n = 1 biological replicate).
Extended Data Fig. 8 Lymphocyte populations in the decidua.
a, Heat map showing z-scores of the mean log-transformed, normalized expression of selected genes in the lymphocyte populations. Proliferating dNK cells (dNKp) are excluded from the analysis (n = 11 deciduas). b, FACS gating strategy in Fig. 5 applied in matched blood. Matched blood for the sample shown in Fig. 5 (n = 2 biological replicates). c, Morphology of dNK1, dNK2 and dNK3 subsets by Giemsa–Wright stain after cytospin (representative data from 1 of n = 2 biological replicates are shown). Scale bar, 10 μm.
Extended Data Fig. 9 Expression of ligands and receptors at the maternal–fetal interface.
a, Heat map showing z-scores of the mean log-transformed, normalized expression of genes annotated as cytokines, growth factors, hormones and angiogenic factors with a log-mean > 0.1 in the selected decidual immune populations (n = 11 deciduas). b, Violin plots showing log-transformed, normalized expression levels of selected ligands expressed in the three dNK cells and their corresponding receptors expressed on other decidual cells and EVT (CD39, CD73, ADORA3, CSF1, CSF1R, CCL5, CCR1, XCL1 and XCR1; n = 11 deciduas, n = 5 placentas) c, Immunohistochemistry images of serial decidual sections stained for the EVT marker HLA-G and the inhibitory ligand PDL1. Bottom panels shown the areas in white boxes in the top panels at higher power. HLA-G+ cells are only present at the site of placentation (decidua basalis) and are absent elsewhere (decidua parietalis). SpA, spiral arteries. The EVT is strongly PDL1+. We show representative data from one individual of n = 5 biological replicates. d, Immunohistochemistry images of decidual serial sections of the decidual implantation site (at 10 weeks of gestation), stained for the trophoblast cell marker, cytokeratin-7 (red arrow) and the inhibitory receptor KIR2DL1 on a natural killer cell (black arrow). The asterisk marks the lumen of a spiral artery that supplies the conceptus. We show representative data from one individual of n = 5 samples).
Extended Data Fig. 10 Encyclopaedia of cells at the maternal–fetal interface.
a, Summary of populations from our scRNa-seq data. Blue, fetal; red, maternal.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables 1-10 and an SI Tables Guide.
Rights and permissions
About this article
Cite this article
Vento-Tormo, R., Efremova, M., Botting, R.A. et al. Single-cell reconstruction of the early maternal–fetal interface in humans. Nature 563, 347–353 (2018). https://doi.org/10.1038/s41586-018-0698-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0698-6
Keywords
This article is cited by
-
Decoding leukemia at the single-cell level: clonal architecture, classification, microenvironment, and drug resistance
Experimental Hematology & Oncology (2024)
-
Molecular epidemiology of pregnancy using omics data: advances, success stories, and challenges
Journal of Translational Medicine (2024)
-
Increased miR-3074-5p expression promotes M1 polarization and pyroptosis of macrophages via ERα/NLRP3 pathway and induces adverse pregnancy outcomes in mice
Cell Death Discovery (2024)
-
Single-cell profiling of the microenvironment in human bone metastatic renal cell carcinoma
Communications Biology (2024)
-
The transcriptional regulatory network modulating human trophoblast stem cells to extravillous trophoblast differentiation
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.