Abstract
The putative Plasmodium translocon of exported proteins (PTEX) is essential for transport of malarial effector proteins across a parasite-encasing vacuolar membrane into host erythrocytes, but the mechanism of this process remains unknown. Here we show that PTEX is a bona fide translocon by determining structures of the PTEX core complex at near-atomic resolution using cryo-electron microscopy. We isolated the endogenous PTEX core complex containing EXP2, PTEX150 and HSP101 from Plasmodium falciparum in the ‘engaged’ and ‘resetting’ states of endogenous cargo translocation using epitope tags inserted using the CRISPR–Cas9 system. In the structures, EXP2 and PTEX150 interdigitate to form a static, funnel-shaped pseudo-seven-fold-symmetric protein-conducting channel spanning the vacuolar membrane. The spiral-shaped AAA+ HSP101 hexamer is tethered above this funnel, and undergoes pronounced compaction that allows three of six tyrosine-bearing pore loops lining the HSP101 channel to dissociate from the cargo, resetting the translocon for the next threading cycle. Our work reveals the mechanism of P. falciparum effector export, and will inform structure-based design of drugs targeting this unique translocon.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
World Health Organization. World Malaria Report 2017. (World Health Organization, Geneva, 2017).
Miller, L. H., Ackerman, H. C., Su, X. Z. & Wellems, T. E. Malaria biology and disease pathogenesis: insights for new treatments. Nat. Med. 19, 156–167 (2013).
Cowman, A. F., Healer, J., Marapana, D. & Marsh, K. Malaria: biology and disease. Cell 167, 610–624 (2016).
Spillman, N. J., Beck, J. R. & Goldberg, D. E. Protein export into malaria parasite-infected erythrocytes: mechanisms and functional consequences. Annu. Rev. Biochem. 84, 813–841 (2015).
Boddey, J. A. & Cowman, A. F. Plasmodium nesting: remaking the erythrocyte from the inside out. Annu. Rev. Microbiol. 67, 243–269 (2013).
Przyborski, J. M., Nyboer, B. & Lanzer, M. Ticket to ride: export of proteins to the Plasmodium falciparum-infected erythrocyte. Mol. Microbiol. 101, 1–11 (2016).
de Koning-Ward, T. F., Dixon, M. W., Tilley, L. & Gilson, P. R. Plasmodium species: master renovators of their host cells. Nat. Rev. Microbiol. 14, 494–507 (2016).
Marti, M., Good, R. T., Rug, M., Knuepfer, E. & Cowman, A. F. Targeting malaria virulence and remodeling proteins to the host erythrocyte. Science 306, 1930–1933 (2004).
Hiller, N. L. et al. A host-targeting signal in virulence proteins reveals a secretome in malarial infection. Science 306, 1934–1937 (2004).
Heiber, A. et al. Identification of new PNEPs indicates a substantial non-PEXEL exportome and underpins common features in Plasmodium falciparum protein export. PLoS Pathog. 9, e1003546 (2013).
Lingelbach, K. & Joiner, K. A. The parasitophorous vacuole membrane surrounding Plasmodium and Toxoplasma: an unusual compartment in infected cells. J. Cell Sci. 111, 1467–1475 (1998).
Ansorge, I., Benting, J., Bhakdi, S. & Lingelbach, K. Protein sorting in Plasmodium falciparum-infected red blood cells permeabilized with the pore-forming protein streptolysin O. Biochem. J. 315, 307–314 (1996).
Gehde, N. et al. Protein unfolding is an essential requirement for transport across the parasitophorous vacuolar membrane of Plasmodium falciparum. Mol. Microbiol. 71, 613–628 (2009).
de Koning-Ward, T. F. et al. A newly discovered protein export machine in malaria parasites. Nature 459, 945–949 (2009).
Beck, J. R., Muralidharan, V., Oksman, A. & Goldberg, D. E. PTEX component HSP101 mediates export of diverse malaria effectors into host erythrocytes. Nature 511, 592–595 (2014).
Elsworth, B. et al. PTEX is an essential nexus for protein export in malaria parasites. Nature 511, 587–591 (2014).
