Abstract
The origin of the eukaryotic cell is unresolved1,2. Metagenomics sequencing has recently identified several potential eukaryotic gene homologues in Asgard archaea3,4, consistent with the hypothesis that the eukaryotic cell evolved from within the Archaea domain. However, many of these eukaryotic-like sequences are highly divergent and the organisms have yet to be imaged or cultivated, which brings into question the extent to which these archaeal proteins represent functional equivalents of their eukaryotic counterparts. Here we show that Asgard archaea encode functional profilins and thereby establish that this archaeal superphylum has a regulated actin cytoskeleton, one of the hallmarks of the eukaryotic cell5. Loki profilin-1, Loki profilin-2 and Odin profilin adopt the typical profilin fold and are able to interact with rabbit actin—an interaction that involves proteins from species that diverged more than 1.2 billion years ago6. Biochemical experiments reveal that mammalian actin polymerizes in the presence of Asgard profilins; however, Loki, Odin and Heimdall profilins impede pointed-end elongation. These archaeal profilins also retard the spontaneous nucleation of actin filaments, an effect that is reduced in the presence of phospholipids. Asgard profilins do not interact with polyproline motifs and the profilin–polyproline interaction therefore probably evolved later in the Eukarya lineage. These results suggest that Asgard archaea possess a primordial, polar, profilin-regulated actin system, which may be localized to membranes owing to the sensitivity of Asgard profilins to phospholipids. Because Asgard archaea are also predicted to encode potential eukaryotic-like genes involved in membrane-trafficking and endocytosis3,4, imaging is now necessary to elucidate whether these organisms are capable of generating eukaryotic-like membrane dynamics that are regulated by actin, such as are observed in eukaryotic cell movement, podosomes and endocytosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Eme, L., Spang, A., Lombard, J., Stairs, C. W. & Ettema, T. J. G. Archaea and the origin of eukaryotes. Nat. Rev. Microbiol. 15, 711–723 (2017).
López-García, P. & Moreira, D. Open questions on the origin of eukaryotes. Trends Ecol. Evol. 30, 697–708 (2015).
Spang, A. et al. Complex archaea that bridge the gap between prokaryotes and eukaryotes. Nature 521, 173–179 (2015).
Zaremba-Niedzwiedzka, K. et al. Asgard archaea illuminate the origin of eukaryotic cellular complexity. Nature 541, 353–358 (2017).
Gunning, P. W., Ghoshdastider, U., Whitaker, S., Popp, D. & Robinson, R. C. The evolution of compositionally and functionally distinct actin filaments. J. Cell Sci. 128, 2009–2019 (2015).
Dacks, J. B. et al. The changing view of eukaryogenesis—fossils, cells, lineages and how they all come together. J. Cell Sci. 129, 3695–3703 (2016).
Spang, A. et al. Asgard archaea are the closest prokaryotic relatives of eukaryotes. PLoS Genet. 14, e1007080 (2018).
Da Cunha, V., Gaia, M., Gadelle, D., Nasir, A. & Forterre, P. Lokiarchaea are close relatives of Euryarchaeota, not bridging the gap between prokaryotes and eukaryotes. PLoS Genet. 13, e1006810 (2017).
Xue, B. & Robinson, R. C. Guardians of the actin monomer. Eur. J. Cell Biol. 92, 316–332 (2013).
Courtemanche, N. & Pollard, T. D. Interaction of profilin with the barbed end of actin filaments. Biochemistry 52, 6456–6466 (2013).
Thompson, M. J. & Eisenberg, D. Transproteomic evidence of a loop-deletion mechanism for enhancing protein thermostability. J. Mol. Biol. 290, 595–604 (1999).
Ferron, F., Rebowski, G., Lee, S. H. & Dominguez, R. Structural basis for the recruitment of profilin–actin complexes during filament elongation by Ena/VASP. EMBO J. 26, 4597–4606 (2007).
Senju, Y. et al. Mechanistic principles underlying regulation of the actin cytoskeleton by phosphoinositides. Proc. Natl Acad. Sci. USA 114, E8977–E8986 (2017).
Michell, R. H. Inositol lipids: from an archaeal origin to phosphatidylinositol 3,5-bisphosphate faults in human disease. FEBS J. 280, 6281–6294 (2013).
Lane, N. & Martin, W. The energetics of genome complexity. Nature 467, 929–934 (2010).
