Abstract
The emergence of multiple axes is an essential element in the establishment of the mammalian body plan. This process takes place shortly after implantation of the embryo within the uterus and relies on the activity of gene regulatory networks that coordinate transcription in space and time. Whereas genetic approaches have revealed important aspects of these processes1, a mechanistic understanding is hampered by the poor experimental accessibility of early post-implantation stages. Here we show that small aggregates of mouse embryonic stem cells (ESCs), when stimulated to undergo gastrulation-like events and elongation in vitro, can organize a post-occipital pattern of neural, mesodermal and endodermal derivatives that mimic embryonic spatial and temporal gene expression. The establishment of the three major body axes in these ‘gastruloids’2,3 suggests that the mechanisms involved are interdependent. Specifically, gastruloids display the hallmarks of axial gene regulatory systems as exemplified by the implementation of collinear Hox transcriptional patterns along an extending antero-posterior axis. These results reveal an unanticipated self-organizing capacity of aggregated ESCs and suggest that gastruloids could be used as a complementary system to study early developmental events in the mammalian embryo.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
References
Rossant, J. & Joyner, A. L. Towards a molecular-genetic analysis of mammalian development. Trends Genet. 5, 277–283 (1989).
Simunovic, M. & Brivanlou, A. H. Embryoids, organoids and gastruloids: new approaches to understanding embryogenesis. Development 144, 976–985 (2017).
van den Brink, S. C. et al. Symmetry breaking, germ layer specification and axial organisation in aggregates of mouse embryonic stem cells. Development 141, 4231–4242 (2014).
Sasai, Y., Eiraku, M. & Suga, H. In vitro organogenesis in three dimensions: self-organising stem cells. Development 139, 4111–4121 (2012).
Turner, D. A., Baillie-Johnson, P. & Martinez Arias, A. Organoids and the genetically encoded self-assembly of embryonic stem cells. BioEssays 38, 181–191 (2016).
Turner, D. A. et al. Anteroposterior polarity and elongation in the absence of extraembryonic tissues and spatially localised signalling in gastruloids, mammalian embryonic organoids. Development 144, 3894–3906 (2017).
Zhao, R. et al. Loss of both GATA4 and GATA6 blocks cardiac myocyte differentiation and results in acardia in mice. Dev. Biol. 317, 614–619 (2008).
Carninci, P. et al. The transcriptional landscape of the mammalian genome. Science 309, 1559–1563 (2005).
Chal, J. et al. Recapitulating early development of mouse musculoskeletal precursors of the paraxial mesoderm in vitro. Development 345, dev157339 (2018).
Olivera-Martinez, I. et al. Major transcriptome re-organisation and abrupt changes in signalling, cell cycle and chromatin regulation at neural differentiation in vivo. Development 141, 3266–3276 (2014).
Dessaud, E., McMahon, A. P. & Briscoe, J. Pattern formation in the vertebrate neural tube: a sonic hedgehog morphogen-regulated transcriptional network. Development 135, 2489–2503 (2008).
Wang, P. et al. A molecular signature for purified definitive endoderm guides differentiation and isolation of endoderm from mouse and human embryonic stem cells. Stem Cells Dev. 21, 2273–2287 (2012).
Beck, F., Erler, T., Russell, A. & James, R. Expression of Cdx-2 in the mouse embryo and placenta: possible role in patterning of the extra-embryonic membranes. Dev. Dyn. 204, 219–227 (1995).
Blum, M. et al. Gastrulation in the mouse: the role of the homeobox gene goosecoid. Cell 69, 1097–1106 (1992).
Lee, J. D. & Anderson, K. V. Morphogenesis of the node and notochord: The cellular basis for the establishment and maintenance of left–right asymmetry in the mouse. Dev. Dyn. 237, 3464–3476 (2008).
Deschamps, J. & van Nes, J. Developmental regulation of the Hox genes during axial morphogenesis in the mouse. Development 132, 2931–2942 (2005).
Deschamps, J. & Duboule, D. Embryonic timing, axial stem cells, chromatin dynamics, and the Hox clock. Genes Dev. 31, 1406–1416 (2017).
