Abstract
Inspired by the period-four oscillation in flash-induced oxygen evolution of photosystem II discovered by Joliot in 1969, Kok performed additional experiments and proposed a five-state kinetic model for photosynthetic oxygen evolution, known as Kok’s S-state clock or cycle1,2. The model comprises four (meta)stable intermediates (S0, S1, S2 and S3) and one transient S4 state, which precedes dioxygen formation occurring in a concerted reaction from two water-derived oxygens bound at an oxo-bridged tetra manganese calcium (Mn4CaO5) cluster in the oxygen-evolving complex3,4,5,6,7. This reaction is coupled to the two-step reduction and protonation of the mobile plastoquinone QB at the acceptor side of PSII. Here, using serial femtosecond X-ray crystallography and simultaneous X-ray emission spectroscopy with multi-flash visible laser excitation at room temperature, we visualize all (meta)stable states of Kok’s cycle as high-resolution structures (2.04–2.08 Å). In addition, we report structures of two transient states at 150 and 400 µs, revealing notable structural changes including the binding of one additional ‘water’, Ox, during the S2→S3 state transition. Our results suggest that one water ligand to calcium (W3) is directly involved in substrate delivery. The binding of the additional oxygen Ox in the S3 state between Ca and Mn1 supports O–O bond formation mechanisms involving O5 as one substrate, where Ox is either the other substrate oxygen or is perfectly positioned to refill the O5 position during O2 release. Thus, our results exclude peroxo-bond formation in the S3 state, and the nucleophilic attack of W3 onto W2 is unlikely.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
20 November 2018
Figs. 1, 2 and 3 files initially published online were corrupted in the HTML.
References
Kok, B., Forbush, B. & McGloin, M. Cooperation of charges in photosynthetic O2 evolution—I. A linear four step mechanism. Photochem. Photobiol. 11, 457–475 (1970).
Joliot, P., Barbieri, G. & Chabaud, R. A new model of photochemical centers in system-2. Photochem. Photobiol. 10, 309–329 (1969).
Hillier, W. & Wydrzynski, T. 18O-water exchange in photosystem II: substrate binding and intermediates of the water splitting cycle. Coord. Chem. Rev. 252, 306–317 (2008).
Yano, J. & Yachandra, V. Mn4Ca cluster in photosynthesis: where and how water is oxidized to dioxygen. Chem. Rev. 114, 4175–4205 (2014).
Cox, N. & Messinger, J. Reflections on substrate water and dioxygen formation. Biochim. Biophys. Acta 1827, 1020–1030 (2013).
Debus, R. J. FTIR studies of metal ligands, networks of hydrogen bonds, and water molecules near the active site Mn4CaO5 cluster in photosystem II. Biochim. Biophys. Acta 1847, 19–34 (2015).
Klauss, A., Haumann, M. & Dau, H. Seven steps of alternating electron and proton transfer in photosystem II water oxidation traced by time-resolved photothermal beam deflection at improved sensitivity. J. Phys. Chem. B 119, 2677–2689 (2015).
Young, I. D. et al. Structure of photosystem II and substrate binding at room temperature. Nature 540, 453–457 (2016).
Suga, M. et al. Light-induced structural changes and the site of O=O bond formation in PSII caught by XFEL. Nature 543, 131–135 (2017).
Kern, J. et al. Simultaneous femtosecond X-ray spectroscopy and diffraction of photosystem II at room temperature. Science 340, 491–495 (2013).
Kulik, L. V., Epel, B., Lubitz, W. & Messinger, J. Electronic structure of the Mn4OxCa cluster in the S0 and S2 states of the oxygen-evolving complex of photosystem II based on pulse 55Mn-ENDOR and EPR spectroscopy. J. Am. Chem. Soc. 129, 13421–13435 (2007).
Dau, H. & Haumann, M. The manganese complex of photosystem II in its reaction cycle—basic framework and possible realization at the atomic level. Coord. Chem. Rev. 252, 273–295 (2008).