Ezezika, O. C. et al. Incompatibility with formin Cdc12p prevents human profilin from substituting for fission yeast profilin: insights from crystal structures of fission yeast profilin. J. Biol. Chem. 284, 2088–2097 (2009).
Nag, S. et al. Ca2+ binding by domain 2 plays a critical role in the activation and stabilization of gelsolin. Proc. Natl Acad. Sci. USA 106, 13713–13718 (2009).
Xue, B., Leyrat, C., Grimes, J. M. & Robinson, R. C. Structural basis of thymosin-β4/profilin exchange leading to actin filament polymerization. Proc. Natl Acad. Sci. USA 111, E4596–E4605 (2014).
Lee, W. L., Grimes, J. M. & Robinson, R. C. Yersinia effector YopO uses actin as bait to phosphorylate proteins that regulate actin polymerization. Nat. Struct. Mol. Biol. 22, 248–255 (2015).
Wang, H., Robinson, R. C. & Burtnick, L. D. The structure of native G-actin. Cytoskeleton (Hoboken) 67, 456–465 (2010).
Kabsch, W. Xds. Acta Crystallogr. D 66, 125–132 (2010).
Adams, P. D. et al. The Phenix software for automated determination of macromolecular structures. Methods 55, 94–106 (2011).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Pei, J. & Grishin, N. V. PROMALS3D: multiple protein sequence alignment enhanced with evolutionary and three-dimensional structural information. Methods Mol. Biol. 1079, 263–271 (2014).
Zhang, Y. I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9, 40 (2008).
Pagès, H. et al. DebRoyBiostrings: Efficient manipulation of biological strings. https://rdrr.io/bioc/Biostrings/ (2018).
Finn, R. D. et al. InterPro in 2017-beyond protein family and domain annotations. Nucleic Acids Res. 45, D190–D199 (2017).
Acknowledgements
We thank A*STAR for support; W. Burkholder for reagents; A. Kaya, M. Magnitov and E. Balıkçı for technical advice; and B. Venkatesh for valuable discussions. We thank the experimental facility and the technical services provided by: The Synchrotron Radiation Protein Crystallography Facility of the National Core Facility Program for Biotechnology, Ministry of Science and Technology and the National Synchrotron Radiation Research Center, a national user facility supported by the Ministry of Science and Technology, Taiwan; and the Australian Synchrotron, part of ANSTO.
Reviewer information
Nature thanks T. Ettema, T. Pollard and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
C.A. and R.C.R. conceived experiments, analysed data and wrote the paper. C.A. performed experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Asgard actins.
a, Phylogenetic tree of the polymerizing actin fold. This phylogenetic tree reveals that the variability observed between the Asgard and eukaryotic actins is approximately similar in magnitude to the variability found within bacterial MreBs and lower than that observed within bacterial FtsAs or ParMs, which indicates a probable conservation in function between the Asgard and eukaryotic actins. Nevertheless, Asgard and eukaryotic actins form two separate clusters, which indicates that the Asgard actin genes are unlikely to be contaminants from eukaryotes. The actin, MreB, FtsA and ParM sequences were aligned in PROMALS3D using accession codes for the actins shown in Supplementary Table 1, and the structures: 1JCE, 3WT0, 1MWM and actin 3HBT (PDB codes). The MreBs are ABO56305.1, ABC18867.1, ACB86382.1, AAC72350.1, ABC76501.1, AAO36573.1 and AAN82446.1; the FtsAs are AAN78610.1, AAO35699.1, ABC76564.1, AAF95541.1, ACB84890.1, ABC19166.1, and KVD45461.1; and the ParMs are ABO60264.1, ABC18679.1, ACB86228.1, BAC79052.1, ABC77529.1, AAO37409.1, and ABV16243.1 (GenBank codes). Heimdall actin, Loki actin, Odin actin and Thor actin are highly similar to eukaryotic actins, as judged by the branch points and identities (Supplementary Table 1a), and are here referred to as the ‘Asgard actins’. b, Models of the Asgard actins. The Asgard actins were modelled using I-TASSER. The ‘C-score’ is a confidence score for estimating the quality of predicted models, which is typically in the range of −5 to 2 and in which a high C-score signifies a model with a high confidence. Template modelling scores (‘TM-scores’) range between 0 and 1, and values greater than 0.5 indicate models of the correct topology and a value of 1 indicates an exact match. The TM-score and r.m.s.d. are estimated by linear regression, and the estimated errors are the root-mean-squared TM-score or r.m.s. deviations. The models of Heimdall actin, Loki actin, Odin actin and Thor actin are of high confidence and are notably similar to the structure of rabbit muscle actin, as judged by these metrics. Models of Loki ARP1 and Loki ARP2 are slightly less reliable and their sequences are more divergent (Supplementary Table 1a). The major differences between the Loki ARP1 and Loki ARP2 models and the structure of rabbit muscle actin lie in the DNAse I binding loop (D-loop), which is often disordered in G-actin structures (indicated by the oval) and becomes ordered on forming F-actin. This region has insertions in Loki ARP1 and Loki ARP2.
