Abstract
Nicotinamide adenine dinucleotide (NAD+) is a co-substrate for several enzymes, including the sirtuin family of NAD+-dependent protein deacylases. Beneficial effects of increased NAD+ levels and sirtuin activation on mitochondrial homeostasis, organismal metabolism and lifespan have been established across species. Here we show that α-amino-β-carboxymuconate-ε-semialdehyde decarboxylase (ACMSD), the enzyme that limits spontaneous cyclization of α-amino-β-carboxymuconate-ε-semialdehyde in the de novo NAD+ synthesis pathway, controls cellular NAD+ levels via an evolutionarily conserved mechanism in Caenorhabditis elegans and mouse. Genetic and pharmacological inhibition of ACMSD boosts de novo NAD+ synthesis and sirtuin 1 activity, ultimately enhancing mitochondrial function. We also characterize two potent and selective inhibitors of ACMSD. Because expression of ACMSD is largely restricted to kidney and liver, these inhibitors may have therapeutic potential for protection of these tissues from injury. In summary, we identify ACMSD as a key modulator of cellular NAD+ levels, sirtuin activity and mitochondrial homeostasis in kidney and liver.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The authors declare that all the data supporting the findings of this study are available from the corresponding author upon request.
References
Houtkooper, R. H., Pirinen, E. & Auwerx, J. Sirtuins as regulators of metabolism and healthspan. Nat. Rev. Mol. Cell Biol. 13, 225–238 (2012).
Imai, S. & Guarente, L. It takes two to tango: NAD+ and sirtuins in aging/longevity control. NPJ Aging Mech. Dis. 2, https://doi.org/10.1038/npjamd.2016.17 (2016).
Belenky, P., Bogan, K. L. & Brenner, C. NAD+ metabolism in health and disease. Trends Biochem. Sci. 32, 12–19 (2007).
Yang, Y. & Sauve, A. A. NAD+ metabolism: bioenergetics, signaling and manipulation for therapy. Biochim. Biophys. Acta 1864, 1787–1800 (2016).
Katsyuba, E. & Auwerx, J. Modulating NAD+ metabolism, from bench to bedside. EMBO J. 36, 2670–2683 (2017).
Bender, D. A. Biochemistry of tryptophan in health and disease. Mol. Aspects Med. 6, 101–197 (1983).
Fukuoka, S. I. Identification and expression of a cDNA encoding human α-amino-β-carboxymuconate-ε-semialdehyde decarboxylase (ACMSD). A key enzyme for the tryptophan-niacine pathway and “quinolinate hypothesis”. J. Biol. Chem. 277, 35162–35167 (2002).
Vrablik, T. L., Huang, L., Lange, S. E. & Hanna-Rose, W. Nicotinamidase modulation of NAD+ biosynthesis and nicotinamide levels separately affect reproductive development and cell survival in C. elegans. Development 136, 3637–3646 (2009).
Rongvaux, A., Andris, F., Van Gool, F. & Leo, O. Reconstructing eukaryotic NAD metabolism. BioEssays 25, 683–690 (2003).
McReynolds, M. R., Wang, W., Holleran, L. M. & Hanna-Rose, W. Uridine monophosphate synthetase enables eukaryotic de novo NAD+ biosynthesis from quinolinic acid. J. Biol. Chem. (2017).
Mouchiroud, L. et al. The NAD+/sirtuin pathway modulates longevity through activation of mitochondrial UPR and FOXO signaling. Cell 154, 430–441 (2013).
Hashimoto, T., Horikawa, M., Nomura, T. & Sakamoto, K. Nicotinamide adenine dinucleotide extends the lifespan of Caenorhabditis elegans mediated by sir-2.1 and daf-16. Biogerontology 11, 31–43 (2010).
Gebauer, J. et al. A genome-scale database and reconstruction of Caenorhabditis elegans metabolism. Cell Syst. 2, 312–322 (2016).
Burnett, C. et al. Absence of effects of Sir2 overexpression on lifespan in C. elegans and Drosophila. Nature 477, 482–485 (2011).