Peloquin, J. M. et al. 55Mn ENDOR of the S2-state multiline EPR signal of photosystem II: Implications on the structure of the tetranuclear cluster. J. Am. Chem. Soc. 122, 10926–10942 (2000).
Krewald, V. et al. Metal oxidation states in biological water splitting. Chem. Sci. (Camb.) 6, 1676–1695 (2015).
Chernev, P. et al. Merging structural information from X-ray crystallography, quantum chemistry, and EXAFS spectra: The oxygen-evolving complex in PSII. J. Phys. Chem. B 120, 10899–10922 (2016).
Tanaka, A., Fukushima, Y. & Kamiya, N. Two different structures of the oxygen-evolving complex in the same polypeptide frameworks of photosystem II. J. Am. Chem. Soc. 139, 1718–1721 (2017).
Yamaguchi, K. et al. Theory of chemical bonds in metalloenzymes XXI. Possible mechanisms of water oxidation in oxygen evolving complex of photosystem II. Mol. Phys. 116, 717–745 (2018).
Yano, J. et al. Where water is oxidized to dioxygen: structure of the photosynthetic Mn4Ca cluster. Science 314, 821–825 (2006).
Pantazis, D. A., Ames, W., Cox, N., Lubitz, W. & Neese, F. Two interconvertible structures that explain the spectroscopic properties of the oxygen-evolving complex of photosystem II in the S2 state. Angew. Chem. Int. Ed. 51, 9935–9940 (2012).
Retegan, M. et al. A five-coordinate Mn(IV) intermediate in biological water oxidation: spectroscopic signature and a pivot mechanism for water binding. Chem. Sci. 7, 72–84 (2016).
Dau, H. & Haumann, M. Considerations on the mechanism of photosynthetic water oxidation — dual role of oxo-bridges between Mn ions in (i) redox-potential maintenance and (ii) proton abstraction from substrate water. Photosynth. Res. 84, 325–331 (2005).
Cox, N. et al. Photosynthesis. Electronic structure of the oxygen-evolving complex in photosystem II prior to O-O bond formation. Science 345, 804–808 (2014).
Siegbahn, P. E. M. Structures and energetics for O2 formation in photosystem II. Acc. Chem. Res. 42, 1871–1880 (2009).
de Wijn, R. & van Gorkom, H. J. Kinetics of electron transfer from QA to QB in photosystem II. Biochemistry 40, 11912–11922 (2001).
Sakamoto, H., Shimizu, T., Nagao, R. & Noguchi, T. Monitoring the reaction process during the S2→S3 transition in photosynthetic water oxidation using time-resolved infrared spectroscopy. J. Am. Chem. Soc. 139, 2022–2029 (2017).
Rossini, E. & Knapp, E.-W. Protonation equilibria of transition metal complexes: from model systems toward the Mn-complex in photosystem II. Coord. Chem. Rev. 345, 16–30 (2017).
Askerka, M., Wang, J., Vinyard, D. J., Brudvig, G. W. & Batista, V. S. S3 state of the O2-evolving complex of photosystem II: insights from QM/MM, EXAFS, and femtosecond X-ray diffraction. Biochemistry 55, 981–984 (2016).
Boussac, A. & Rutherford, A. W. Nature of the inhibition of the oxygen-evolving enzyme of photosystem II induced by NaCl washing and reversed by the addition of Ca2+ or Sr2+. Biochemistry 27, 3476–3483 (1988).
Tso, J., Sivaraja, M. & Dismukes, G. C. Calcium limits substrate accessibility or reactivity at the manganese cluster in photosynthetic water oxidation. Biochemistry 30, 4734–4739 (1991).
Suzuki, H., Sugiura, M. & Noguchi, T. Monitoring water reactions during the S-state cycle of the photosynthetic water-oxidazing center: detection of the DOD bending vibration by means of Fourier transform infrared spectroscopy. Biochemistry 47, 11024–11030 (2008).