Scotti, M. & Kmita, M. Recruitment of 5′ Hoxa genes in the allantois is essential for proper extra-embryonic function in placental mammals. Development 139, 731–739 (2012).
Harrison, S. E., Sozen, B., Christodoulou, N., Kyprianou, C. & Zernicka-Goetz, M. Assembly of embryonic and extra-embryonic stem cells to mimic embryogenesis in vitro. Science 356, eaal1810 (2017).
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
Kadoshima, T. et al. Self-organization of axial polarity, inside-out layer pattern, and species-specific progenitor dynamics in human ES cell-derived neocortex. Proc. Natl Acad. Sci. USA 110, 20284–20289 (2013).
Eiraku, M. et al. Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature 472, 51–56 (2011).
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
Rivron, N. C. et al. Blastocyst-like structures generated solely from stem cells. Nature 557, 106–111 (2018).
Heisenberg, C. P. & Bellaiche, Y. Forces in tissue morphogenesis and patterning. Cell 153, 948–962 (2013).
Hooper, M., Hardy, K., Handyside, A., Hunter, S. & Monk, M. HPRT-deficient (Lesch–Nyhan) mouse embryos derived from germline colonization by cultured cells. Nature 326, 292–295 (1987).
Fehling, H. J. et al. Tracking mesoderm induction and its specification to the hemangioblast during embryonic stem cell differentiation. Development 130, 4217–4227 (2003).
Freyer, L. et al. A loss-of-function and H2B-Venus transcriptional reporter allele for Gata6 in mice. BMC Dev. Biol. 15, 38 (2015).
Papanayotou, C. et al. A novel Nodal enhancer dependent on pluripotency factors and Smad2/3 signaling conditions a regulatory switch during epiblast maturation. PLoS Biol. 12, e1001890 (2014).
Deluz, C. et al. A role for mitotic bookmarking of SOX2 in pluripotency and differentiation. Genes Dev. 30, 2538–2550 (2016).
Turner, D. A. et al. Wnt/β-catenin and FGF signalling direct the specification and maintenance of a neuromesodermal axial progenitor in ensembles of mouse embryonic stem cells. Development 141, 4243–4253 (2014).
Ring, D. B. et al. Selective glycogen synthase kinase 3 inhibitors potentiate insulin activation of glucose transport and utilization in vitro and in vivo. Diabetes 52, 588–595 (2003).
Baillie-Johnson, P., van den Brink, S. C., Balayo, T., Turner, D. A. & Martinez Arias, A. Generation of aggregates of mouse embryonic stem cells that show symmetry breaking, polarization and emergent collective behaviour in vitro. J. Vis. Exp. 105, e53252 (2015).
Girgin, M. et al. Generating gastruloids from mouse embryonic stem cells. Protoc. Exch. https://doi.org/10.1038/protex.2018.094 (2018).
Guerreiro, I. et al. Reorganisation of Hoxd regulatory landscapes during the evolution of a snake-like body plan. eLife 5, e16087 (2016).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Blankenberg, D. et al. Galaxy: a web-based genome analysis tool for experimentalists. Curr. Protoc. Mol. Biol. 89, 19.10.1–19.10.21 (2010).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A. R. BEDTools: The Swiss-army tool for genome feature analysis. Curr. Protoc. Bioinformatics 47, 1–34 (2014).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2014).
Yates, A. et al. Ensembl 2016. Nucleic Acids Res. 44, D710–D716 (2016).
Roberts, A., Trapnell, C., Donaghey, J., Rinn, J. L. & Pachter, L. Improving RNA-seq expression estimates by correcting for fragment bias. Genome Biol. 12, R22 (2011).
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics 10, 48 (2009).
Supek, F., Bosnjak, M., Skunca, N. & Smuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE 6, e21800 (2011).
Woltering, J. M., Noordermeer, D., Leleu, M. & Duboule, D. Conservation and divergence of regulatory strategies at Hox loci and the origin of tetrapod digits. PLoS Biol. 12, e1001773 (2014).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Shawlot, W., Deng, J. M. & Behringer, R. R. Expression of the mouse cerberus-related gene, Cerr1, suggests a role in anterior neural induction and somitogenesis. Proc. Natl Acad. Sci. USA 95, 6198–6203 (1998).
