Abstract
Immune checkpoint blockade therapy has been successful in treating some types of cancer but has not shown clinical benefits for treating leukaemia1. This result suggests that leukaemia uses unique mechanisms to evade this therapy. Certain immune inhibitory receptors that are expressed by normal immune cells are also present on leukaemia cells. Whether these receptors can initiate immune-related primary signalling in tumour cells remains unknown. Here we use mouse models and human cells to show that LILRB4, an immunoreceptor tyrosine-based inhibition motif-containing receptor and a marker of monocytic leukaemia, supports tumour cell infiltration into tissues and suppresses T cell activity via a signalling pathway that involves APOE, LILRB4, SHP-2, uPAR and ARG1 in acute myeloid leukaemia (AML) cells. Deletion of LILRB4 or the use of antibodies to block LILRB4 signalling impeded AML development. Thus, LILRB4 orchestrates tumour invasion pathways in monocytic leukaemia cells by creating an immunosuppressive microenvironment. LILRB4 represents a compelling target for the treatment of monocytic AML.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The TCGA datasets analysed are available in the UCSC Xena Browser (https://xena.ucsc.edu). The RNA-seq datasets generated in the current study have been deposited in NCBI SRA database with the SRA accession number SRP155049.
References
Curran, E. K., Godfrey, J. & Kline, J. Mechanisms of immune tolerance in leukemia and lymphoma. Trends Immunol. 38, 513–525 (2017).
Kang, X. et al. Inhibitory leukocyte immunoglobulin-like receptors: Immune checkpoint proteins and tumor sustaining factors. Cell Cycle 15, 25–40 (2016).
Hirayasu, K. & Arase, H. Functional and genetic diversity of leukocyte immunoglobulin-like receptor and implication for disease associations. J. Hum. Genet. 60, 703–708 (2015).
Trowsdale, J., Jones, D. C., Barrow, A. D. & Traherne, J. A. Surveillance of cell and tissue perturbation by receptors in the LRC. Immunol. Rev. 267, 117–136 (2015).
Dobrowolska, H. et al. Expression of immune inhibitory receptor ILT3 in acute myeloid leukemia with monocytic differentiation. Cytometry B Clin. Cytom. 84, 21–29 (2013).
Vlad, G. et al. Membrane and soluble ILT3 are critical to the generation of T suppressor cells and induction of immunological tolerance. Int. Rev. Immunol. 29, 119–132 (2010).
Mosier, D. E., Gulizia, R. J., Baird, S. M. & Wilson, D. B. Transfer of a functional human immune system to mice with severe combined immunodeficiency. Nature 335, 256–259 (1988).
Ha, S. et al. Isolation and characterization of IgG1 with asymmetrical Fc glycosylation. Glycobiology 21, 1087–1096 (2011).
Straus, D. J. et al. The acute monocytic leukemias: multidisciplinary studies in 45 patients. Medicine (Baltimore) 59, 409–425 (1980).
Colovai, A. I. et al. Expression of inhibitory receptor ILT3 on neoplastic B cells is associated with lymphoid tissue involvement in chronic lymphocytic leukemia. Cytometry B Clin. Cytom. 72, 354–362 (2007).
Crispe, I. N. et al. Cellular and molecular mechanisms of liver tolerance. Immunol. Rev. 213, 101–118 (2006).
Carson, M. J., Doose, J. M., Melchior, B., Schmid, C. D. & Ploix, C. C. CNS immune privilege: hiding in plain sight. Immunol. Rev. 213, 48–65 (2006).
Fujisaki, J. et al. In vivo imaging of Treg cells providing immune privilege to the haematopoietic stem-cell niche. Nature 474, 216–219 (2011).
Castells, M. C. et al. gp49B1-αvβ3 interaction inhibits antigen-induced mast cell activation. Nat. Immunol. 2, 436–442 (2001).
Grainger, D. J., Reckless, J. & McKilligin, E. Apolipoprotein E modulates clearance of apoptotic bodies in vitro and in vivo, resulting in a systemic proinflammatory state in apolipoprotein E-deficient mice. J. Immunol. 173, 6366–6375 (2004).
Ali, K., Middleton, M., Puré, E. & Rader, D. J. Apolipoprotein E suppresses the type I inflammatory response in vivo. Circ. Res. 97, 922–927 (2005).
DiDonato, J. A., Mercurio, F. & Karin, M. NF-κB and the link between inflammation and cancer. Immunol. Rev. 246, 379–400 (2012).
You, M., Flick, L. M., Yu, D. & Feng, G. S. Modulation of the nuclear factor κb pathway by Shp-2 tyrosine phosphatase in mediating the induction of interleukin (IL)-6 by IL-1 or tumor necrosis factor. J. Exp. Med. 193, 101–110 (2001).
Baumgartner, B. et al. Increased IκB kinase activity is associated with activated NF-κB in acute myeloid blasts. Leukemia 16, 2062–2071 (2002).
Béné, M. C. et al. CD87 (urokinase-type plasminogen activator receptor), function and pathology in hematological disorders: a review. Leukemia 18, 394–400 (2004).
Su, S. C., Lin, C. W., Yang, W. E., Fan, W. L. & Yang, S. F. The urokinase-type plasminogen activator (uPA) system as a biomarker and therapeutic target in human malignancies. Expert Opin. Ther. Targets 20, 551–566 (2016).
Wang, Y. et al. Identification of a novel nuclear factor-kappaB sequence involved in expression of urokinase-type plasminogen activator receptor. Eur. J. Biochem. 267, 3248–3254 (2000).