Viswanathan, M. & Guarente, L. Regulation of Caenorhabditis elegans lifespan by sir-2.1 transgenes. Nature 477, E1–E2 (2011).
Houtkooper, R. H. et al. Mitonuclear protein imbalance as a conserved longevity mechanism. Nature 497, 451–457 (2013).
Benedetti, C., Haynes, C. M., Yang, Y., Harding, H. P. & Ron, D. Ubiquitin-like protein 5 positively regulates chaperone gene expression in the mitochondrial unfolded protein response. Genetics 174, 229–239 (2006).
Haynes, C. M., Yang, Y., Blais, S. P., Neubert, T. A. & Ron, D. The matrix peptide exporter HAF-1 signals a mitochondrial UPR by activating the transcription factor ZC376.7 in C. elegans. Mol. Cell 37, 529–540 (2010).
Berdichevsky, A., Viswanathan, M., Horvitz, H. R. & Guarente, L. C. elegans SIR-2.1 interacts with 14-3-3 proteins to activate DAF-16 and extend life span. Cell 125, 1165–1177 (2006).
Honda, Y. & Honda, S. The daf-2 gene network for longevity regulates oxidative stress resistance and Mn-superoxide dismutase gene expression in Caenorhabditis elegans. FASEB J. 13, 1385–1393 (1999).
Pucci, L., Perozzi, S., Cimadamore, F., Orsomando, G. & Raffaelli, N. Tissue expression and biochemical characterization of human 2-amino 3-carboxymuconate 6-semialdehyde decarboxylase, a key enzyme in tryptophan catabolism. FEBS J. 274, 827–840 (2007).
Uhlén, M. et al. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Fukuwatari, T. Phthalate esters enhance quinolinate production by inhibiting α-amino-β-carboxymuconate-ε-semialdehyde decarboxylase (ACMSD), a key enzyme of the tryptophan pathway. Toxicol. Sci. 81, 302–308 (2004).
Saito, K. et al. Mechanism of increases in l-kynurenine and quinolinic acid in renal insufficiency. Am. J. Physiol. Renal Physiol. 279, F565–F572 (2000).
Pellicciari, R. et al. α-amino-β-carboxymuconate-ε-semialdehyde decarboxylase (ACMSD) inhibitors as novel modulators of de novo nicotinamide adenine dinucleotide (NAD+) biosynthesis. J. Med. Chem. 61, 745–759 (2018).
Chen, Y. & Guillemin, G. J. Kynurenine pathway metabolites in humans: disease and healthy states. Int. J. Tryptophan Res. 2, 1–19 (2009).
Michelotti, G. A., Machado, M. V. & Diehl, A. M. NAFLD, NASH and liver cancer. Nat. Rev. Gastroenterol. Hepatol. 10, 656–665 (2013).
Gariani, K. et al. Eliciting the mitochondrial unfolded protein response by nicotinamide adenine dinucleotide repletion reverses fatty liver disease in mice. Hepatology 63, 1190–1204 (2016).
Gariani, K. et al. Inhibiting poly-ADP ribosylation increases fatty acid oxidation and protects against fatty liver disease. J. Hepatol. (2016).
Lewington, A. J., Cerda, J. & Mehta, R. L. Raising awareness of acute kidney injury: a global perspective of a silent killer. Kidney Int. 84, 457–467 (2013).
Tran, M. T. et al. PGC1α drives NAD biosynthesis linking oxidative metabolism to renal protection. Nature 531, 528–532 (2016).
Liu, L. et al. Quantitative analysis of NAD synthesis-breakdown fluxes. Cell Metab. 27, 1067–1080 (2018).
Shi, H. et al. NAD deficiency, congenital malformations, and niacin supplementation. N. Engl. J. Med. 377, 544–552 (2017).
Kamath, R. S., Martinez-Campos, M., Zipperlen, P., Fraser, A. G. & Ahringer, J. Effectiveness of specific RNA-mediated interference through ingested double-stranded RNA in Caenorhabditis elegans. Genome Biol. 12, RESEARCH0002 (2001).