Acknowledgements
We thank members of the Duboule, Lutolf and Martinez-Arias laboratories for sharing material and discussions, J. Deschamps for DNA clones, D. Trono for the Oct4::GFP iPSC line, the histology platform and the Gene Expression Core Facility (EPFL) as well as the genomics platform at Geneva University. This work was supported by funds from the BBSRC (No. BB/M023370/1 and BB/P003184/1 to A.M.A.), an NC3Rs David Sainsbury Fellowship (No. NC/P001467/1 to D.A.T.), an Engineering and Physical Sciences Research Council (EPSRC) Studentship (to P.B.-J.), a Company of Biologists Development Travelling Fellowship (DEVTF-151210 to P.B.-J.), a Newnham College Constance Work Junior Research Fellowship (to N.M.), the École Polytechnique Fédérale de Lausanne (D.D. and M.P.L.), the University of Geneva (D.D.), the Swiss National Research Fund (No. 310030B_138662 to D.D.) and the European Research Council grants ERC MOFDH (No. 250316 to A.M.A.), RegulHox (No 588029 to D.D.) and STEMCHIP (No 311422 to M.P.L.).
Reviewer information
Nature thanks J. Briscoe and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
L.B., M.G. and D.A.T. produced gastruloids. L.B. performed WISH, imaged and analysed gastruloids, performed RNA extraction and qPCR analysis, analysed and interpreted RNA-seq and wrote the manuscript. N.M. performed bioinformatics analyses, analysed, interpreted and produced figures, and contributed to the manuscript. M.G. contributed to the gastruloid culture protocol, and performed immunostaining and confocal analysis. D.A.T. contributed to the protocols, collected, stained, imaged and analysed gastruloids, and contributed to the manuscript. P.B.-J. contributed to the protocol and collected gastruloids. A.-C.C. cultured gastruloids and performed WISH. M.P.L., D.D. and A.M.A. conceived the project. M.P.L. corrected the manuscript. D.D. and A.M.A. analysed and interpreted the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 z stacks used for 3D rendering of gastruloids.
a–d, Gastruloids produced using Gata6H2B-Venus ESCs treated with a pulse of the GSK3 inhibitor Chi between 48 h and 72 h AA and fixed at 48 h (a), 72 h (b), 96 h (c) or 120 h AA (d) and imaged by confocal microscopy. BRA and SOX2 proteins are stained in red and white, respectively. VENUS signal (green) reports Gata6 expression and Hoechst 33342 (blue) marks the nuclei. Gastruloids correspond to the 3D renderings shown in Fig. 1a. Each fluorescent channel is displayed to the right of the merged image. Gata6 (a) or Gata6 and SOX2 (b) signals were undetectable, and are therefore not shown. Three z sections are shown for each gastruloid. The bright-field outline of each gastruloid is indicated by the dashed lines. Each panel is representative of an experiment perfomed in parallel in seven independent biological replicates showing the same expression pattern. Scale bars are as indicated.
Extended Data Fig. 2 Transcriptional profiling of mouse embryos and gastruloids.
a. Heat map showing the temporal evolution of 97 out of the 250 most variable genes throughout embryonic development from E6.5 to E9.5 (left) and their corresponding expression over the gastruloid time course, from 24 h to 168 h AA (right). Expression levels are indicated by colour scale from blue to red (bottom left). Genes were clustered according to their expression behaviour in the embryo. Enriched GO term categories were identified for each cluster using the Gorilla and REVIGO tools (see Supplementary Information dataset 1). Finally, a functional classification of each cluster was established based on the identified GO term categories and literature-based evidence. b, Expression of markers for different embryonic tissues through the gastruloid time course. The two replicates of each time point are represented by a triangle and a circle, respectively. The black dotted line in each plot represents the average behaviour of the genes displayed in the plot. For gastruloids, n = 2 independent biological replicates per time point; for E6.5 and E7.8 embryos, n = 3 independent biological replicates; for E8.5 and E9.5 embryos, n = 2 independent biological replicates. c, PCA analysis of RNA-seq datasets from either pooled or individual gastruloids using the top 1,000 most highly expressed genes. Despite different strategies used for RNA-seq of pooled versus individual gastruloids (accounting for the sample segregation across PC1), their clustering illustrates both the homogeneity of gastruloid cultures and the representativeness of pooled samples to single gastruloid samples. For individual gastruloid RNA-seq: n = 10 independent biological replicates per time point.