Extended Data Fig. 2 Titration data.
Titration data for the polymerization profiles that are shown in Fig. 2a–f.
Extended Data Fig. 3 Comparisons of polymerization profiles.
Polymerization profiles are shown in Fig. 2a–f. a, Enlargement of the lag phases and comparison between non-seeded and seeded actin polymerization are also shown as titrations for each profilin. b, Comparison of the inhibition of actin nucleation in the pyrene–actin assay of Asgard profilins, relative to human profilin-1. These profiles complement the Odin actin–human profilin-1 comparison shown in Fig. 2h. c, The quality of the purified Asgard profilin proteins. SDS–PAGE gels indicating the purity of the profilins (abbreviated as ‘P’ in this panel) used in this study. M indicates the protein molecular weight (MW) marker lane.
Extended Data Fig. 4 Comparison of profilin structures.
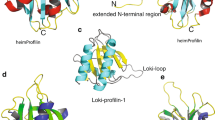
a, Ribbon representations of the profilin–actin complexes. The overlays were created by superimposing the actin structures to show the relative binding geometries of the profilins (abbreviated as P). For clarity, only one actin is displayed. The positions of Loki profilin-1 and Loki profilin-2 overlay well with the major interaction with subdomain 3. The rabbit α-actin subdomains are indicated as black numbers. Some flexibility is observed in the binding mode of Loki profilin-2, as seen by a slight shift between the two complexes in the asymmetric unit (labelled LokiP2A and LokiP2B). Human profilin-1 and Odin profilin adopt binding positions that also interact with subdomain 3, but which additionally have more-intimate contacts with rabbit α-actin subdomain 1. The human profilin-1–rabbit α-actin and Odin profilin–rabbit α-actin complexes are shown separately to aid in interpretation of the overlay. Bound calcium ions are shown as black spheres and ATP in stick representation. b, Latrunculin B (LatB, cyan) binds to analogous positions in rabbit α-actin in each complex. The Odin profilin–rabbit α-actin binds to a second molecule of latrunculin B at the edge of the Odin profilin–rabbit α-actin interface. For clarity, two views of Odin profilin–rabbit α-actin are shown. c–f, Cartoons of the profilin structures rotated by 180° around the y axis with respect to Fig. 1b, d, e, f. g–i, Cα traces of the superimpositions of human, Odin, Loki profilin-1 and Loki profilin-2 structures (g), Loki profilin-1 and Loki profilin-2 structures (h) and the human and Odin profilins (i) to highlight their similarities and differences. j–l, The 2Fo−Fc electron density contoured at 1.0σ—indicating the quality of the structures—is shown surrounding the Loki-loop regions of Loki profilin-1 (j) and Loki profilin-2 (k), along with the absence of the Loki-loop in this region in Odin profilin (l).
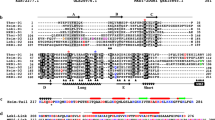
Extended Data Fig. 5 Structure-based sequence alignments.
a, Eukaryotic and Asgard profilins. The sequences were aligned using profilin structures (PDB codes: 2PAV, 1ACF, 3D9Y and 1A0K) and Loki and Odin profilin structures as guides. The Loki profilin-1 secondary structure is indicated below the alignment in green. Actin-binding sites are indicated by red stars (human profilin-1), blue stars (Loki profilin-1 and Loki profilin-2) and brown stars (Odin profilin). These appear in the same regions but are not well-conserved between the eukaryotes and archaea in amino acid sequences. Black arrows indicate polyproline-binding resides, which are conserved in the eukaryotic sequences but not in the archaea sequences. b, Eukaryotic and Asgard actins. The sequences were aligned using the native rabbit α-actin structure as a guide (PDB code: 3HBT). Profilin-binding sites observed in the structures are indicated by red (human profilin-1; PDB code: 2PAV), blue (Loki profilin-1 and Loki profilin-2) and cyan (Odin profilin) bars above the alignment. The profilins bind to regions of actin that display conservation between human cytoplasmic β-actin and the Asgard actins. Loki ARP1 is included here to demonstrate that other actin-like sequences are found in the Asgard metagenomes, and that these sequences share less sequence identity with human cytoplasmic β-actin. The red arrow indicates an alternative start methionine in the Odin actin sequence, and the Odin actin sequence displayed here takes account of a potential frame-shift in the nucleotide sequence (GenBank accession code: MDVT01000007).