Mouchiroud, L. et al. Pyruvate imbalance mediates metabolic reprogramming and mimics lifespan extension by dietary restriction in Caenorhabditis elegans. Aging Cell 10, 39–54 (2011).
Mouchiroud, L. et al. The Movement Tracker: a flexible system for automated movement analysis in invertebrate model organisms. Curr. Protoc. Neurosci. 77, 8.37.1–8.37.21 (2016).
Zamporlini, F. et al. Novel assay for simultaneous measurement of pyridine mononucleotides synthesizing activities allows dissection of the NAD+ biosynthetic machinery in mammalian cells. FEBS J. 281, 5104–5119 (2014).
Yang, T. & Sauve, A. A. NAD metabolism and sirtuins: metabolic regulation of protein deacetylation in stress and toxicity. AAPS J. 8, E632–E643 (2006).
Oosterveer, M. H. et al. LRH-1-dependent glucose sensing determines intermediary metabolism in liver. J. Clin. Invest. 122, 2817–2826 (2012).
Jha, P., Wang, X. & Auwerx, J. Analysis of mitochondrial respiratory chain supercomplexes using blue native polyacrylamide gel electrophoresis (BN-PAGE). Curr. Protoc. Mouse Biol. 6, 1–14 (2016).
Lagouge, M. et al. Resveratrol improves mitochondrial function and protects against metabolic disease by activating SIRT1 and PGC-1α. Cell 127, 1109–1122 (2006).
Jha, P. et al. Role of adipose tissue in methionine-choline-deficient model of non-alcoholic steatohepatitis (NASH). Biochim. Biophys. Acta 1842, 959–970 (2014).
Schock-Kusch, D. et al. Transcutaneous measurement of glomerular filtration rate using FITC-sinistrin in rats. Nephrol. Dial. Transplant. 24, 2997–3001 (2009).
Schreiber, A. et al. Transcutaneous measurement of renal function in conscious mice. Am. J. Physiol. Renal Physiol. 303, F783–F788 (2012).
Melnikov, V. Y. et al. Neutrophil-independent mechanisms of caspase-1- and IL-18-mediated ischemic acute tubular necrosis in mice. J. Clin. Invest. 110, 1083–1091 (2002).
Acknowledgements
We thank P. Gönczy and the Caenorhabditis Genetics Center for providing reagents, the Bioimaging and Optics Core Facility and the Phenotyping Unit of EPFL, N. Moullan, T. Clerc and S. Bichet for technical assistance. E.K. was supported by Fondation Romande pour la Recherche sur le Diabète. M.Z. was supported by the KNOW consortium ‘Healthy Animal—Safe Food’ MS&HE no. 05-1/KNOW2/2015 and the Foundation for Polish Science. This work was supported by funds from EPFL and Swiss National Science Foundation (grant 310030B_160318). J.I. acknowledges funding from the Foundation Pierre-Mercier pour la Science. We thank H. Gallant-Ayala for advice on analytical methods.
Reviewer information
Nature thanks W. Hanna-Rose, S. Parikh, J. Tam and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
E.K., J.A. and R.P. conceived and designed the project. E.K., A.M., M.Z., F.D.F., K.G., D.R. and N.G. performed experiments. P.L. synthesized ACMSD inhibitors. O.M. made the acsd-1::GFP C. elegans reporter strain. N.R. and L.C. performed activity assays for the enzymes in de novo NAD+ biosynthesis. V.v.d.V. and J.I. performed the metabolomics and N.S.-R. carried out histopathology. S.d.S. and D.L. assisted with GFR measurements. E.K., K.S. and J.A. wrote the manuscript with the contributions from all other authors.
Corresponding authors
Ethics declarations
Competing interests
J.A., R.P. and N.R. are inventors on US patent 9,708,272 (18 July, 2017), filed by TES Pharma S.r.l., Corciano, Italy. The patent covers the results obtained with the ACMSD inhibitors, TES-991 and TES-1025, described in Figs. 3–5. R.P., F.D.F., N.G. and P.L. are employed by TES Pharma.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 acsd-1 LOF improves NAD+ levels, mitochondrial function, and lifespan through de novo synthesis in C. elegans.