Barber, J. A mechanism for water splitting and oxygen production in photosynthesis. Nat. Plants 3, 17041 (2017).
Hellmich, J. et al. Native-like photosystem II superstructure at 2.44 Å resolution through detergent extraction from the protein crystal. Structure 22, 1607–1615 (2014).
Ibrahim, M. et al. Improvements in serial femtosecond crystallography of photosystem II by optimizing crystal uniformity using microseeding procedures. Struct. Dyn. 2, 041705 (2015).
Guskov, A. et al. Cyanobacterial photosystem II at 2.9-Å resolution and the role of quinones, lipids, channels and chloride. Nat. Struct. Mol. Biol. 16, 334–342 (2009).
Krivanek, R., Kern, J., Zouni, A., Dau, H. & Haumann, M. Spare quinones in the QB cavity of crystallized photosystem II from Thermosynechococcus elongatus. Biochim. Biophys. Acta 1767, 520–527 (2007).
Fransson, T. et al. X-ray emission spectroscopy as an in situ diagnostic tool for X-ray crystallography of metalloproteins using an X-ray free-electron laser. Biochemistry 57, 4629–4637 (2018).
Fuller, F. D. et al. Drop-on-demand sample delivery for studying biocatalysts in action at X-ray free-electron lasers. Nat. Methods 14, 443–449 (2017).
Kern, J. et al. Taking snapshots of photosynthetic water oxidation using femtosecond X-ray diffraction and spectroscopy. Nat. Commun. 5, 4371 (2014).
Yano, J. et al. in Sustaining Life on Planet Earth: Metalloenzymes Mastering Dioxygen and Other Chewy Gases, Metal Ions in Life Sciences (eds Kroneck, P. M. H. & Sosa Torres, M. E.) 13–43 (Springer International Publishing, 2015).
Beckmann, K., Messinger, J., Badger, M. R., Wydrzynski, T. & Hillier, W. On-line mass spectrometry: membrane inlet sampling. Photosynth. Res. 102, 511–522 (2009).
Emma, P. et al. First lasing and operation of an Ångström-wavelength free-electron laser. Nat. Photon. 4, 641–647 (2010).
Boutet, S., Cohen, A. & Wakatsuki, S. The new macromolecular femtosecond crystallography (MFX) instrument at LCLS. Synchr. Radiat. News 29, 23–28 (2016).
Sauter, N. K. XFEL diffraction: developing processing methods to optimize data quality. J. Synchr. Radiat. 22, 239–248 (2015).
Winter, G. et al. DIALS: implementation and evaluation of a new integration package. Acta Crystallogr. D Struct. Biol. 74, 85–97 (2018).
Sauter, N. K., Hattne, J., Grosse-Kunstleve, R. W. & Echols, N. New Python-based methods for data processing. Acta Crystallogr. D Biol. Crystallogr. 69, 1274–1282 (2013).
Hattne, J. et al. Accurate macromolecular structures using minimal measurements from X-ray free-electron lasers. Nat. Methods 11, 545–548 (2014).
Sauter, N. K. et al. Improved crystal orientation and physical properties from single-shot XFEL stills. Acta Crystallogr. D Biol. Crystallogr. 70, 3299–3309 (2014).
Waterman, D. G. et al. Diffraction-geometry refinement in the DIALS framework. Acta Crystallogr. D Struct. Biol. 72, 558–575 (2016).
Zeldin, O. B. et al. Data Exploration Toolkit for serial diffraction experiments. Acta Crystallogr. D Biol. Crystallogr. 71, 352–356 (2015).
Andrews, L. C. & Bernstein, H. J. The geometry of Niggli reduction: BGAOL - embedding Niggli reduction and analysis of boundaries. J. Appl. Crystallogr. 47, 346–359 (2014).
Suga, M. et al. Native structure of photosystem II at 1.95 Å resolution viewed by femtosecond X-ray pulses. Nature 517, 99–103 (2015).