Hu, J. et al. uPAR induces expression of transforming growth factor β and interleukin-4 in cancer cells to promote tumor-permissive conditioning of macrophages. Am. J. Pathol. 184, 3384–3393 (2014).
Ilkovitch, D. & Lopez, D. M. Urokinase-mediated recruitment of myeloid-derived suppressor cells and their suppressive mechanisms are blocked by MUC1/sec. Blood 113, 4729–4739 (2009).
Baitsch, D. et al. Apolipoprotein E induces antiinflammatory phenotype in macrophages. Arterioscler. Thromb. Vasc. Biol. 31, 1160–1168 (2011).
Hagemann, T. et al. “Re-educating” tumor-associated macrophages by targeting NF-κB. J. Exp. Med. 205, 1261–1268 (2008).
Mussai, F. et al. Acute myeloid leukemia creates an arginase-dependent immunosuppressive microenvironment. Blood 122, 749–758 (2013).
de Goeje, P. L. et al. Immunoglobulin-like transcript 3 is expressed by myeloid-derived suppressor cells and correlates with survival in patients with non-small cell lung cancer. OncoImmunology 4, e1014242 (2015).
Chang, C. C. et al. Tolerization of dendritic cells by T(S) cells: the crucial role of inhibitory receptors ILT3 and ILT4. Nat. Immunol. 3, 237–243 (2002).
Suciu-Foca, N. et al. Soluble Ig-like transcript 3 inhibits tumor allograft rejection in humanized SCID mice and T cell responses in cancer patients. J. Immunol. 178, 7432–7441 (2007).
Piedrahita, J. A., Zhang, S. H., Hagaman, J. R., Oliver, P. M. & Maeda, N. Generation of mice carrying a mutant apolipoprotein E gene inactivated by gene targeting in embryonic stem cells. Proc. Natl Acad. Sci. USA 89, 4471–4475 (1992).
Zheng, J. et al. Inhibitory receptors bind ANGPTLs and support blood stem cells and leukaemia development. Nature 485, 656–660 (2012).
Kang, X. et al. The ITIM-containing receptor LAIR1 is essential for acute myeloid leukaemia development. Nat. Cell Biol. 17, 665–677 (2015).
Deng, M. et al. A motif in LILRB2 critical for Angptl2 binding and activation. Blood 124, 924–935 (2014).
Zheng, J. et al. Ex vivo expanded hematopoietic stem cells overcome the MHC barrier in allogeneic transplantation. Cell Stem Cell 9, 119–130 (2011).
Lu, Z. et al. Fasting selectively blocks development of acute lymphoblastic leukemia via leptin-receptor upregulation. Nat. Med. 23, 79–90 (2017).
Zhang, C. C., Kaba, M., Iizuka, S., Huynh, H. & Lodish, H. F. Angiopoietin-like 5 and IGFBP2 stimulate ex vivo expansion of human cord blood hematopoietic stem cells as assayed by NOD/SCID transplantation. Blood 111, 3415–3423 (2008).
Zheng, J., Huynh, H., Umikawa, M., Silvany, R. & Zhang, C. C. Angiopoietin-like protein 3 supports the activity of hematopoietic stem cells in the bone marrow niche. Blood 117, 470–479 (2011).
Cawthorne, C., Swindell, R., Stratford, I. J., Dive, C. & Welman, A. Comparison of doxycycline delivery methods for Tet-inducible gene expression in a subcutaneous xenograft model. J. Biomol. Tech. 18, 120–123 (2007).
Denisov, I. G., Grinkova, Y. V., Lazarides, A. A. & Sligar, S. G. Directed self-assembly of monodisperse phospholipid bilayer Nanodiscs with controlled size. J. Am. Chem. Soc. 126, 3477–3487 (2004).
Acknowledgements
We thank the National Cancer Institute (1R01CA172268 and 5P30CA142543), the Leukemia & Lymphoma Society (1024-14 and TRP-6024-14), the March of Dimes Foundation (1-FY14-201), the Cancer Prevention and Research Institute of Texas (RP140402, DP150056, RP180435, PR150551, and RR150072), the Robert A. Welch Foundation (I-1834 and AU-0042-20030616), the National Natural Science Foundation of China (81570093, 81422001, and 81721004), the National Basic Research Program of China (2014CB965000), and the China Scholarship Council (201608330307) for support. We also thank G. Salazar for editing the manuscript and Y. Dang for RStudio coding.
Author information
Authors and Affiliations
Contributions
M.D. and C.C.Z. designed the study and wrote the manuscript. M.D., C.C.Z., X.G., N.Z., Z.A. and X.C.L. contributed to the experimental plan and data interpretation. M.D., Z.Li. and L.H. performed mouse experiments. M.D., X.G., L.X., Z.Li., Y.C., Z.Lu., Y.X., Z.Z., C.Y., L.L., H.D., Z.A., J.Z. and N.Z. performed antibody characterizations. M.D., X.G. and L.L. measured APOE–LILRB4 binding affinity. K.H., H.A., M.D., J.K., L.H. and J.X. performed reporter assays. W.C., H.C., R.C., M.J.Y., J.H., N.U. and C.L. provided primary patient samples. M.D., W.C., L.H. and H.C. performed flow cytometry analysis of primary patient cells. M.D., Z.Li. and H.C. performed CRISPR–Cas9 experiments. M.D., X.G., Z.Li., L.H., H.C., W.L. and G.W. performed plasmid constructions. M.D., Z.Li., L.H., H.C., W.L., J.X., S.J., X.L. and L.Z. performed in vitro T cell assays. N.N. and Y.J. produced lipid-bound APOE protein. M.D., Z.Li., L.H. and F.H. performed western blotting. H.T., A.H.S., T.H., Y.Z., B.C., N.X., G.-Q.C., Y.-X.F., X.C.L., Z.A., N.Z. and C.C.Z. helped with or advised on experiments and provided reagents. M.D. and H.Z. performed statistical analysis.
