Abstract
Algae produce massive amounts of dimethylsulfoniopropionate (DMSP), which fuel the organosulfur cycle1,2. On a global scale, several petagrams of this sulfur species are produced annually, thereby driving fundamental processes and the marine food web1. An important DMSP transformation product is dimethylsulfide, which can be either emitted to the atmosphere3,4 or oxidized to dimethylsulfoxide (DMSO) and other products5. Here we report the discovery of a structurally unusual metabolite, dimethylsulfoxonium propionate (DMSOP), that is synthesized by several DMSP-producing microalgae and marine bacteria. As with DMSP, DMSOP is a low-molecular-weight zwitterionic metabolite that carries both a positively and a negatively charged functional group. Isotope labelling studies demonstrate that DMSOP is produced from DMSP, and is readily metabolized to DMSO by marine bacteria. DMSOP was found in near nanomolar amounts in field samples and in algal culture media, and thus represents—to our knowledge—a previously undescribed biogenic source for DMSO in the marine environment. The estimated annual oceanic production of oxidized sulfur from this pathway is in the teragram range, similar to the calculated dimethylsulfide flux to the atmosphere3. This sulfoxonium metabolite is therefore a key metabolite of a previously undescribed pathway in the marine sulfur cycle. These findings highlight the importance of DMSOP in the marine organosulfur cycle.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The datasets generated and analysed during the current study are available from the corresponding authors upon reasonable request.
References
Ksionzek, K. B. et al. Dissolved organic sulfur in the ocean: biogeochemistry of a petagram inventory. Science 354, 456–459 (2016).
Sievert, S. M., Kiene, R. P. & Schulz-Vogt, H. N. The sulfur cycle. Oceanography (Wash. DC) 20, 117–123 (2007).
Lana, A. et al. An updated climatology of surface dimethlysulfide concentrations and emission fluxes in the global ocean. Glob. Biogeochem. Cycles 25, GB1004 (2011).
Charlson, R. J., Lovelock, J. E., Andreae, M. O. & Warren, S. G. Oceanic phytoplankton, atmospheric sulfur, cloud albedo and climate. Nature 326, 655–661 (1987).
Lee, P. A. & de Mora, S. J. A review of dimethylsulfoxide in aquatic environments. Atmosphere–ocean 37, 439–456 (1999).
Sunda, W., Kieber, D. J., Kiene, R. P. & Huntsman, S. An antioxidant function for DMSP and DMS in marine algae. Nature 418, 317–320 (2002).
Kiene, R. P., Linn, L. J. & Bruton, J. A. New and important roles for DMSP in marine microbial communities. J. Sea Res. 43, 209–224 (2000).
Alcolombri, U. et al. Identification of the algal dimethyl sulfide-releasing enzyme: a missing link in the marine sulfur cycle. Science 348, 1466–1469 (2015).
Todd, J. D. et al. Structural and regulatory genes required to make the gas dimethyl sulfide in bacteria. Science 315, 666–669 (2007).
Yoch, D. C. Dimethylsulfoniopropionate: its sources, role in the marine food web, and biological degradation to dimethylsulfide. Appl. Environ. Microbiol. 68, 5804–5815 (2002).
Asher, E. C., Dacey, J. W. H., Stukel, M., Long, M. C. & Tortell, P. D. Processes driving seasonal variability in DMS, DMSP, and DMSO concentrations and turnover in coastal Antarctic waters. Limnol. Oceanogr. 62, 104–124 (2017).
Hatton, A. D., Shenoy, D. M., Hart, M. C., Mogg, A. & Green, D. H. Metabolism of DMSP, DMS and DMSO by the cultivable bacterial community associated with the DMSP-producing dinoflagellate Scrippsiella trochoidea. Biogeochemistry 110, 131–146 (2012).
Lee, P. A. & de Mora, S. J. Intracellular dimethylsulfoxide (DMSO) in unicellular marine algae: speculations on its origin and possible biological role. J. Phycol. 35, 8–18 (1999).
Lidbury, I. et al. A mechanism for bacterial transformation of dimethylsulfide to dimethylsulfoxide: a missing link in the marine organic sulfur cycle. Environ. Microbiol. 18, 2754–2766 (2016).
Boden, R., Murrell, J. C. & Schäfer, H. Dimethylsulfide is an energy source for the heterotrophic marine bacterium Sagittula stellata. FEMS Microbiol. Lett. 322, 188–193 (2011).
Gebser, B. & Pohnert, G. Synchronized regulation of different zwitterionic metabolites in the osmoadaption of phytoplankton. Mar. Drugs 11, 2168–2182 (2013).
Spielmeyer, A. & Pohnert, G. Direct quantification of dimethylsulfoniopropionate (DMSP) with hydrophilic interaction liquid chromatography/mass spectrometry. J. Chromatogr. B 878, 3238–3242 (2010).
Carlé, J. S. & Christophersen, C. Dogger Bank Itch. 4. An eczema-causing sulfoxonium ion from the marine animal, Alcyonidium gelatinosum [Bryozoa]. Toxicon 20, 307–310 (1982).