Bullen, H. E. et al. Biosynthesis, localization, and macromolecular arrangement of the Plasmodium falciparum translocon of exported proteins (PTEX). J. Biol. Chem. 287, 7871–7884 (2012).
Johnson, D. et al. Characterization of membrane proteins exported from Plasmodium falciparum into the host erythrocyte. Parasitology 109, 1–9 (1994).
Gold, D. A. et al. The Toxoplasma dense granule proteins GRA17 and GRA23 mediate the movement of small molecules between the host and the parasitophorous vacuole. Cell Host Microbe 17, 642–652 (2015).
Garten, M. et al. R. EXP2 is a nutrient-permeable channel in the vacuolar membrane of Plasmodium and is essential for protein export via PTEX. Nat. Microbiol. https://doi.org/10.1038/s41564-018-0222-7 (2018).
Chisholm, S. A. et al. The malaria PTEX component PTEX88 interacts most closely with HSP101 at the host-parasite interface. FEBS J. 285, 2037–2055 (2018).
Holm, L. & Laakso, L. M. Dali server update. Nucleic Acids Res. 44, W351–W355 (2016).
Gibrat, J. F., Madej, T. & Bryant, S. H. Surprising similarities in structure comparison. Curr. Opin. Struct. Biol. 6, 377–385 (1996).
Wiederstein, M., Gruber, M., Frank, K., Melo, F. & Sippl, M. J. Structure-based characterization of multiprotein complexes. Structure 22, 1063–1070 (2014).
Krissinel, E. & Henrick, K. Secondary-structure matching (SSM), a new tool for fast protein structure alignment in three dimensions. Acta Crystallogr. D Biol. Crystallogr. 60, 2256–2268 (2004).
Dosztanyi, Z., Csizmok, V., Tompa, P. & Simon, I. IUPred: web server for the prediction of intrinsically unstructured regions of proteins based on estimated energy content. Bioinformatics 21, 3433–3434 (2005).
Dosztanyi, Z., Csizmok, V., Tompa, P. & Simon, I. The pairwise energy content estimated from amino acid composition discriminates between folded and intrinsically unstructured proteins. J. Mol. Biol. 347, 827–839 (2005).
Schirmer, E. C., Glover, J. R., Singer, M. A. & Lindquist, S. HSP100/Clp proteins: A common mechanism explains diverse functions. Trends Biochem. Sci. 21, 289–296 (1996).
Hanson, P. I. & Whiteheart, S. W. AAA+ proteins: have engine, will work. Nat. Rev. Mol. Cell Biol. 6, 519–529 (2005).
Deville, C. et al. Structural pathway of regulated substrate transfer and threading through an Hsp100 disaggregase. Sci. Adv. 3, e1701726 (2017).
Gates, S. N. et al. Ratchet-like polypeptide translocation mechanism of the AAA+ disaggregase Hsp104. Science 357, 273–279 (2017).
AhYoung. A. P., Koehl, A., Cascio, D. & Egea, P. F. Structural mapping of the ClpB ATPases of Plasmodium falciparum: targeting protein folding and secretion for antimalarial drug design. Protein Sci. 24, 1508–1520 (2015).
Puchades, C. et al. Structure of the mitochondrial inner membrane AAA+ protease YME1 gives insight into substrate processing. Science 358, (2017).
Yokom, A. L. et al. Spiral architecture of the Hsp104 disaggregase reveals the basis for polypeptide translocation. Nat. Struct. Mol. Biol. 23, 830–837 (2016).
Seyffer, F. et al. Hsp70 proteins bind Hsp100 regulatory M domains to activate AAA+ disaggregase at aggregate surfaces. Nat. Struct. Mol. Biol. 19, 1347–1355 (2012).
Lipinska, N. et al. Disruption of ionic interactions between the nucleotide binding domain 1 (NBD1) and middle (M) domain in Hsp100 disaggregase unleashes toxic hyperactivity and partial independence from Hsp70. J. Biol. Chem. 288, 2857–2869 (2013).
Elsworth, B. et al. Proteomic analysis reveals novel proteins associated with the Plasmodium protein exporter PTEX and a loss of complex stability upon truncation of the core PTEX component, PTEX150. Cell. Microbiol. 18, 1551–1569 (2016).