Corresponding authors
Ethics declarations
Competing interests
The Board of Regents of the University of Texas System has filed patent applications with PCT Application Nos. PCT/US2016/020838, which covers anti-LILRB antibodies and their uses in detecting and treating cancer, and PCT/US2017/044171, which covers the methods for identifying LILRB-blocking antibodies. Authors C.C.Z., M.D., Z.A., N.Z., X.G. and J.Z. are listed as inventors of PCT/US2016/020838. Authors C.C.Z., Z.A., N.Z., M.D., J.K. and X.G. are listed as inventors of PCT/US2017/044171. Both patent applications have been exclusively licensed to Immune-Onc Therapeutics by the Board of Regents of the University of Texas System. Authors Z.A. and C.C.Z. are Scientific Advisory Board members with Immune-Onc Therapeutics, who also own equities and have a sponsored research agreement with Immune-Onc Therapeutics. Authors T.H. and X.C.L. are employees of and hold equities in Immune-Onc Therapeutics.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 LILRB4 expression in patients with AML negatively correlated with overall survival and T cell proliferation.
a, Analysis of correlation between mRNA levels of immune-modulating molecules and the overall survival of patients with AML (n = 160, divided into three groups based on gene expression) in TCGA database (https://xena.ucsc.edu) by Kaplan–Meier long-rank test. b, Kaplan–Meier analysis of correlations between LILRB4 mRNA level and the overall survival of patients (n = 160) from the TCGA database, performed in Xena browser (https://xena.ucsc.edu). Low, n = 57; medium, n = 48; high, n = 55. The P value was from Kaplan–Meier log-rank test. c, mRNA expression data from the TCGA database were analysed as a function of the AML subtype of the patient. M0, n = 16; M1, n = 42; M2, n = 39; M3, n = 16; M4, n = 35; M5, n = 18; M6, n = 2; M7, n = 3; and two not-classified AML samples. Pairwise comparisons between M4 and each one of the other subtypes (all P < 0.0001), as well as between M5 and each one of the other subtypes (all P < 0.0001), using two-sample t-test. Mean and s.e.m. values are shown. d, A multivariable Cox regression analysis to assess the association, with adjustment for confounders that include age, cytogenetics and PML-RAR mutation in TCGA database. The total sample size was 79. *P < 0.05 is considered significant. e, f, Autologous T cells isolated from individual patients with monocytic AML or B-ALL were incubated with irradiated LILRB4-positive or LILRB4-negative primary leukaemia cells from the same patients. pT, patient T cells. Allogeneic T cells isolated from healthy donors were incubated with irradiated LILRB4-positive or LILRB4-negative primary leukaemia cells from indicated patients with AML or B-ALL at an E:T of 10:1. nT, normal T cells. After culture with anti-CD3/CD28/CD137-coated beads and rhIL-2 for 14 days, T cells were stained with anti-CD3, anti-CD4, and anti-CD8 antibodies and analysed by flow cytometry. e, f, P values from two-tailed Student’s t-test. P values in black indicate significance of CD3+CD8+ cells; P values in red indicate significance of CD3+CD4+ cells. n = 2 or 3 biologically independent samples with mean and s.e.m. See raw data for e and f in Source Data.
Extended Data Fig. 2 LILRB4 suppresses T cell proliferation in vitro.