Extended Data Fig. 3 Gastruloids display spatio-temporal organization in the expression profiles of neural, mesodermal and endodermal marker genes.
a–f, The expression profiles of several genes usually expressed in the embryonic neural, mesodermal and endodermal domains were analysed by plotting RNA-seq data from the pooled gastruloids in heat maps of scaled gene expression (2 independent biological replicates per time point) (a, c, e) and/or by WISH (b, d, f). a, b, Genes usually expressed in different types of mesoderm precursors in the embryo (for example, Tcf15 in paraxial somatic mesoderm, Osr1 in intermediate mesoderm, Bra in tail bud, notochord and presomitic mesoderm, and Pecam1 in lateral plate mesoderm) were expressed in reproducible and spatially restricted domains within the gastruloids. c, d, Expression of different neural markers was detected in our RNA-seq (c). Transcripts of genes such as Lfng or Irx3 formed continuous and homogenous domains located in the central and dorsal portion of the gastruloids, reminiscent of their expression domains in the embryo (d, top panels). Genes involved in Notch signalling in neural progenitors (Hes5, Dll1) and in the terminal differentiation of neural precursors (Phox2a, Mnx1) displayed a salt-and-pepper expression pattern, consistent with the lack of an organized neural-tube structure (see also Extended Data Figs. 4a, c, 5). However, the latter mRNAs also displayed a graded distribution along the anterior-to-posterior extension of the gastruloid axis and were absent from its posterior half (empty red arrowheads). e, f, Endoderm-specific genes were also expressed in gastruloids. In particular, genes expressed in the embryonic digestive tract were consistently found on the ventral side of gastruloids. For each gene, the proportion of gastruloids displaying the reported expression pattern is shown in the upper right corner of the image, expressed as a fraction of the total number. Experimental statistics are provided in Supplementary Information dataset 3. Scale bar, 100 µm.
Extended Data Fig. 4 Tissue organization in gastruloids.
a, Gastruloids formed from Sox1GFP;BramCherry (SBR) line and stained for Sox2 expression (Sox1GFP and SOX2 signals are displayed in green and magenta, respectively). White arrowheads indicate tubular SOX2/Sox1-positive neural structures. Red arrowheads point to the presumptive digestive tube. b, WISH on 8-µm transverse cryosections of gastruloids at 144 h AA using Sox2 and Meox1 antisense probes, counter-stained with Nuclear Fast Red. Sox2-positive cells localized predominantly in a compact dorsal domain, whereas Meox1 signals were found in two bilateral domains. The domain of expression of each gene is outlined with white dashed lines. c, Haematoxylin and eosin staining of transverse paraffin sections of different gastruloids at 120 h AA, showing the diversity of cell types and several levels of tissue organization. d, Gastruloids formed from Sox1GFP;BramCherry ESCs were fixed and stained at 168 h AA for OLIG2 (top, white), PAX3 (middle, red) and PAX7 (bottom, red). Scale bars as indicated. c, d, Gastruloids formed from Sox1GFP; BramCherry ESCs collected at 168 h AA and stained for SOX17 (magenta, c) or CDX2 (magenta, d). Scale bars as indicated. All immunostaining experiments were repeated twice, with three biological replicates per experiment, with similar results.
Extended Data Fig. 5 Double-FISH staining shows organized gene expression across the three main gastruloid axes.
a–e, Double-FISH staining of gastruloids at 144 h AA with Meox1 and Cyp26a1 (a), Sox2 and Shh (b), Sox2 and Meox1 (c), Meox1 and Hoxd4 (d) or Sox2 and Hoxd4 (e). a–e, Experiments were repeated twice in three biological replicates with similar results. Scale bar, 200 µm.