Kulzer, S. et al. Plasmodium falciparum-encoded exported hsp70–hsp40 chaperonen–co-chaperone complexes within the host erythrocyte. Cell. Microbiol. 14, 1784–1795 (2012).
Batinovic, S. et al. An exported protein-interacting complex involved in the trafficking of virulence determinants in Plasmodium-infected erythrocytes. Nat. Commun. 8, 16044 (2017).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. P. HOLE: A program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360 (1996).
Adjalley, S. H. et al. Quantitative assessment of Plasmodium falciparum sexual development reveals potent transmission-blocking activity by methylene blue. Proc. Natl Acad. Sci. USA 108, E1214–E1223 (2011).
Klemba, M., Beatty, W., Gluzman, I. & Goldberg, D. E. Trafficking of plasmepsin II to the food vacuole of the malaria parasite Plasmodium falciparum. J. Cell Biol. 164, 47–56 (2004).
Spillman, N. J., Beck, J. R., Ganesan, S. M., Niles, J. C. & Goldberg, D. E. The chaperonin TRiC forms an oligomeric complex in the malaria parasite cytosol. Cell Microbiol. 19, e12719 (2017).
Scheres, S. H. W. A Bayesian view on cryo-EM structure determination. J. Mol. Biol. 415, 406–418 (2012).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2015).
Pettersen, E. F. et al. UCSF chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
NCBI Resource Coordinators. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 44, D7–D19 (2016).
Aurrecoechea, C. et al. PlasmoDB: a functional genomic database for malaria parasites. Nucleic Acids Res. 37, D539–D543 (2009).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. E. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EMEM density maps. Nat. Methods 11, 63–65 (2014).
Ganesan, S. M. et al. Yeast dihydroorotate dehydrogenase as a new selectable marker for Plasmodium falciparum transfection. Mol. Biochem. Parasitol. 177, 29–34 (2011).
Blisnick, T. et al. Pfsbp 1, a Maurer’s cleft Plasmodium falciparum protein, is associated with the erythrocyte skeleton. Mol. Biochem. Parasitol. 111, 107–121 (2000).
Puchades, C. et al. Structure of the mitochondrial inner membrane AAA+ protease YME1 gives insight into substrate processing. Science 358, eaao0464 (2017).
Acknowledgements
This research was supported in part by grants from National Institutes of Health (R21AI125983 to P.F.E., R01GM071940/AI094386/DE025567 to Z.H.Z. and K99/R00 HL133453 to J.R.B.). P.F.E. is the Alexander and Renee Kolin Endowed Chair in Molecular Biology and Biophysics. C.-M.H. acknowledges funding from the Ruth L. Kirschstein National Research Service Award (AI007323). We thank the UCLA Proteome Research Center for assistance in mass spectrometry and acknowledge the use of instruments in the Electron Imaging Center for Nanomachines supported by UCLA and grants from NIH (S10RR23057, S10OD018111 and U24GM116792) and NSF (DBI-1338135 and DMR-1548924). We thank A. W. P. Fitzpatrick for input on cryo-EM aspects of the project and J. Su for helping with Fig. 1a.
Reviewer information
Nature thanks A. Cowman, P. da Fonseca and W. Hol for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
C.-M.H., P.F.E. and Z.H.Z. initiated the project; J.R.B. generated P. falciparum lines, collected P. falciparum, performed complementation experiments and helped to write the paper; C.-M.H. purified the sample, screened purified samples by negative stain, prepared cryo-EM grids, acquired and processed the cryo-EM data, interpreted the structures and wrote the paper; M.L. built and refined the atomic models and helped to interpret the structures; Y.C. helped with sample freezing; Z.H.Z. supervised the cryo-EM aspects of the project, interpreted the structures and wrote the paper; P.F.E. supervised biochemical aspects of the project and helped to interpret the structures; D.E.G. supervised parasitology aspects of the project. D.E.G., M.L. and P.F.E. helped to edit the paper. Queries regarding cryo-EM and atomic modelling should be addressed to Z.H.Z. and queries on biochemistry should be addressed to P.F.E.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
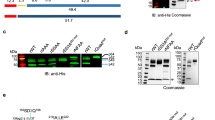
Extended Data Fig. 1 Generation of HSP101–3×Flag P. falciparum and analysis of purified PTEX.