a, Schematic of preparation of LILRB4-modulated THP-1 cells and examination of LILRB4 expression on the cell surfaces by flow cytometry. WT, THP-1 cells treated with scrambled control; LILRB4KO, LILRB4-knockout THP-1 cells; LILRB4KO-wt, forced expression of wild-type LILRB4 in LILRB4KO THP-1 cells; LILRB4KO-intΔ, forced expression of intracellular domain-deleted mutant LILRB4 in LILRB4KO THP-1 cells. b, Loss of LILRB4 on THP-1 cells reduces T cell suppression. Representative photograph of Fig. 1c (scale bar, 100 µm). c, Loss of LILRB4 on THP-1 cells does not affect cell proliferation (n = 3 biologically independent samples with mean and s.e.m.). d, Examination of LILRB4 expression on cell surface of LILRB4KO MV4-11 cells by flow cytometry. e, f, Loss of LILRB4 on MV4-11 cells reduces T cell suppression. T cells isolated from healthy donors incubated in the lower chambers of a 96-well transwell plate with irradiated MV4-11 cells (E:T of 2:1) in the upper chamber separated by a membrane with 3-µm pores. After culture with anti-CD3/CD28-coated beads and rhIL-2 for 7 days, representative cells were photographed using an inverted microscope (scale bar, 100 µm) (e) and T cells were stained with anti-CD3 and analysed by flow cytometry (f). n = 4 biologically independent samples. g, Loss of LILRB4 on MV4-11 cells does not affect cell proliferation (n = 3 biologically independent samples with mean and s.e.m.). h, i, T cells (E, effector cells) isolated from healthy donors were incubated with indicated irradiated THP-1 cells (T, target cells) without direct contact in transwells for 2 days. E:T = 2:1. T cells were treated with BrdU for 30 min followed by BrdU and 7-AAD staining for flow cytometry analysis. Representative flow cytometry plots are shown in h and the cell cycle status is summarized in i. T control, T cells cultured without THP-1 cells. n = 3 biologically independent samples with mean and s.e.m. j, k, T cells (E, effector cells) isolated from healthy donors were stained with CFSE and incubated with indicated irradiated THP-1 cells (T, target cells) without direct contact in transwells for 2 days. A representative flow cytometry plot is shown in j and the percentages of proliferating T cells indicated by CFSE-low staining is shown in k. n = 3 biologically independent samples with mean and s.e.m. l, LILRB4 increases PD-1 expression on T cells in coculture of leukaemia cells and T cells. T cells (E, effector cells) isolated from healthy donors were incubated with indicated irradiated THP-1 cells (T, target cells) in a non-contact manner for 5 days. E:T = 2:1. T cells were stained with anti-LAG-3, anti-TIM-3, anti-TIGIT, anti-PD-1 and anti-FasL antibodies for flow cytometry analysis. Representative flow cytometry plots and the mean of fluorescence intensities, at the right-upper corner (black, WT; red, KO), are shown. Experiments were performed three times with similar results. m, n, Anti-LILRB4 antibody had no effect on proliferation of THP-1 cells (m) or T cells (n). m, The growth of THP-1 cells during 7 days treatment with IgG or anti-LILRB4 antibody (n = 3 biologically independent samples with mean and s.e.m.). n, The numbers of human primary T cells after 5 days treatment with IgG or anti-LILRB4 antibody in vitro (n = 3 biologically independent samples with mean and s.e.m.). o, p, Primary T cells and irradiated THP-1 cells (E:T ratio, 2:1) were placed in the lower and upper chamber, respectively, and treated with 10 µg ml−1 control IgG or anti-LILRB4 antibodies. o, Representative photographs of T cells (scale bar, 100 µm). p, T cells stained with anti-CD3 and analysed by flow cytometry. n = 4 biologically independent samples. q, Primary T cells stimulated with anti-CD3/CD28/CD137-coated beads were co-cultured with wild-type or LILRB4KO THP-1 cells with indicated E:T ratios for 4 h (n = 3 biologically independent samples with mean and s.e.m.). Cytotoxity of leukaemia cells was determined by PI staining in flow cytometry analysis. r–u, CD8+ T cells (5 × 104 cells) stimulated with anti-CD3/CD28/CD137-coated beads were co-cultured with 5 × 103 THP-1 cells that stably express GFP and treated with 100 µg ml−1 anti-LILRB4 antibodies or control IgG for 5 days. s, t, n = 4 biologically independent samples; u, n = 3 biologically independent samples with mean and s.e.m. Representative flow plots (r) of the percentages of T cells (GFP–) and surviving leukaemia cells (GFP+), and quantification of T cells (s), GFP+ leukaemia cells (t), and secretion of IFNγ (u), are shown. b, d, e, h, j, o, r, Experiments repeated independently three times with similar results. See Methods for definition of box plot elements in f, p, s, t. All P values were from two-tailed Student’s t-test.
Extended Data Fig. 3 Inhibition of LILRB4 reduces leukaemia development in humanized immunocompromised mice and syngeneic mice.
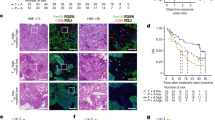
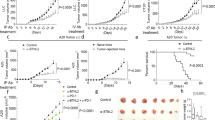