Uervirojnangkoorn, M. et al. Enabling X-ray free electron laser crystallography for challenging biological systems from a limited number of crystals. eLife 4, e05421 (2015).
Lyubimov, A. Y. et al. Advances in X-ray free electron laser (XFEL) diffraction data processing applied to the crystal structure of the synaptotagmin-1/SNARE complex. eLife 5, e18740 (2016).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D Biol. Crystallogr. 68, 352–367 (2012).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Wieghardt, K. The active-sites in manganese-containing metalloproteins and inorganic model complexes. Angew. Chem. Int. Edn Engl. 28, 1153–1172 (1989).
Cinco, R. M. et al. Comparison of the manganese cluster in oxygen-evolving photosystem II with distorted cubane manganese compounds through X-ray absorption spectroscopy. Inorg. Chem. 38, 5988–5998 (1999).
Mukhopadhyay, S., Mandal, S. K., Bhaduri, S. & Armstrong, W. H. Manganese clusters with relevance to photosystem II. Chem. Rev. 104, 3981–4026 (2004).
Law, N. A., Caudle, M. T. & Pecoraro, V. L. Manganese redox enzymes and model systems: Properties, structures, and reactivity. Adv. Inorg. Chem. 46, 305–440 (1999).
Tsui, E. Y., Kanady, J. S. & Agapie, T. Synthetic cluster models of biological and heterogeneous manganese catalysts for O2 evolution. Inorg. Chem. 52, 13833–13848 (2013).
Schrödinger LLC. The pymol molecular graphics system, version 1.8. (2015).
Lang, P. T., Holton, J. M., Fraser, J. S. & Alber, T. Protein structural ensembles are revealed by redefining X-ray electron density noise. Proc. Natl Acad. Sci. USA 111, 237–242 (2014).
Brewster, A. S. et al. Improving signal strength in serial crystallography with DIALS geometry refinement. Acta Crystallogr. D Struct. Biol. 74, 877–894 (2018).
Kupitz, C. et al. Serial time-resolved crystallography of photosystem II using a femtosecond X-ray laser. Nature 513, 261–265 (2014).
Ho, F. M. & Styring, S. Access channels and methanol binding site to the CaMn4 cluster in Photosystem II based on solvent accessibility simulations, with implications for substrate water access. Biochim. Biophys. Acta 1777, 140–153 (2008).
Vassiliev, S., Zaraiskaya, T. & Bruce, D. Exploring the energetics of water permeation in photosystem II by multiple steered molecular dynamics simulations. Biochim. Biophys. Acta 1817, 1671–1678 (2012).
Murray, J. W. & Barber, J. Structural characteristics of channels and pathways in photosystem II including the identification of an oxygen channel. J. Struct. Biol. 159, 228–237 (2007).
Gabdulkhakov, A. et al. Probing the accessibility of the Mn4Ca cluster in photosystem II: channels calculation, noble gas derivatization, and cocrystallization with DMSO. Structure 17, 1223–1234 (2009).
Umena, Y., Kawakami, K., Shen, J.-R. & Kamiya, N. Crystal structure of oxygen-evolving photosystem II at a resolution of 1.9 Å. Nature 473, 55–60 (2011).
Sakashita, N., Watanabe, H. C., Ikeda, T. & Ishikita, H. Structurally conserved channels in cyanobacterial and plant photosystem II. Photosynth. Res. 133, 75–85 (2017).