a, Schematic showing strategy for endogenous tagging of P. falciparum hsp101 with 3×Flag using CRISPR–Cas9 editing. Diagnostic PCR primers and expected amplicon following successful integration are shown. sgRNA, single-guide RNA; UTR, untranslated region; CAM, calmodulin promoter; PfU6, P. falciparum U6 promoter; hDHFR, human dihydrofolate reductase. b, Diagnostic PCR with genomic DNA template from NF54attB parent or two independent populations of HSP101–3×Flag P. falciparum. The experiment was performed once. c, Western blot of NF54attB and HSP101–3×Flag P. falciparum probed with mouse Flag M2 antibody (Sigma) and goat anti-mouse IRDye 680 secondary antibody (Li-Cor). Arrowhead indicates full-length HSP101–3×Flag (predicted molecular weight 102.9 kDa after signal peptide cleavage). Data represent two independent experiments. d, Giemsa staining of parasite-infected human erythrocytes from which PTEX was purified. Scale bar, 5 μm. For source data, see Supplementary Fig. 3. e, Silver-stained SDS–PAGE of the Flag-purified PTEX sample. Identities of the bands labelled EXP2, PTEX150 and HSP101 were confirmed by tryptic digest liquid chromatography–mass spectrometry (LC–MS). f, Tryptic digest LC–MS analysis of the Flag-purified PTEX sample. The PTEX core components are among the five most abundant species detected in the purified sample. For gel and blot source data, see Supplementary Fig. 1.
Extended Data Fig. 2 Detailed views of the PTEX protein-conducting channel and symmetry mismatch in the engaged state.
a–c, Cryo-EM densities and atomic models of cargo and pore loops from the near-atomic resolution structures of Clp/HSP100 ATPases YME159 (a), PTEX HSP101 (b) and HSP10431 (c). Tyrosine sidechain densities are clearly visibly intercalating with the cargo densities. The modelled engaged state PTEX cargo has a calculated r.m.s.d. of 1.09 Å and 1.25 Å to the published YME1 and HSP104 cargo models, respectively. Pore loops are labelled by NBD and protomer (for example, D2PL,P1 is NBD2 pore loop, protomer 1). d, Side view of the bisected engaged state PTEX cryo-EM map. The protein-conducting channel, calculated using HOLE40, is shown superimposed over the bisected map in translucent white with a navy outline. The HSP101 NBD2 pore loop densities are coloured by HSP101 protomer, and the cargo density is coloured pink. e–j, The transition from the asymmetric HSP101 spiral to the C7-pseudosymmetric PTEX150(668–823)–EXP2 heptamer is depicted using a series of cross-sections taken perpendicular to the central axis of the translocon, spanning the area of symmetry mismatch. The section of the translocon corresponding to each cross-sectional image is indicated with a bracket in d.
Extended Data Fig. 3 Detailed comparisons of the engaged and resetting states.
a, Side and top views of the EXP2 heptamer in the engaged state. Symmetric portions that remain constant between protomers are coloured in mint. Portions that vary between protomers are coloured and labelled by protomer. b, c, Superposition of the seven EXP2 protomers, labelled A–G, in the engaged (b) and resetting (c) states, coloured as in a. d, e, Top view of HSP101 NBD1 (d) and NBD2 (e) in the engaged and resetting states, shown in simplified surface representation. The hinge point at the interface between HSP101 protomers 3 and 4 is indicated. f, g, Ribbon diagrams of the resetting state (f) and engaged state (g) nucleotide binding pockets are shown for each protomer. ATPγS in each pocket is shown with corresponding cryo-EM density (mesh). The R859 arginine finger (sidechain shown in red-orange) is positioned approximately 3–5.5 Å from the phosphorus atom in the γ-phosphate of the ATPγS in the binding pocket of the neighbouring protomer in all protomers except R859 in protomer 3 in the resetting state (sidechain shown in gold), where the ATPγS bound in the protomer 4 NBD2 nucleotide pocket has shifted approximately 7.5 Å away from the protomer 3 R859 arginine finger. h, i, Enlarged side view of the atomic models of the HSP101 NBD2 pore loops and unfolded cargo polypeptide backbone in the engaged (h) and resetting (i) states, shown with corresponding cryo-EM densities. Tyrosine sidechain densities are clearly visibly intercalating with the cargo densities. The modelled PTEX cargo has a calculated r.m.s.d. of 1.09 Å and 1.25 Å to the published YME1 and HSP104 cargo models, respectively. Pore loops are labelled by NBD and protomer (for example, D2PL,P1 is NBD2 pore loop, protomer 1).