a–c, Wild-type or LILRB4KO THP-1 cells (3 × 106 cells per mouse) were subcutaneously implanted into hPBMC-repopulated NSG mice (WT, n = 14 mice with mean and s.e.m.; LILRB4KO, n = 10 mice, mean and s.e.m.; also see Source Data). Tumour size (a), quantification of CD3+ cells at day 31 in peripheral blood of recipient mice (b) and representative flow plots showing CD4+ and CD8+ T cells (c) are shown. d, e, LILRB4 increases PD-1 expression on tumour-infiltrated T cells. Wild-type or LILRB4KO THP-1 cells were subcutaneously implanted into hPBMC-repopulated NSG mice. Three weeks after implantation, 7 out of 10 WT group mice had large tumours and 3 out of 10 knockout-group mice had tiny tumours. These tumours were dissected for immunohistochemistry and flow cytometry staining with anti-LILRB4, anti-CD3, anti-PD-1 and anti-ARG1 antibodies. Left corner images are magnified from yellow highlighted regions. In CD3 and PD-1 staining images, orange dashed lines indicate the tumour boundary. Black arrowheads indicate PD-1 positive cells. Scale bar, 100 μm. e, Tumours were dissected and cells in tumour region were stained with anti-CD3 and anti-PD-1 antibodies for flow cytometry analysis. The percentages of PD-1+ T cells (ratio of PD-1+CD3+ cells to CD3+ cells) were calculated. f–i, THP-1 cells were transplanted into hPBMC-repopulated NSG mice, and mice were treated with control IgG or anti-LILRB4 antibodies after 6 days (10 mg kg−1; n = 5). Leukaemia development was monitored by luminescence imaging (f); luminescence flux (radiance) at day 26 (g; n = 5) and T cell numbers at day 26 in representative mice (h, i). j, k, Engraftment of human T cells and intravenously transplanted Dox-inducible LILRB4-knockout THP-1 cells (GFP+) in NSG mice at day 7 before Dox administration (n = 5). l, Representative flow plot shows that LILRB4 was successfully deleted in engrafted leukaemia cells in bone marrow of Dox-fed mouse at the endpoint. n.s., not significant. m–w, Mouse AML C1498 cells (3 × 106 cells per mouse) that stably express LILRB4–IRES–GFP were subcutaneously implanted into C57BL/6 mice. Anti-LILRB4-N297A antibodies or control IgG were intravenously injected at 6, 9, 12, 15, 18 and 21 days after implantation of tumour cells. Two groups of mice were treated with anti-CD8 antibodies at 3, 6, 9 and 12 days after implantation of tumour cells to achieve CD8+ T cell depletion. m, Tumour growth of subcutaneously implanted human LILRB4-expressing mouse AML C1498 cells (hLILRB4 C1498) in C57BL/6 mice with anti-LILRB4-N297A antibodies or control antibody treatment (n = 5 mice). Also see Source Data. n, Survival curve of subcutaneous hLILRB4 C1498 tumour-bearing mice (n = 12 mice). As for tumour size, anti-LILRB4 antibodies decreased the tumour weight (o, n = 5 mice) but did not do so in the absence of CD8+ T cells (p, n = 5 mice). The percentage of CD8+ T cells in spleen was significantly negatively correlated with tumour size (q, n = 5 mice) but not in the absence of CD8+ T cells (r, n = 5 mice). s, Adoptive transplantation of spleen cells from control mice or tumour-bearing mice that were cured by anti-LILRB4-N297A treatment (n = 5 mice). Tumour size was monitored as a function of time. Arrow indicates day of rechallenge in mice that had eliminated leukaemia with three times the number of AML cells (n = 4 mice). Also see Source Data. Anti-LILRB4 antibodies reduced the infiltration of leukaemia cells into host tissues (t–v, n = 5 mice) and even CD8+ cells were depleted (w, n = 5 mice). x–z, C57BL/6 mice were intravenously injected with human LILRB4-expressing mouse AML C1498 cells (3 × 106 cells per mouse) that expressed GFP. Anti-LILRB4-N297A antibodies (n = 9 mice) or control IgG (n = 9 mice) were intravenously injected at 6, 9, 12, 15 and 18 days after implantation of tumour cells. Anti-LILRB4 antibodies decreased the percentage of leukaemia cells in bone marrow (x). Anti-LILRB4 antibodies increased CD8+ T cells (y). The percentage of CD8+ T cells in bone marrow was significantly negatively correlated with the percentage of leukaemia cells (z). c, i, l, Experiments repeated independently three times with similar results. See Methods for definition of box plot elements in b, e, g, h, j, k, o, p, t–y. All P values (except n, long-rank test; and q, r, z, Pearson’s correlation) from two-tailed Student’s t-test.
Extended Data Fig. 4 Anti-LILRB4 antibodies reduce leukaemia development by restoring autologous T cells in PDX mice and inhibiting primary AML cell infiltration.
a, Primary peripheral blood or bone marrow mononuclear AML cells (5 × 106 to 1 × 107 cells per mouse) from each of sixteen human patients (three shown in Fig. 1g–i, also see Supplementary Table 5) were injected into NSG mice followed by treatment with IgG or anti-LILRB4 antibodies (10 mg kg−1 twice a week by intravenous injection). Percentages of human CD45+LILRB4+ AML cells collected from haematopoietic tissues including bone marrow, spleen, liver and peripheral blood 2–4 months after transplantation, as determined by flow cytometry, are shown. b, Percentages of autologous human T cells collected from haematopoietic tissues including bone marrow, spleen, liver and peripheral blood 2–4 months after transplantation, as determined by flow cytometry; and representative flow plots of CD3+CD8+ T cells in bone marrow of mice in three PDXs. n = 8 biologically independent samples for all PDXs except AML#11 (n = 20 biologically independent samples) in a, b. c–e, Comparison of infiltration of human primary monocytic AML cells in NSG mice (n = 5 mice) after treatment with anti-LILRB4 antibody or IgG control. c, d, Primary human peripheral blood mononuclear cells from patients with monocytic AML were injected. The quantifications in c are also shown in Fig. 2l–n. e, Mouse liver cells with xenografted primary human monocytic AML cells (human CD45+LILRB4+ cells) were injected. See Methods for definition of box plot elements in a–e. All P values from two-tailed Student’s t-test.
Extended Data Fig. 5 LILRB4 promotes infiltration of AML cells.