Acknowledgements
This work was supported by the Director, Office of Science, Office of Basic Energy Sciences (OBES), Division of Chemical Sciences, Geosciences, and Biosciences (CSGB), Department of Energy (DOE) (J.Y., V.K.Y.), by National Institutes of Health (NIH) grants GM055302 (V.K.Y.), GM110501 (J.Y.) GM126289 (J.K.), GM117126 (N.K.S.), GM124149 and GM124169 (J.M.H.), the Ruth L. Kirschstein National Research Service Award (GM116423-02, F.D.F.), and Human Frontiers Science Project RGP0063/2013 (J.Y., U.B., A.Z.). We acknowledge the DFG-Cluster of Excellence “UniCat” coordinated by T.U. Berlin and Sfb1078, TP A5 (A.Z., H.D.); the Artificial Leaf Project (K&A Wallenberg Foundation 2011.0055) and Vetenskapsrådet (2016-05183) (J.M.); Diamond Light Source, Biotechnology and Biological Sciences Research Council (grant 102593) and a Strategic Award from the Wellcome Trust (A.M.O.). This research used NERSC, supported by DOE, under Contract No. DE-AC02-05CH11231. Synchrotron facilities at the ALS, Berkeley and SSRL, Stanford, were funded by DOE OBES. The SSRL Structural Molecular Biology Program is supported by the DOE OBER, and NIH (P41GM103393). LCLS and SSRL, SLAC National Accelerator Laboratory, are supported by DOE, OBES under Contract No. DE-AC02-76SF00515. We thank the staff at LCLS/SLAC and SSRL (BL 6-2, 7-3) and ALS (BL 5.01, 5.0.2, 8.2.1, 8.3.1).
Author information
Authors and Affiliations
Contributions
U.B., V.K.Y. and J.Y. conceived the experiment; R.A.-M., A.Z., J.M., U.B., N.K.S., J.K., V.K.Y. and J.Y. designed the experiment; R.C., M.I., L.L., R.H., M.Z., L.D., J.W., I.S., A.Z. and J.K. prepared samples; A. Batyuk, M.L., S.B., R.A.-M., J.E.K. and S.C. operated the MFX instrument; F.D.F., S.G., E.P., P.A., A.M.O., J.M. and J.K. developed, tested and ran the sample delivery system; M.H.C., D. Shevela, R.C., C.d.L., J.Y. and J.M. performed and analysed O2 evolution and EPR measurements; R.C., F.D.F., S.G., M.I., C.d.L., M.H.C., I.D.Y., A.S.B., R.A.-M., R.H., M.Z., L.L., L.D., D. Sokaras, E.P., C.W., T.F., T.K., R.G.S., P.A., A. Butryn, A. Batyuk, M.L., S.B., J.E.K., S.C., A.M.O., A.Z., J.M., U.B., N.K.S., J.K., V.K.Y. and J.Y. performed the LCLS experiment; I.D.Y., A.S.B., N.W.M., J.M.H., P.D.A. and N.K.S. developed new software for data processing; I.D.Y., A.S.B., L.L., F.D.F., C.W., T.F., P.A., H.D., J.M.H., N.K.S. and J.K. processed and analysed XFEL data; J.M., J.K., V.K.Y. and J.Y. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Overview of the PSII structure and electron density maps of the 3F state.
a, Structure of the native PSII homodimer. In the left monomer the location of cofactors for the initial charge separation (P680, PheoD1), and for the electron transfer leading to the reduction of the plastoquinone (QA, QB) at the acceptor side and to the oxidation of the OEC at the donor side by P680+ are indicated. In the right monomer, the locations of the protein subunits are displayed. b–d, 2mFobs − DFcalc map (blue, 1.5σ contour) obtained from the room temperature 3F data set. b, Density around the main chain and a chlorophyll. c, Well-resolved ordered water molecules. d, Chlorophyll and pheophytin molecules with well-resolved tails. e, Clear density in hydrophobic regions and along cofactor hydrocarbon tails.
Extended Data Fig. 2 Flash-induced S-state turnover of PSII micro crystals.
a, Change of the first moment of the in situ-measured Mn Kβ XES as a function of flashes and fit to the data. b, Flash-induced O2 yield as measured by MIMS as a function of flash number and fit to the data. c, The estimated S-state population (%) for each of the flash states from fitting of the XES data and of the flash-induced O2 evolution pattern of a suspension of PSII crystals at pH 6.5. Two different fits were performed: a global fit of both O2 and XES data using an equal miss parameter of 22% and 100% S1 population in the 0F sample (black traces in a, b; S-state distribution listed in the columns headed O2), and a direct fit of the XES data using a 8% miss parameter in the S1→S2 and a 27% miss parameter for the S2→S3 and S3→S0 transitions (XES in c). For the XES fit, shifts of −0.06 eV per oxidation state increase for all S states were assumed. The XES raw spectra are published elsewhere36.