Extended Data Fig. 4 Resolution assessments of the two PTEX states.
a–c, e, f, Local resolution evaluations of the full PTEX map (a) and the focus-refined maps of the upper/hexameric (b) and lower/heptameric (c) halves of PTEX in the engaged state, the full PTEX map (e) and the focus-refined map of the lower/heptameric (f) half of PTEX in the resetting state, calculated by Resmap56 and coloured according to resolution. Maps are displayed at higher thresholds where the detergent belt is not visible for clarity, to avoid obscuring details of the transmembrane helices. d, Global resolution assessment of the engaged and resetting state maps as measured using the ‘Gold-standard’ Fourier shell correlation (FSC) curves generated by RELION44,45 by comparison of two independently refined ‘half-maps’. g, Map-to-model FSC curves demonstrating the degree of correlation between the refined PTEX models and the experimental cryo-EM maps for the engaged and resetting states.
Extended Data Fig. 5 Representative regions of cryo-EM density and atomic models.
Additional cryo-EM densities (mesh) superposed with our atomic models for HSP101 (a), PTEX150(668–823) (b) and EXP2 (c). Displayed regions correspond to areas circled in magenta on guide figures (inset, upper right), and are coloured as in the guide figure: HSP101 (cornflower blue), PTEX150(668-823) (salmon), EXP2 (mint). Terminal residues for each segment are labelled with the amino acid, residue number, protomer, and state.
Extended Data Fig. 6 Experimentally determined secondary structure elements and detected mass-spectrometry fragments mapped to the primary sequences of the three PTEX proteins.
For EXP2 (a), PTEX150 (b) and HSP101 (c), secondary structure elements are shown as tubes (helices), lines (loops), and arrows (strands) above the corresponding sequence and are coloured as in Figs. 2a, 3a, 4a. In the sequences shown below, residues resolved in our structures are coloured according to protein colours in Fig. 1c–f: EXP2 (mint), PTEX150 (salmon) and HSP101 (cornflower). Signal peptide residues are coloured gold. All residues in the mature proteins that are not resolved in our structures are shown in grey. The 3×Flag residues at the C terminus of HSP101 are coloured green. Peptides detected in tryptic digest liquid chromatography–tandem mass spectrometry (LC–MS/MS) analysis of the purified PTEX sample are shown as black lines below the corresponding sequences. Arrowheads above the EXP2 sequence indicate truncations sites described in this work and previously20 immediately before (∆222–287, red arrowhead) and after (∆234–287, green arrowhead) the assembly strand. Arrowheads above the PTEX150 sequence indicate previously described truncation sites37 (∆847–993, red arrowhead; ∆869–993, green arrowhead).
Extended Data Fig. 7 Electron microscopy of the PTEX core complex.
a–c, Representative negative stain micrograph (a), enlarged portion of micrograph (b) and two-dimensional class averages (c) of the PTEX core complex in multiple orientations. d, e, Representative cryo-EM micrograph (d) and two-dimensional class averages (e) of the PTEX core complex in multiple orientations. Individual particles in d are circled in yellow (top views) and green (side and oblique views) (for source data, see Supplementary Fig. 4). Arrow in upper left panel of e indicates the detergent belt, which is visible as a less-dense (dimmer) halo surrounding the denser (brighter) densities of the α-helices visible in the transmembrane domain in side views. Scale bars: 700 Å (a), 700 Å (b), 100 Å (c), 200 Å (d) and 100 Å (e).