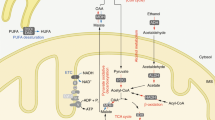
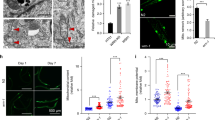
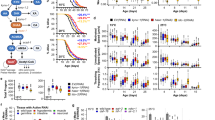
a, De novo synthesis of NAD+ from tryptophan. Names of the worm’s orthologues are in blue. b, acsd-1 expression pattern across different developmental stages in wild-type worms expressing extrachromosomal array of acsd-1::GFP transgene. Scale bar, 100 μm. c, acsd-1 expression pattern in adult wild-type worms expressing extrachromosomal array of acsd-1::GFP transgene. i, intestine; m, muscle; p, pharynx; v, vulva. d, acsd-1 mRNA levels in wild-type and rrf-3(pk1426) mutants (n = 6, each n represents a pool of ~600 worms). e, ACSD-1 activity in control (empty vector) versus acsd-1 RNAi-fed worms quantified in both wild-type and rrf-3 mutants (n = 3, where each n represents a pool of ~3,600 worms) with compensation for negative controls. f, QPRT-like activity can be detected in both wild-type worms and rrf-3 mutants (n = 3, each n represents a pool of ~3,600 worms). g, h, Effects of acsd-1 knockdown throughout the entire life on N2 (g) and rrf-3 mutant (h) worm lifespan. i, Lifespan of rrf-3(pk1426) mutants exposed to control or acsd-1 RNAi upon tryptophan supplementation. P*, ctrl versus ctrl + Trp 50 μM; P^, ctrl versus acsd-1 RNAi; P°, ctrl + Trp 50 μM versus acsd-1 RNAi + Trp 50 μM. j, Quantification of GFP signal in ges-1::mito::GFP reporter strain, expressing mitochondria-targeted GFP in the intestine at day 1 and 3 of adulthood (n = 4, each n represents a pool of 20 worms). k, Blue native PAGE on mitochondria extracted from rrf-3 mutant worms fed with either empty vector or acsd-1 RNAi bacteria at day 2 of adulthood (n = 3, each n represents mitochondria extracted from a pool of ~10,000 worms). l, Mitochondrial morphology in the Pmyo-3::mito::GFP reporter strain fed with control or acsd-1 RNAi. Stars represent nuclei. Scoring includes the total perimeter of the mitochondrial network, its total area, the area occupied by the mitochondria within the cell and the circularity assessment, in which 1 is a perfect circle and 0 is a line (n = 6 worms). m, Epistasis between RNAi for acsd-1 and the UPRmt regulator, ubl-5. P*, ctrl versus ctrl/acsd-1 RNAi; P°, ctrl/ubl-5 RNAi versus ubl-5/acsd-1 RNAi. n, Quantification of the GFP signal in hsp-4::GFP reporter strain (n = 4, each n represents a pool of 20 worms) at day 1 and 3 of adulthood. o, Quantification of the GFP signal in hsp-16.2::GFP reporter strain (n = 4, each n represents a pool of 20 worms). After the first time point sampled at 20 °C, worms were exposed to 37 °C, and the measurement was repeated every hour for 6 h. p, Expression of UPRmt genes in worms at day 2 of adulthood fed with control or acsd-1 RNAi (n = 6, each n represents a pool of ~600 worms). q, Expression of sod-3 mRNA at day 1 of adulthood in control or acsd-1 RNAi-fed worms (n = 3, each n represents a pool of ~600 worms). r, Survival of wild-type (N2) worms exposed to 4 mM paraquat starting at the L4 stage, in which the knockdown of acsd-1 was performed at different life stages. P*, ctrl versus acsd-1 RNAi whole life; P^, ctrl versus acsd-1 RNAi development; P°, ctrl versus acsd-1 RNAi adulthood. s, Epistasis between RNAi for acsd-1 and daf-16 in wild-type (N2) worms exposed to 4 mM paraquat. P*, ctrl versus ctrl/acsd-1 RNAi; P^, ctrl/daf-16 RNAi versus daf-16/acsd-1 RNAi. All worm assays, except for hsp-16.2::GFP reporter strain, were performed at 20 °C and repeated at least once. Data are mean ± s.e.m. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001. P values calculated using two-tailed t-test (d, e, j, l, n–q) or log-rank test (g–i, m, r, s). For individual P values, see Source Data. For lifespan values, see Extended Data Table 1.