Extended Data Fig. 6 A node-like structure and left–right asymmetry in gastruloids.
a, b, Gastruloids formed from NodalYFP ESCs were fixed at 120 h AA. They were stained for CDX2, YFP (NodalYFP) and E-cadherin (a, top panel), CDX2, YFP (NodalYFP, green) and phalloidin (a, bottom panel) or for CDX2, YFP and E-cadherin (both with an Alexa-488 secondary antibody), and SOX2 (b). Maximum intensity projection of a representative gastruloid in b, with the node-like structure highlighted. Hoechst 33342 labels the nuclei (greyscale in a, blue in b). Data are representative of one experiment performed in three independent biological replicates. c, d, In situ hybridization showing expression of the indicated genes in gastruloids at different time-points AA. *, presumptive node-like cells. White arrowheads point towards Nodal-expressing cells distributed asymmetrically, on the lateral side of the gastruloid. Whereas Nodal was expressed in the presumptive node region from 96 h AA, no clear asymmetry in transcript distribution was observed at that stage. e, Three-dimensional renderings of confocal stacks of 120 h gastruloids containing a NodalYFP reporter gene (green) and stained for SOX2 (white) and BRA (red) proteins. SOX2 signal identifies dorsal cells. Left and right panels show the same gastruloid, imaged from two different polar directions that is, top (dorsal) and bottom (ventral) or ‘left’ and ‘right’, depending on the orientation of the gastruloid. Insets in specific panels show a cross-section through the gastruloid at the indicated z plane. White arrowheads indicate the region of biased Nodal expression. Empty white arrowheads point to the node-like cells marked by the NodalYFP reporter gene (see also Fig. 4d). These results are consistent with the asymmetric distribution of Nodal transcripts at 120–144 h AA. f, In situ hybridization showing expression of Cer1 in 120 h AA (left) and 144 h AA (right) gastruloids. The gastruloid midline is marked by a dashed white line. At this stage, Cer1 is expressed in the presumptive embryonic somitic territory47 and the pattern in gastruloids may reflect this specificity. In c, d and f, the proportion of gastruloids displaying the reported expression pattern is shown at the bottom left corner of each image, expressed as a fraction of the total number of specimens analysed (see Supplementary Information dataset 3 for a complete statistical report).
Extended Data Fig. 7 z stacks used for 3D rendering of gastruloids.
a, b, Dorsal (a) and ventral (b) sections of the same representative gastruloid shown in the 3D renderings in Fig. 3d, fixed and stained at 120 h for NodalYFP (green), BRA (red) and SOX2 (white). Hoechst 33342 was used to label nuclei. Data are representative of two independent experiments with n = 13 biological replicates in total (see Supplementary Information dataset 3 for a detailed statistical report). Scale bar, 100 µm.
Extended Data Fig. 8 Hox expression profiles in mouse embryos and gastruloids.
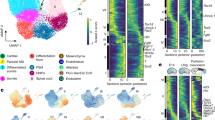
a, Heat map of unscaled gene expression in E6.5–E9.5 mouse embryos, showing levels of Hox gene transcripts over time. Between 2 and 3 independent biological replicates were used for each time point (indicated below each graph). b, RNA-seq mapping showing Hoxa and Hoxd gene expression in these embryos. After a first wave of transcription of 5′ Hoxa and Hoxd genes, which is likely to reflect their activation in extra-embryonic tissues, the HoxA and HoxD clusters were progressively transcribed between E7.8 and E9.5, when expression of Hox13 paralogues was detected. Each profile was averaged from independent biological replicates indicated in a. c, Heat map of unscaled gene expression in pooled gastruloids, showing Hox gene transcript levels over time. Two independent biological replicates were used per time point. d, RNA-seq mapping showing Hoxd gene expression in pooled gastruloids at different time points. Sub-groups of Hoxd genes are progressively activated between 72 h and 168 h AA, when expression of Hoxd13 starts to be detected (e). This resembles the temporal activation described in vivo (a, b). Each profile represents the average of two independent biological replicates. e, WISH of gastruloids collected at different time points, showing the detectable initiation of expression of different Hoxd genes. Each panel shows the earliest stage at which the indicated gene was detected (black arrowhead). Expression of Hoxd4 was already strong at 96 h AA, indicating that its transcripts are rapidly upregulated compared to Hoxd9, which is expressed at low levels at this stage. Scale bar, 100 µm. The fraction of gastruloids displaying the reported expression pattern is indicated in the upper right corner of each image. Experimental statistics are provided in Supplementary Information dataset 3.