Extended Data Fig. 3 Isomorphous difference maps in the second monomer at the QB site.
a–c, Fobs − Fobs maps contoured at 3σ at plastoquinone QB in monomer a. a, 1F − 0F difference map matching reduction of the plastoquinone to a semiquinone and concomitant slight geometry change. b, 2F − 0F difference map matching replacement of the fully reduced quinol with another quinone at the original position. c, 3F − 0F difference map, showing again structural changes similar to the 1F − 0F map, indicating formation of the semiquinone. Similar views are shown for monomer A in Fig. 1d–f and comparison of both monomers indicates similar flash-induced changes in both monomers.
Extended Data Fig. 4 Movement of ligands around the OEC in the different S states.
a, Overview of the ligand environment of the OEC, showing the dark state (0F) structure. Coordination of the OEC by nearby side chains and water molecules is indicated by dashed lines. b–g, Trends for selected individual side chains in both monomers (b–d, monomer A; e–g, monomer a). Overlays of the refined models at the OEC following least-squares fitting of subunit D1 residues 55–65, 160–190 and 328, subunit CP43 residues 328 and 354–358, and chain D residue 352 of each other model to the 0F model. The largest and most consistent motions of side chains near the OEC through the sequence of illuminated state models are annotated with arrows indicating the trend. A motion observed in only one monomer is indicated by a dashed line.
Extended Data Fig. 5 Impact of the data quality on the resolving power of the maps.
a–f, The data quality evidenced by 2F state models and 2mFobs − DFcalc maps contoured at 1.5σ. a, 5TIS (2F, 2.25 Å) model and map. Overlays indicate atom numbering in the OEC and the identities of selected coordinating sidechains. b, Current 2F model and map cut to 2.25 Å. c, Current 2F model and map at the full 2.07 Å resolution. Emergence of locations of O4 with improved data quality is indicated by bold arrow. d, As in a from a different angle and with mFobs − DFcalc density at 3σ indicating the lack of sufficient evidence for inserting an additional O atom at a chemically reasonable position. e, f, As in b, c from the same direction as d and with mFobs − DFcalc density at 3σ shown to 2.25 and 2.05 Å, respectively, after omitting the inserted Ox atom. Centring of the refined Ox position within the omit density gives a clear indication of the position of the inserted water in the S3 state with the current, higher-quality data, even when artificially cut to the same resolution as the previous data set. g, mFobs − DFcalc maps of the 2F data that compare the O6 model from Suga et al.9 and the Ox model from the current study. Map shows the mFobs − DFcalc density calculated with our current 2F data and our model adding the O6 position of Suga et al.9 (with the occupancy of 0.7 and B-factor of 30) (g-1), and with our Ox model (g-2). We see clearly a positive density for the missing Ox and a negative density at the O6 position in g-1. Schematics of the O6 and Ox S3 models are shown on the left.
Extended Data Fig. 6 Isomorphous difference maps in the second monomer at the OEC.
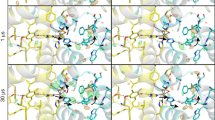
Isomorphous difference density OEC sites in monomer a. Fobs − Fobs difference densities between the various illuminated states and the 0F data are contoured at +3σ (blue) and −3σ (orange). The model for the 0F data is shown in light grey whereas carbons are coloured as follows: 1F (cyan), 2F (150 μs) (green), 2F (400 μs) (yellow) and 2F (0.2 s) (blue).