Extended Data Fig. 2 Pathways activated by Acmsd knockdown in worms are conserved in mammalian cells.
a, Acmsd transcript levels reflected by the Ct values in different hepatic and renal cells and cell lines (n = 4). Ct values larger than 35 reflect very low transcript levels. b, Efficiency of Acmsd shRNA in mouse primary hepatocytes 48 h post adenoviral transduction (n = 6). c, NAD+ levels in mitochondria of AML-12 cells transduced with either shRNA control or shRNA against Acmsd (n = 5). d, e, Blue native PAGE followed by in-gel activity assay for complex II (blue) (d), and complex I (purple) and IV (brown) (e) on mitochondria extracted from mouse primary hepatocytes transduced with either shRNA control or shRNA against Acmsd for 48 h. The experiment was performed once. f, Primary hepatocytes extracted from a Sirt1L2/L2 mouse were transduced either with an adenovirus encoding GFP (wild-type condition) or the Cre recombinase to generate Sirt1 knockout primary hepatocytes. These hepatocytes were exposed to an shRNA targeting a random sequence or shRNA targeting Acmsd. Transcript levels of Acmsd and Sirt1 (n = 3). g, FOXO1 acetylation levels in mouse primary hepatocytes transduced with either shRNA control or shRNA against Acmsd for 48 h. The experiment was independently performed twice. Data are mean ± s.e.m.; each n represents a biologically independent sample. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001. P values calculated using two-tailed t-test. For gel source images see Supplementary Fig. 1. For individual P values, see Source Data.
Extended Data Fig. 3 Pharmacological inhibition of ACMSD has similar effects to genetic downregulation.
a, b, mRNA levels of mitochondrial genes in mouse primary hepatocytes treated for 24 h with DMSO or TES-1025 (a) or TES-991 (b), at the indicated concentrations (n = 3). c, SOD2 activity in mouse primary hepatocytes treated for 24 h with DMSO or TES-1025, at indicated concentrations (n = 4). d, Fatty acid oxidation assessed in mouse primary hepatocytes treated with DMSO or TES-991 for 24 h at the indicated concentrations (n = 5). FCCP (2 μM) was used as an uncoupler to reach maximal respiration. e, mRNA levels of mitochondrial genes in HK-2 cells after 24 h of treatment with TES-1025 or TES-1025 in combination with SIRT1 inhibitor, EX527, at the indicated concentrations (n = 5–8). f, Apoptosis rate in HK-2 cells assessed 16 h after addition of 50 μM cisplatin by caspase-3/7 activity. TES-1025 was added simultaneously with the cisplatin. g, Biochemical analysis of plasma from mice fed with chow diet or chow diet supplemented with TES-991 or TES-1025 at 15 mg kg−1 body weight day−1 (ctrl and TES-991, n = 10; TES-1025, n = 9 mice). h, i, Quinolinic acid (QA) (h) and nicotinic acid (NA) (i) levels in livers (n = 9), kidneys (ctrl, n = 10; TES-991, TES-1025, n = 9 mice) and brains (ctrl, n = 11; TES-991, TES-1025, n = 12 mice), from mice fed with control chow diet or chow diet supplemented with TES-991 or TES-1025 at the dose of 15 mg kg−1 body weight day−1. j, k, mRNA levels of β-oxidation, mitochondrial and oxidative stress defence genes in livers (j) and kidneys (k) of mice described in g (n = 6 mice). Data are mean ± s.e.m. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001. P values calculated using two-tailed t-test (a–d, g–k) or one-way ANOVA (e, f). For individual P values, see Source Data.