a, c, LILRB4 expression on mouse C1498 (a) or WEHI-3 (c) AML cells that stably express Lilrb4 (also known as Lilrb4a). b, d, Forced expression of LILRB4 did not affect proliferation of mouse C1498 (b, n = 3 biologically independent samples with mean and s.e.m.) or WEHI-3 (d, n = 3 biologically independent samples with mean and s.e.m.) AML cells. e, Forced expression of human LILRB4 promoted transendothelial migration of mouse AML WEHI-3 cells (n = 3 biologically independent samples with mean and s.e.m.). f, NSG mice (n = 6 mice) were injected with 1 × 106 THP-1 cells followed immediately by IgG or anti-LILRB4 antibody treatment and were monitored by bioluminescence imaging. g, h, Anti-LILRB4 antibodies decreased AML cell infiltration into internal organs. Mice were killed at 21 days for ex vivo bioluminescence imaging of internal organs after transplantation of 1 × 106 luciferase-expressing THP-1 cells. Images of luminescence flux (radiance) from representative mice are shown (g). 1, GI tract; 2, legs; 3, lung; 4, spleen; 5, liver; 6, kidneys; 7, brain; 8, heart. Infiltrated leukaemia cells formed tumour nodules in liver (h). i, j, Anti-LILRB4 antibodies did not affect LILRB4-negative cancer cells. LILRB4 is expressed on THP-1 and MV4-11 human AML cells but not on U937 cells as analysed by flow cytometry (i). Isotype IgG was used as control. NSG mice were injected with U937 human AML cells, which do not express LILRB4, and then treated with anti-LILRB4 antibodies (j). IgG served as control antibodies. Mice were killed at day 25 post-transplant for analysis of LV, BM, SP and PB by flow cytometry. The presence of human AML cells was detected by anti-human CD45 antibody staining (n = 4 mice with mean and s.e.m.). k–t, Anti-LILRB4 antibodies decreased infiltration of THP-1 (k–o) or MV4-11 (p–t) human AML cells. Comparison of transendothelial migration abilities of GFP-expressing THP-1 (k) or CFSE-labelled MV4-11 (p) cells after treatment with anti-LILRB4 antibody or IgG control in a transwell assay (n = 3 biologically independent samples with mean and s.e.m.). Comparison of the homing abilities of GFP-expressing THP-1 or CFSE-labelled MV4-11 cells (5 × 106 per mouse) that were injected into NSG mice followed immediately by IgG or anti-LILRB4 antibody treatment. Numbers of leukaemia cells (GFP+ in l, CFSE+ in q) in LV, SP and BM normalized to that in PB as determined by flow cytometry 20 h after injection (n = 5 mice). NSG mice were injected with 1 × 106 THP-1 or MV4-11 cells followed immediately by IgG or anti-LILRB4 antibody treatment (n = 6 mice for THP-1 or 5 mice for MV4-11 xenografts). Percentages of MV4-11 cells (stained with anti-human CD45) as determined by flow cytometry in indicated organs at day 21 post-transplant (m, r), overall survival (n, s) and body weights as a function of time (o, t) are shown. u, Targeted immune cell populations were depleted in NSG mice. Representative flow cytometry plots demonstrating successful reduction of NK cell (CD45+CD49b+), macrophage (CD11b+F4/80+), and neutrophil (CD11b+CD11c–) frequency in NSG mice depleted of the respective immune cell subtype by treatment with anti-asialo GM1 antibodies, clodronate liposomes and anti-Ly6G antibodies, respectively, compared to non-depleted (wild-type) NSG mice. v, CFSE-labelled MV4-11 cells (5 × 106 per mouse) were injected into NSG mice in which the respective innate immune cells were depleted, followed immediately by IgG or anti-LILRB4-N297A antibody treatment (n = 5 mice). Numbers of leukaemia cells (CFSE positive) in LV, SP and BM normalized to that in PB at 20 h post-injection. a, c, i, u, Experiments repeated independently three times with similar results. See Methods for definition of box plot elements in l, m, o, q, r, t, v. All P values (except n, s, log-rank test) are from two-tailed Student’s t-test.
Extended Data Fig. 6 APOE induces LILRB4 activation to suppress T cells and support AML cell migration in vitro.
a, Schematic of the LILRB4 reporter system. b, Human and mouse integrin heterodimer proteins cannot activate the LILRB4 reporter (n = 3 biologically independent samples with mean and s.e.m.). Human and mouse sera were used as positive controls. The threshold of activation is twice that of negative control treatment. c, Flow cytometry demonstrating that anti-LILRB4 antibody binds to human LILRB4 reporter cells. d, LILRB4 activation as indicated by percentage of GFP+ cells in the presence and absence of 10% human serum (HS) with or without anti-LILRB4 antibody or control IgG (n = 3 biologically independent samples with mean and s.e.m.). e, Flow chart of ligand identification of potential ligands of LILRB4 in human serum. f, Fractionation of LILRB4-stimulating activities from human serum by fast protein liquid chromatography. The positive control was 10% human serum. g, Proteins identified from the LILRB4 stimulating fractions by mass spectrometry. PSMs, peptide spectrum matches. h, Both human and mouse APOE proteins can activate LILRB4 reporter (n = 3 biologically independent samples with mean and s.e.m.). Human and mouse sera were used as positive controls. The threshold of activation is twice that of negative control treatment. i, APOE proteins from different sources all activate LILRB4. APOE (20 µg ml−1) purified from human plasma, His-tagged or tag-free recombinant human APOE (rhAPOE) (20 µg ml−1) expressed by 293T mammalian cells, or rhAPOE (20 µg ml−1) expressed by bacteria all activate the LILRB4 reporter. These APOE all represent human APOE3 (n = 3 biologically independent samples with mean and s.e.m.). j, APOE2, APOE3 and APOE4 all activate the LILRB4 reporter (n = 3 biologically independent samples with mean and s.e.m.). Forty micrograms per millilitre APOEs were coated on plates or directly added in cell culture medium (soluble). k, l, Three APOE isoforms bind to human LILRB4. k, Binding kinetics of APOE2, 3 and 4 to LILRB4-Fc were measured using SPR. LILRB4-Fc was immobilized on protein A biosensor tips and incubated with APOE concentrations ranging from 1.5625 nM to 100 nM. l, Binding kinetics of APOE2, 3, and 4 to LILRB4-Fc were measured using bio-layer interferometry (Octet). LILRB4-Fc was immobilized on protein A biosensor tips and incubated with APOE concentrations ranging from 44 nM to 1,176 nM. m, As shown in Fig. 3h, mutation of W106 and Y121, located in the first immunoglobulin domain and in the linker between two immunoglobulin domains, respectively, significantly reduced activation of LILRB4 by APOE. n, p, Examination of APOE expression in APOE-knockout THP-1 and MV4-11 cells by immunoblots. Primary T cells and irradiated THP-1 or MV4-11 cells (E:T = 2:1) were incubated in the lower and upper chambers, respectively. T cells were photographed (o, q, scale bar, 100 µm) and quantified by flow cytometry (Fig. 3i and r, n = 4 biologically independent samples) after 7 days. s, t, Loss of APOE suppresses transendothelial migration of human AML THP-1 and MV4-11 cells (n = 4 biologically independent samples with mean and s.e.m.). c, k, l, n–q, These experiments were repeated independently three times with similar results. See Methods for definition of box plot elements in r. All P values are from two-tailed Student’s t-test.