Extended Data Fig. 7 Water environment of the OEC.
a, Extended schematic of the hydrogen bonding network connecting the OEC to the solvent-exposed surface of PSII and identification of several channels for either possible water movement or proton transfer. Top right, locations of four selected channels in the PSII monomer. b–e, Movements within the water networks across monomers. Coloured spheres are shown for each ordered water or chloride ion across the four metastable states, 0F through 3F, and for both monomers, with the stronger colour matching the first (A) monomer and the lighter colour matching the second (a) monomer. For ordered solvent, residue number is shown; for OEC atoms, the atom identifier is shown; and for the Cl2 site, the Cl− 680 label is shown. b, The O1 water chain. Positional disagreement between monomers is visible especially near waters 77 (2F) and 27 (3F) and is on the same scale as changes between illuminated states, both of which may indicate a more dynamic water channel. c, The O4 water chain. With the notable exception of water 20, most water positions are stable across monomers and illuminated states. Water 20 is highly unstable in position in the two states (0F, 3F) in which it is modelled, and there is not sufficient density in the remaining states to model a water 20 position. d, The Cl1 site water channel with no notable movements. e, The Cl2 site water channel with no notable movements. f, Indication of a split position of W3 in the S0 state. mFobs − DFcalc difference density (green mesh) in the 3F state suggests an alternate position near W3 (W3b in Fig. 4d). g, h, Possible access to W3/Ox side from the Cl1 or the O1 channel. The surface of the protein is shown in grey to visualize the extent of the cavities around the OEC, and Van der Waals radii are indicated for selected residues or atoms by dotted spheres. Shown are two different views for each channel. The direction of the Cl1 channel is indicated by a green arrow and the O1 channel by a pink arrow. Water W2 is shown in purple, W3 in cyan and Ox in orange. Yellow spheres indicate other waters. Mn are shown in magenta, other bridging oxygens as red spheres.
Supplementary information
41586_2018_681_MOESM2_ESM.mov
Video 1 Fobs-Fobs isomorphous difference density around Mn1 and Mn4 of the OEC from the 0F to the 2F state. The video shows the isomorphous difference density Fobs-Fobs for the 1F, 2F (150 μs), 2F (400 μs) and 2F (200 ms) data with the 0F data in the region of Mn1 and Mn4 of the OEC. Density is shown as orange (negative) and blue (positive) surfaces at 3 σ contour level.
41586_2018_681_MOESM3_ESM.mov
Video 2 Fobs-Fobs isomorphous difference density between the 1F and 0F state at the Water 20 site. The video shows the extent and location of the main peak in the isomorphous difference density in the vicinity of the OEC. It is located around water W20, indicating disappearance/disordering of this water upon the formation of the S2 state. Density is shown as orange (negative) and blue (positive) surfaces at 3 σ contour level.
Source data
Rights and permissions
About this article
Cite this article
Kern, J., Chatterjee, R., Young, I.D. et al. Structures of the intermediates of Kok’s photosynthetic water oxidation clock. Nature 563, 421–425 (2018). https://doi.org/10.1038/s41586-018-0681-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0681-2
Keywords
This article is cited by
-
Towards understanding the crystallization of photosystem II: influence of poly(ethylene glycol) of various molecular sizes on the micelle formation of alkyl maltosides
Photosynthesis Research (2024)
-
International conference on “Photosynthesis and Hydrogen Energy Research for Sustainability-2023”: in honor of Robert Blankenship, Győző Garab, Michael Grätzel, Norman Hüner and Gunnar Öquist
Photosynthesis Research (2024)
-
On the simulation and interpretation of substrate-water exchange experiments in photosynthetic water oxidation
Photosynthesis Research (2024)
-
Indirect interactions involving the PsbM or PsbT subunits and the PsbO, PsbU and PsbV proteins stabilize assembly and activity of Photosystem II in Synechocystis sp. PCC 6803
Photosynthesis Research (2024)
-
Oxygen-evolving photosystem II structures during S1–S2–S3 transitions
Nature (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.