Warabi, K. et al. Dogger Bank Itch revisited: isolation of (2-hydroxyethyl) dimethylsulfoxonium chloride as a cytotoxic constituent from the marine sponge Theonella aff. mirabilis. Comp. Biochem. Physiol. B 128, 27–30 (2001).
Curson, A. R. J. et al. Dimethylsulfoniopropionate biosynthesis in marine bacteria and identification of the key gene in this process. Nat. Microbiol. 2, 17009 (2017).
Kinsey, J. D., Kieber, D. J. & Neale, P. J. Effects of iron limitation and UV radiation on Phaeocystis antarctica growth and dimethylsulfoniopropionate, dimethylsulfoxide and acrylate concentrations. Environ. Chem. 13, 195–211 (2016).
Howard, E. C. et al. Bacterial taxa that limit sulfur flux from the ocean. Science 314, 649–652 (2006).
Curson, A. R. J., Todd, J. D., Sullivan, M. J. & Johnston, A. W. B. Catabolism of dimethylsulphoniopropionate: microorganisms, enzymes and genes. Nat. Rev. Microbiol. 9, 849–859 (2011).
Ansede, J. H., Pellechia, P. J. & Yoch, D. C. Metabolism of acrylate to beta-hydroxypropionate and its role in dimethylsulfoniopropionate lyase induction by a salt marsh sediment bacterium, Alcaligenes faecalis M3A. Appl. Environ. Microbiol. 65, 5075–5081 (1999).
Kirkwood, M., Le Brun, N. E., Todd, J. D. & Johnston, A. W. B. The dddP gene of Roseovarius nubinhibens encodes a novel lyase that cleaves dimethylsulfoniopropionate into acrylate plus dimethyl sulfide. Microbiology 156, 1900–1906 (2010).
Curson, A. R. J., Sullivan, M. J., Todd, J. D. & Johnston, A. W. B. DddY, a periplasmic dimethylsulfoniopropionate lyase found in taxonomically diverse species of Proteobacteria. ISME J. 5, 1191–1200 (2011).
Forrester, J., Jones, R. V. H., Preston, P. N. & Simpson, E. S. C. Generation of trimethylsulfonium cation from dimethyl sulfoxide and dimethyl sulfate: implications for the synthesis of epoxides from aldehydes and ketones. J. Chem. Soc. Perkin Trans. I 0, 2289–2291 (1995).
Ayres, D. C. & Hossain, A. M. M. Oxidation of aromatic substrates. Part II. The action of ruthenium tetraoxide on some derivatives of naphtalene and its monoaza-analogues. J. Chem. Soc. Perkin Trans. I 0, 707–710 (1975).
Chambers, S. T., Kunin, C. M., Miller, D. & Hamada, A. Dimethylthetin can substitute for glycine betaine as an osmoprotectant molecule for Escherichia coli. J. Bacteriol. 169, 4845–4847 (1987).
Maier, I. & Calenberg, M. Effect of extracellular Ca2+ and Ca2+-antagonists on the movement and chemoorientation of male gametes of Ectocarpus siliculosus (Phaeophyceae). Bot. Acta 107, 451–460 (1994).
Guillard, R. R. & Ryther, J. H. Studies of marine planktonic diatoms. I. Cyclotella nana Hustedt, and Detonula confervacea (Cleve) Gran. Can. J. Microbiol. 8, 229–239 (1962).
Guillard, R. R. L. & Hargraves, P. E. Stichochrysis immobilis is a diatom, not a chrysophyte. Phycologia 32, 234–236 (1993).
Spielmeyer, A., Gebser, B. & Pohnert, G. Investigations of the uptake of dimethylsulfoniopropionate by phytoplankton. ChemBioChem 12, 2276–2279 (2011).
Verity, P. G. et al. Relationships between cell volume and the carbon and nitrogen content of marine photosynthetic nanoplankton. Limnol. Oceanogr. 37, 1434–1446 (1992).
Welschmeyer, N. A. Fluorimetric analysis of chorophyll a in the presence of chlorophyll b and pheopigments. Limnol. Oceanogr. 39, 1985–1992 (1994).
Kiene, R. P. & Slezak, D. Low dissolved DMSP concentrations in seawater revealed by small-volume gravity filtration and dialysis sampling. Limnol. Oceanogr. Methods 4, 80–95 (2006).
Howard, A. G. & Russell, D. W. Borohydride-coupled HPLC–FPD instrumentation and its use in the determination of dimethylsulfonium compounds. Anal. Chem. 69, 2882–2887 (1997).
Spielmeyer, A., Gebser, B. & Pohnert, G. Dimethylsulfide sources from microalgae: improvement and application of a derivatization-based method for the determination of dimethylsulfoniopropionate and other zwitterionic osmolytes in phytoplankton. Mar. Chem. 124, 48–56 (2011).