Extended Data Fig. 8 Detergent belt, amino-terminal domain, and claw densities visible in maps at lower thresholds.
a, The engaged state PTEX150–EXP2 heptamer, displayed in surface representation and coloured by electrostatic potential. The bottom half of the full engaged state density map is superimposed, showing the location of the detergent belt in relation to the EXP2 transmembrane domain. A ring of positively charged residues is clearly visible directly above where the PVM surface would normally lie. b, c, Engaged state (b) and resetting state (c) maps were low-pass filtered to 6 Å to improve clarity of low-resolution details, and are shown overlaid, at two different thresholds to improve visibility of the detergent belt and the poorly resolved N-terminal domains of HSP101 (teal, higher threshold; peach, lower threshold). d, Resetting state map of PTEX displayed at a lower threshold to show the strong claw-shaped densities extending from the PTEX150(668–823) shaft up to the HSP101 M domain. e, Enlarged view of the interaction between HSP101 Y488 and Y491 and the three-turn helix, shown with corresponding cryo-EM density (mesh).
Extended Data Fig. 9 Data processing workflow.
Illustration of workflow for 3D classification, focused classification and refinement. Maps are displayed at higher thresholds where the detergent belt is not visible for clarity, to avoid obscuring details of the transmembrane helices.
Supplementary information
Supplementary Figures
This file contains Supplementary Figs. 1–4. Supplementary Fig. 1: Full length western blots, Supplementary Fig. 2: Flow cytometry gating strategy, Supplementary Fig. 3: Giemsa-staining of P. falcipaurm infected human erythrocytes and Supplementary Fig. 4: original representative cryoEM micrograph of vitrified PTEX particles in various orientations.
Video 1
Details of the PTEX cryoEM Density Map. 360° view of the density map (mesh) and models of the PTEX complex in the engaged state. Enlarged views of the atomic models of EXP2 (mint), PTEX150 (salmon), and HSP101 (cornflower blue).
Video 2
Details of the PTEX Core Components. Detailed views of the cryoEM density and atomic models of the individual PTEX core components in the engaged state.
Video 3
PTEX Stoichiometry and Architecture. 360° view of the engaged state PTEX complex cryoEM map, colored by protomer, illustrating the stoichiometry and global organization of the three core components, EXP2, PTEX150, and HSP101.
Video 4
Flythrough of the PTEX channel. 360° view of density map and ribbon models of the PTEX complex in the engaged state, followed by a flythrough of the channel entering via HSP101, with focus on the NBD2 tyrosine staircase and cargo peptide (yellow).
Video 5
HSP101 Pore Loops and Endogenous Cargo. Video highlighting the details of the HSP101 tyrosine pore loops and endogenous cargo in the cryoEM map of PTEX in the engaged state.
Video 6
PTEX Engaged and Resetting States Morph. Morph between the engaged and resetting states, calculated using the “morph conformations” function in UCSF Chimera. This morph is not representative of the exact motion of the mechanism, but is intended to give a sense of scale for the dramatic conformational shift.
Video 7
PTEX NBD Tyrosine Loops Engaged and Resetting States Morph. Morph of the tyrosine loops of NBD1 (gold) and NBD2 (cyan) between the engaged and resetting states, calculated using the “morph conformations” function in UCSF Chimera. Note the collapsing of the NBD2 tyrosine loops from a staircase to planar “C” shape. This morph is not representative of the exact motion of the mechanism, but is intended to give a sense of scale for the dramatic conformational shift.
Rights and permissions
About this article
Cite this article
Ho, CM., Beck, J.R., Lai, M. et al. Malaria parasite translocon structure and mechanism of effector export. Nature 561, 70–75 (2018). https://doi.org/10.1038/s41586-018-0469-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0469-4
Keywords
This article is cited by
-
Toxoplasma protein export and effector function
Nature Microbiology (2024)
-
Structure, dynamics and assembly of the ankyrin complex on human red blood cell membrane
Nature Structural & Molecular Biology (2022)
-
A ‘Build and Retrieve’ methodology to simultaneously solve cryo-EM structures of membrane proteins
Nature Methods (2021)
-
A single-cell atlas of Plasmodium falciparum transmission through the mosquito
Nature Communications (2021)
-
Composition and stage dynamics of mitochondrial complexes in Plasmodium falciparum
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.