Extended Data Fig. 9 Homogeneity in Hox transcript profiles for individual gastruloids.
a, PCA based on Hox-transcript datasets only, extracted from individually sequenced gastruloids across time points (10 individual organoids per time point, representing independent biological replicates). The analysis was carried out using the log2-transformed FPKM+1 value of all 39 Hox genes. Replicate batches of organoids primarily cluster according to their age at collection. The clustering revealed the low sample-to-sample variation. however, replicates were clearly separated by the temporal parameter, representing 93.6% of total sample variation. b, Comparison of Hoxa (top) and Hoxd (bottom) gene-expression profiles among individual gastruloids confirmed the low inter-sample variation among time points, illustrated with the 120 h AA condition. c, WISH of 168 h AA gastruloids showing the expression of different Hoxa paralogues. The proportion of gastruloids displaying the reported expression pattern is shown in the upper right corner of the image, expressed as a fraction of the total number. Experimental statistics are provided in Supplementary Information dataset 3. Scale bar, 100 µm.
Extended Data Fig. 10 Axial extension and spatio-temporal Hox expression patterns in iPSC-derived gastruloids.
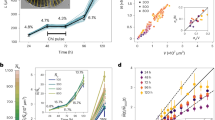
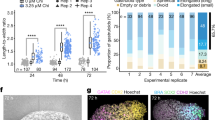
a. Dot plot representing the progression in the measured longitudinal extension of gastruloids produced either from ESCs or from iPSCs. In each case, 10 different gastruloids were measured at the different time points indicated. The median (round points) and the interquartile range (vertical bars) are reported. b, Light microscopy images showing representative examples of gastruloids at the different time points analysed in a. Zoom: 10×. Note that iPSC-derived gastruloids exhibit delay in their longitudinal extension rate and at 120 h AA they are markedly smaller than their ESC-derived counterparts. For this analysis, gastruloids were produced starting from the same number of cells (800 cells per well). c, Dot plots representing the Bra mRNA levels, showing comparable dynamics of this gene in both types of gastruloids. Circles represent individual data points and the short horizontal line represents the mean. The number of biological independent replicates (n) per condition is indicated. d, Confocal images showing the expression of Oct4, SOX2 and BRA (top) or of Oct4, SOX1 and CDX2 (bottom) in 120 h AA gastruloids derived from the iPSC line Oct4::Gfp (IpSL40N). iPSC-derived gastruloids were fixed and stained for SOX2 and BRA (top) and CDX2 and SOX1 (bottom). Oct4::GFP signal is shown in grey. Scale bar, 200 μm. In each case, data are representative of one experiment with three independent biological replicates. e, Dot plots representing the Hoxd mRNA levels in ESC- or iPSC-derived gastruloids collected at different time points AA. Each circle represents an independent biological replicate, the horizontal bars represent the mean value of the replicates. Both sets of gastruloids sequentially activated Hoxd gene expression. However, their temporal activation seemed to be delayed in iPSC gastruloids (especially that of the most 3′ Hoxd paralogues). f, WISH of 144 h AA gastruloids showing the expression of different Hoxd paralogues. Even though iPSC-derived gastruloids reproduced the antero-posterior Hoxd collinear expression, the Hoxd9 expression domain often extended more anteriorly in comparison to that in ESC-derived gastruloids (see Fig. 4c), occupying roughly the same domain as Hoxd4. Patches of Hoxd-negative cells were often observed within the Hoxd4/Hoxd9 expression domain (white). The fraction of gastruloids displaying the reported expression pattern is indicated in the upper right corner of each image. Experimental statistics are provided in Supplementary Information dataset 3. Scale bar, 100 µm.