Extended Data Fig. 7 LILRB4 upregulates phosphorylation of SHP-2, NFκB signalling and expression of uPAR and ARG1 to suppress T cell activity and support leukaemia migration.
a, Phosphorylated SHP-2 and IκBα were downregulated upon LILRB4 knockout in MV4-11 cells. b, Co-immunoprecipitation demonstrated that LILRB4 interacts with SHP-2 in THP-1 cells. c, SHP1, SHP2 and SHIP were individually knocked out by CRISP–Cas9 in THP-1 cells as detected by western blotting. d, Primary T cells and irradiated THP-1 cells (E:T = 2:1) were cultured in the lower and upper chambers, respectively. T cells were photographed (scale bar, 100 µm) after 7 days. e, f, Two different NFκB inhibitors restored T cell proliferation from the suppression by THP-1 cells in an LILRB4-dependent manner (n = 4 biologically independent samples). THP-1 cells were pretreated with various doses of NFκB inhibitors for 1 h. Primary T cells and irradiated pretreated THP-1 cells (E:T = 2:1) were cultured in the lower and upper chambers, respectively. T cells were photographed (e, scale bar, 100 µm) and analysed by flow cytometry (f) after 7 days. g, h, Loss of LILRB4 decreased secreted protein production in THP-1 cells as determined by a human cytokine antibody array (g) and the blot intensities were quantified by ImageJ software (h, n = 3 biologically independent samples with mean and s.e.m.). Red boxes indicate proteins that were changed upon LILRB4 knockout; blue boxes indicate positive controls. i, Surface uPAR was downregulated in LILRB4KO THP-1 and MV4-11 AML cells. j, T cells were incubated with irradiated indicated THP-1 cells supplemented with indicated concentrations of recombinant uPAR proteins for 7 days. T cells were photographed. k, T cells isolated from healthy donors were cultured with anti-CD3/CD28-coated beads and rhIL-2 and supplemented with indicated concentrations of uPAR proteins for 3 days (n = 4 biologically independent samples). Representative cells were photographed using an inverted microscope and T cells were analysed by flow cytometry. l, Expression of uPAR and ARG1 is downregulated in in LILRB4 knockout THP-1 and MV4-11 AML cells. m, Arginase activity as determined by a colorimetric method (DARG100, BioAssay system) was decreased in condition medium of LILRB4KO THP-1 and MV4-11 cells (n = 3 biologically independent samples with mean and s.e.m.). n, Primary T cells and irradiated indicated THP-1 cells (E:T = 2:1) were incubated in the lower and upper chambers, respectively, and were supplemented with 0.002 U/l recombinant ARG1 proteins for 7 days. T cells were photographed. o, T cells isolated from healthy donors were cultured with anti-CD3/CD28-coated beads and rhIL-2 and supplemented with indicated concentrations of ARG1 proteins for 3 days (n = 4 biologically independent samples). Representative cells were photographed using an inverted microscope and T cells were analysed by flow cytometry. p, Autologous T cells isolated from individual patients with monocytic AML were incubated with irradiated LILRB4-positive or LILRB4-negative primary leukaemia cells from the same patients at an E:T of 10:1, supplemented with recombinant anti-LILRB4 antibodies, APOE-VLDL, uPAR or ARG1. pT, patient T cells. After culture with anti-CD3/CD28/CD137-coated beads and rhIL-2 for 14 days, T cells were stained with anti-CD3, anti-CD4, and anti-CD8 antibodies and analysed by flow cytometry. n = 3 biologically independent samples with mean and s.e.m. q, Supplementation of recombinant uPAR or ARG1 to the medium rescued the decrease in transmigration ability of LILRB4KO THP-1 or LILRB4KO MV4-11 cells across endothelium (n = 3 biologically independent samples with mean and s.e.m.). Scale bar, 100 µm. a–e, g, i, j, l, n, Experiments repeated independently three times with similar results. See Methods for definition of box plot elements in f, k, o. All P values from two-tailed Student’s t-test. See raw data for p in Source Data.