Extended Data Fig. 4 ACMSD inhibitors protect hepatic function from NAFLD induced by MCD diet.
a, Plasma aspartate transaminase (AST) levels in 16-week-old C57BL/6J male mice fed for 2.5 weeks with control diet, MCD diet or MCD diet supplemented with 15mg kg−1 day−1 TES-991 (n = 8 mice). b, Representative photomicrographs of liver tissues stained with H&E or Oil red O from the mouse cohorts described in a. The experiment was performed twice independently. c, Representative photomicrographs of liver tissues from the mouse cohorts described in a, stained with CD45 and the corresponding negative control. The experiment was performed twice independently. d, Hepatic SOD2 activity in mouse cohorts described in a (chow diet, n = 8; MCD diet, n = 7; MCD diet + TES-991, n = 6 mice). e–h, Liver NAD+ (e), triglyceride content (f), plasma ALT (g) and AST (h) levels in congenic C57BL/6J Sirt1hep−/− mice that match the mouse cohorts described in a regarding age, gender and treatment duration (chow diet, n = 8; MCD diet, MCD diet + TES-991, n = 10 mice). i, Representative photomicrographs of liver tissues stained with H&E from the Sirt1hep−/− mice described in e–h. The experiment was performed once. j, Hepatic SOD2 activity in congenic C57BL/6J Sirt1hep−/− mice described in e–h (chow diet, n = 8; MCD diet, MCD diet + TES-991, n = 9 mice). k, mRNA levels of oxidative stress defence, mitochondrial, β-oxidation, inflammatory and fibrosis genes in livers of Sirt1hep−/− mice (n = 8 mice). Data are mean ± s.e.m. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001. P values calculated using two-tailed t-test. For individual P values, see Source Data.
Extended Data Fig. 5 ACMSD inhibitors protect renal function in two different models of AKI.
a, Schematic timeline of the cisplatin-induced AKI study. AKI was induced at day 10 after the beginning of the study in male C57BL/6J mice by a single intraperitoneal dose of cisplatin (20 mg kg−1 body weight). Mice in the sham control group were injected with a saline solution. GFR was measured non-invasively 52 h post-cisplatin administration. b–d, Representative photomicrographs of H&E-stained kidney sections (b), histopathological scoring for tubular necrosis (c) and inflammatory cell infiltration (d) of mouse cohorts described in a (sham control, cisplatin-AKI + TES-1025, n = 6; cisplatin-AKI, n = 5 mice). e, Schematic timeline of the IR-AKI study. AKI was induced at day 10 after the beginning of the study in anaesthetized male C57BL/6J mice by a dorsal surgical incision and bilateral occlusion of the renal pedicles for 25 min. Mice in the sham control group underwent the same surgical procedure without application of the occluding clamp on the renal pedicles. f–j, Representative photomicrographs of H&E-stained kidney sections (f) and histopathological scoring for cumulative score (g), tubular necrosis (h), tubular dilation (i), and cast formation (j) of mouse cohorts described in e (n = 5 mice). Tubular cell necrosis (arrows), tubular dilation and casts (asterisk) and interstitial oedema (crescent moon) are indicated on the pictures. k–m, Glutathione protein levels (k), MPO activity (l) and NAD+ content (m) in kidneys of the IR-AKI cohorts described in e (n = 5 mice). n, Protein expression of the respiratory complex subunits in kidneys from the mouse cohorts described in e. The experiment was performed independently twice. Data are mean ± s.e.m. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001. P values calculated using two-tailed t-test. For gel source images see Supplementary Fig. 1. For individual P values, see Source Data. The histopathological scoring was performed independently by two pathologists in a blinded manner (b–d, f–j).
Supplementary information
Supplementary Figure
This file contains the uncropped blots for Figs 1i, 2e, 3l, 5h and Extended Data Figs 2g and 5m.
Rights and permissions
About this article
Cite this article
Katsyuba, E., Mottis, A., Zietak, M. et al. De novo NAD+ synthesis enhances mitochondrial function and improves health. Nature 563, 354–359 (2018). https://doi.org/10.1038/s41586-018-0645-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0645-6
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.