Extended Data Fig. 6 Human profilin-1 binds to polyproline motifs, whereas Asgard profilins do not.
a, ITC binding profiles of full length VASP, which contains multiple polyproline motifs, titrated with human profilin-1. b, ITC binding profiles of a polyproline motif from VASP (PPPAPPLPAAQ) titrated with human profilin-1. c, ITC binding profiles of full-length VASP titrated with Loki profilin-1. d, ITC binding profiles of the single polyproline motif from VASP titrated with Loki profilin-1. e–i, Similar titrations for the single polyproline motif from VASP to the profiles shown in d, for Loki profilin-2 (e), Loki profilin-3 (f), Odin profilin (g), Heimdall profilin (h) and Thor profilin (i) are shown, each of which displays no observable interaction of the Asgard profilins with polyproline sequences. j–l, The 2Fo−Fc electron density, contoured at 1.0σ, surrounding the N- and C-termini of Loki profilin-1 (j), Loki profilin-2 (k) and Odin profilin (l), showing that the terminal helices are tightly packed and well-ordered.
Extended Data Fig. 7 Phospholipid binding.
a, Potential phospholipid-binding sites on the surface of Asgard profilins. Actin complexes are shown rotated by 180° around the y axis, relative to those shown in Fig. 4e–h. Expanded regions show that the profilins each present a basic residue that binds to the C-terminal residue of actin (Phe375) and to Tyr169, which in turn binds to a conserved Val or Ile on the profilins. Phe375 is conserved, whereas Tyr169 has a conservative substitution (Phe) in Loki actin and Odin actin sequences (Extended Data Fig. 5b). Arg117 and interacting residues on actin (Tyr169 and Phe375) adopt slightly different conformations in the Loki profilin-1–rabbit α-actin and Loki profilin-2–rabbit α-actin complexes. Lys60 (Fig. 4f, g) lies at the start of the Loki loop of Loki profilin-1 and Loki profilin-2. b, Pyrene–actin polymerization profiles of rabbit α-actin (2 μM, orange) supplemented with profilins and subsequently with increasing concentrations of a soluble version of PIP2 or inositol trisphosphate (IP3). In each case, PIP2 causes a reduction in the delay of polymerization caused by profilin inhibition of spontaneous actin nucleation, whereas IP3 has no effect. c, ITC was used to verify the interaction between selected profilins and PIP2. ITC binding profile of human profilin-1 titrated with soluble PIP2 demonstrated an interaction, whereas Loki profilin-1 titrated with soluble PIP2 showed marginal binding. Both profilins showed a ΔH = 0 on addition of IP3 in the ITC assay. The marginal binding observed for Loki profilin-1 with soluble PIP2 appears to be above the ΔH = 0 observed for IP3.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables 1-3. Supplementary Table 1: Pairwise percentage sequence identities of actins and profilins. a, Actins. b, Profilins. Supplementary Table 2: Data collection and refinement statistics. a, LokiProfilin1. b, Asgard profilin/rActin complexes. Supplementary Table 3: Structural similarities in profilins and homology between the LokiProfilin1/2 contigs. a, Comparison of profilin structures. b, Nucleotide percentage identities for genes on the LokiProfilin1/2 contigs.
Rights and permissions
About this article
Cite this article
Akıl, C., Robinson, R.C. Genomes of Asgard archaea encode profilins that regulate actin. Nature 562, 439–443 (2018). https://doi.org/10.1038/s41586-018-0548-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0548-6
Keywords
This article is cited by
-
The eukaryotic-like characteristics of small GTPase, roadblock and TRAPPC3 proteins from Asgard archaea
Communications Biology (2024)
-
Actin cytoskeleton and complex cell architecture in an Asgard archaeon
Nature (2023)
-
World War II, Sex and Antibiotics - II
Resonance (2023)
-
Inference and reconstruction of the heimdallarchaeial ancestry of eukaryotes
Nature (2023)
-
Conserved and lineage-specific hypothetical proteins may have played a central role in the rise and diversification of major archaeal groups
BMC Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.