Extended Data Fig. 8 Detection of SHP-2–NFκB signalling and uPAR and ARG1 expression in primary human monocytic AML cells.
a, LILRB4-positive or -high CD33+ AML cells (red box) and LILRB4-negative or -low CD33+ AML cells (blue box) were gated for further intracellular staining of SHP-2 phosphorylated at Y580, IKKα/β phosphorylated at S176/S180, NFκB phosphorylated at S529, uPAR and ARG1. Isotype IgG was used as negative control. Red numbers indicate MFI of LILRB4-positive or -high CD33+ AML cells; blue numbers indicate MFIs of LILRB4-negative or -low CD33+ AML cells. This experiment was repeated with eight individual patient samples with similar results. b, Quantification of individual staining in LILRB4-positive or -high CD33+ AML cells versus in LILRB4-negative or -low CD33+ AML cells. n = 8 independent patients; see Methods for definition of box plot elements. P values from two-tailed Student’s t-test. c, Schematic for the mechanisms by which LILRB4 suppresses T cells and promotes leukaemia infiltration.
Extended Data Fig. 9 Comparison of LILRB4-mediated intracellular signalling in leukaemia cells and in normal haematopoietic cells.
a, Comparison of LILRB4 surface expression on normal monocytes from healthy donors (n = 25 individual donors with mean and s.e.m.) and neoplastic monocytic cells from patients with AML (n = 53 individual patients with mean and s.e.m.). b, Comparison of LILRB4 surface expression on normal monocytes from two healthy donors and on wild-type and LILRB4KO THP-1 cells. This experiment was repeated independently three times with similar results. c, Anti-LILRB4 antibody did not affect homing ability of normal monocytes. Human normal monocytes (b) selected through CD14-positive selection. These isolated monocytes were pooled and stained by CFSE. After staining, monocytes (5 × 106 for each mouse) were injected into NSG mice followed immediately by antibody treatment, and then the mice (n = 4 mice, see Methods for definition of box plot elements) were killed 20 h after transplant. The number of CFSE+ cells in liver, spleen and bone marrow were normalized to that in peripheral blood as determined by flow cytometry. d, e, APOE activates LILRB4 intracellular signalling in leukaemia cells. Indicated THP-1 cells and primary AML (M5) cells were serum-starved overnight and then treated with the indicated concentration of human recombinant APOE protein for the indicated time. Phospho-SHP-2, phospho-NFκB and ARG1 were examined by western blotting. f, The effect of APOE on normal monocytes or in vitro differentiated macrophages. Normal monocytes were isolated from healthy donors and macrophages were derived from these monocytes after one week of differentiation in vitro. Cells were serum-starved overnight and then treated with the indicated concentrations of human recombinant APOE protein for the indicated times. Phospho-SHP-2, phospho-NFκB and ARG1 were examined by western blotting. g, APOE induces uPAR upregulation on AML cells. Normal monocytes were isolated from healthy donors. Indicated primary AML cells and normal monocytes were serum-starved overnight and then treated with 20 μg ml−1 human recombinant APOE protein for eight hours. Surface uPAR was examined by flow cytometry. Representative flow plots are shown and MFIs are shown in top right corner (black, PBS control; red, APOE treatment). Experiments were performed three times with similar results. P values from two-tailed Student’s t-test.
Extended Data Fig. 10 Anti-LILRB4 does not affect engraftment of normal haematopoietic cells.
a, LILRB4 and CD34 co-staining patterns for representative samples of human cord blood mononuclear cells (hCB MNCs). N/G, neutrophils and granulocytes; M/D, monocytes, macrophages and dendritic cells; L/P, lymphocytes, haematopoietic stem and progenitor cells. This experiment was repeated independently three times with similar results. b, Anti-LILRB4 antibody did not affect homing ability of normal haematopoietic progenitor cells. hCB MNCs (1 × 107) were injected into NSG mice followed immediately by antibody treatment, and then the mice (n = 3 mice with mean and s.e.m.) were killed 20 h after transplant. The number of CD45+CD34+ HSCs in liver, spleen, and bone marrow were normalized to that in peripheral blood as determined by flow cytometry. c–e, Anti-LILRB4 antibodies inhibited leukaemia development in hCB-humanized NSG mice. c, Schematic of the experiment to test whether anti-LILRB4 antibody inhibits leukaemia development in hCB-humanized NSG mice. d, Leukaemia development was monitored over time by luminescence imaging. This experiment was repeated independently twice with similar results. e, Frequency of engrafted leukaemia, normal human cells, including human B cells, human myeloid cells and human T cells in peripheral blood over time and haematopoietic tissues of hCB-humanized mice at 24 days after leukaemia transplantation. n = 3 mice with mean and s.e.m. All P values from two-tailed Student’s t-test.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1 and Supplementary Tables 1-5.
Rights and permissions
About this article
Cite this article
Deng, M., Gui, X., Kim, J. et al. LILRB4 signalling in leukaemia cells mediates T cell suppression and tumour infiltration. Nature 562, 605–609 (2018). https://doi.org/10.1038/s41586-018-0615-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0615-z
Keywords
This article is cited by
-
Microglia LILRB4 upregulation reduces brain damage after acute ischemic stroke by limiting CD8+ T cell recruitment
Journal of Neuroinflammation (2024)
-
Sirpα on tumor-associated myeloid cells restrains antitumor immunity in colorectal cancer independent of its interaction with CD47
Nature Cancer (2024)
-
Galectin 7 leads to a relative reduction in CD4+ T cells, mediated by PD-1
Scientific Reports (2024)
-
LILRB4 regulates multiple myeloma development through STAT3-PFKFB1 pathway
Cell Death & Disease (2024)
-
Identification of a novel monocyte/macrophage-related gene signature for predicting survival and immune response in acute myeloid leukemia
Scientific Reports (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.