Acknowledgements
We thank A. Curson for the provision of the A. faecalis mutant; R. Kiene and A. Rellinger, M. Galí, M. Vila, L. Viure and E. Berdalet for collection of field samples and chlorophyll a analyses during field campaigns in the northeast Pacific, Arctic and Mediterranean Sea. We acknowledge the funding by the German Research Foundation (CRC1127 ChemBioSys to G.P. and N.M.), the Max Planck Society (IMPRS BGC) and the National Science Foundation (OCE-1756907 to D.J.K.). This study was co-financed by the State of Thuringia/Thüringer Aufbaubank (2015 FGI 0021) with means of the EU in the framework of the EFRE programme.
Reviewer information
Nature thanks G. Siuzdak and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author contributions
G.P., K.T., B.G. and D.J.K. designed the research. B.G. identified DMSOP signals, performed the synthesis and the initial screening of the metabolite. K.T. performed DMSOP quantification, experiments on the biosynthesis and transformation in algae and bacteria. N.M. carried out experiments on DMSOP production and transformation in algae and performed several analytical measurements. The I. galbana growth experiment and DMSO quantification was performed by L.C. D.J.K. was responsible for field sampling and sample work-up. K.T. and N.M. performed the statistical evaluation of the data. G.P. and D.K. were the principal investigators for their respective research teams. K.T. and G.P. wrote the main drafts of the manuscript. All authors discussed the results and provided feedback and revisions to the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 DMSOP mass spectra.
The HRMS/MS spectra of natural occurring DMSOP and the authentic standard (normalized collision energy of 35) are shown. a, DMSOP standard, molecular ion m/z 151.0421, fragments [C2H7O2S]+ m/z 79.0210 and [C3H5O2]+ m/z 73.0283. b, Isotopic pattern of the molecular ion m/z 151.0421 with the calculated formula C5H11O3S and isotopic fine structure of [M + 1] and [M + 2]. c, DMSOP from a P. parvum extract with added 13C2-DMSOP. d, 13C2-DMSOP, molecular ion m/z 153.0485, fragments [13C2H7O2S]+ m/z 81.0277 and [C3H5O2]+ m/z 73.0282.
Extended Data Fig. 2 Structure of DMSOP.
Arrows show the heteronuclear multiple-bond coherence correlations. Numbers indicate carbon atom positions.
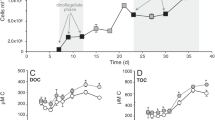
Extended Data Fig. 3 I. galbana growth and cellular DMSOP.
a, b, Growth (a) and photosynthetic efficiency (b) of I. galbana cultures. c, Cellular DMSP and DMSOP content. Data are mean ± s.d. of n = 3 independent cultures. P values are from a one-way repeated-measures ANOVA with Tukey post hoc test. A significant difference in cellular DMSOP concentration compared to day 3 was detected from day 7 onward.
Extended Data Fig. 4 DMSOP stablity in seawater and base.
a, DMSOP is stable over a period of 72 days in seawater (left). It degraded at room temperature under basic pH (pH = 11, monitored over 23 h; right). b, DMSO was released during this base treatment (integration of m/z = 79 in GC/MS). Data are mean ± s.d. of n = 3 independent samples. P values are from one-way repeated-measures ANOVA with Tukey post hoc test compared to t = 0 h.
Extended Data Fig. 5 DMSO release from DMSOP by bacteria.
a, b, DMSOP (1 µM) is degraded by A. faecalis, a dddY knockout mutant of A. faecalis (a) and by Halomonas sp. (b). Data are mean ± s.d. of n = 4 independent cultures. P values result from unpaired two-tailed Student’s t-tests. c, In separate experiments, it was demonstrated that DMSOP is the exclusive source for DMSO production in A. faecalis. Release of labelled DMSO from 13C2-DMSOP was monitored by GC/HRMS. The mass spectrum shows an average over the DMSO peak extracted from an A. faecalis culture that was incubated for 23 h with DMSOP. Integration of the ion traces 80.0200 (13C2-DMSO) and 78.0134 (DMSO) in three independent replicates revealed a degree of labelling of 99.3 ± 0.25%.
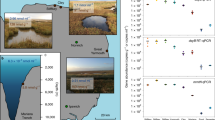
Extended Data Fig. 6 Map of sampling sites.
Sampling sites are indicated in red.
Supplementary information
Rights and permissions
About this article
Cite this article
Thume, K., Gebser, B., Chen, L. et al. The metabolite dimethylsulfoxonium propionate extends the marine organosulfur cycle. Nature 563, 412–415 (2018). https://doi.org/10.1038/s41586-018-0675-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0675-0
Keywords
This article is cited by
-
Decline of a distinct coral reef holobiont community under ocean acidification
Microbiome (2024)
-
Metabolic relationships between marine red algae and algae-associated bacteria
Marine Life Science & Technology (2024)
-
Microbial drivers of DMSO reduction and DMS-dependent methanogenesis in saltmarsh sediments
The ISME Journal (2023)
-
The biogeochemistry of marine dimethylsulfide
Nature Reviews Earth & Environment (2023)
-
Phylogeny and biogeography of the algal DMS-releasing enzyme in the global ocean
ISME Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.