Supplementary information
Supplementary Information
This file contains tables reporting information about seeding cell densities for gastruloid formation, primers used for ISH probe cloning, enzymes used for in vitro transcription of the probes and antibodies and dyes used in this study.
Supplementary Data 1
Provides excel sheets reporting: I) FPKM quantification from pooled gastruloids and embryonic RNAseq data; II) the heatmap plot of the 250 genes identified as the most variable across the different embryonic time-points and the corresponding heatmap of the same genes in pooled gastruloid samples; III) The complete results of the Gorilla and REVIGO analysis over the gene clusters identified in the embryonic heatmap plots.
Supplementary Data 2
Wald test results over the embryonic and gastruloid RNAseq samples. For embryonic samples, the E6.5 replicates were used as the reference group for the comparison. For the pooled and individual gastruloid RNAseq data the 24h AA timepoints were used as reference.
Supplementary Data 3
Experimental and biological replicates used in the different WISH experiments shown in main and Extended Data figures and the statistical analysis of the asymmetric vs. symmetric expression of Nodal, Cerberus and Meox1.
41586_2018_578_MOESM6_ESM.mp4
Video 1: 3D rendering of 96 h AA gastruloid (corresponding to Fig. 1a 96 h AA). a. Gastruloids were produced from Gata6H2B-Venus fixed at 96h AA, stained for Gata6 (green), BRA (red) and SOX2 (white) and imaged by confocal microscopy as indicated in Materials and Methods. Each fluorescent channel is highlighted in turn for clarity. Hoechst was used to stain the nuclei. Data are representative of an experiment perfomed in parallel in seven independent biological replicates showing the same expression pattern.
41586_2018_578_MOESM7_ESM.mp4
Video 2: 3D rendering of 120 h AA gastruloid (corresponding to Fig. 1a 120 h AA). a. Gastruloids were produced from Gata6H2B-Venus fixed at 120 h AA, stained for Gata6 (green), BRA (red) and SOX2 (white) and imaged by confocal microscopy as indicated in Materials and Methods. Each fluorescent channel is highlighted in turn for clarity. Hoechst was used to stain the nuclei. Data are representative of an experiment perfomed in parallel in seven independent biological replicates showing the same expression pattern.
Video 3: Nodal asymmetric expression in 120h AA gastruloid (Fig. 3d).
Gastruloids were produced from NodalYFP ESCs and treated with Chiron between 48h and 72h AA, fixed and stained at 120h for YFP (green), BRA (red). Data are representative of two independent experiments with n = 13 biological replicates in total.
Video 4: Nodal asymmetric expression in 120h AA gastruloid (Fig. 3d).
Gastruloids were produced from NodalYFP ESCs and treated with Chiron between 48h and 72h AA, fixed and stained at 120h for YFP (green), BRA (red). Data are representative of two independent experiments with n = 13 biological replicates in total, both of which yielded similar results.
Rights and permissions
About this article
Cite this article
Beccari, L., Moris, N., Girgin, M. et al. Multi-axial self-organization properties of mouse embryonic stem cells into gastruloids. Nature 562, 272–276 (2018). https://doi.org/10.1038/s41586-018-0578-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0578-0
Keywords
This article is cited by
-
Cellular and molecular control of vertebrate somitogenesis
Nature Reviews Molecular Cell Biology (2024)
-
Precise and scalable self-organization in mammalian pseudo-embryos
Nature Structural & Molecular Biology (2024)
-
Open-top multisample dual-view light-sheet microscope for live imaging of large multicellular systems
Nature Methods (2024)
-
Highly conserved and extremely evolvable: BMP signalling in secondary axis patterning of Cnidaria and Bilateria
Development Genes and Evolution (2024)
-
A 3D “sandwich” co-culture system with vascular niche supports mouse embryo development from E3.5 to E7.5 in vitro
Stem Cell Research & Therapy (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.