Abstract
The folding of newly synthesized proteins to the native state is a major challenge within the crowded cellular environment, as non-productive interactions can lead to misfolding, aggregation and degradation1. Cells cope with this challenge by coupling synthesis with polypeptide folding and by using molecular chaperones to safeguard folding cotranslationally2. However, although most of the cellular proteome forms oligomeric assemblies3, little is known about the final step of folding: the assembly of polypeptides into complexes. In prokaryotes, a proof-of-concept study showed that the assembly of heterodimeric luciferase is an organized cotranslational process that is facilitated by spatially confined translation of the subunits encoded on a polycistronic mRNA4. In eukaryotes, however, fundamental differences—such as the rarity of polycistronic mRNAs and different chaperone constellations—raise the question of whether assembly is also coordinated with translation. Here we provide a systematic and mechanistic analysis of the assembly of protein complexes in eukaryotes using ribosome profiling. We determined the in vivo interactions of the nascent subunits from twelve hetero-oligomeric protein complexes of Saccharomyces cerevisiae at near-residue resolution. We find nine complexes assemble cotranslationally; the three complexes that do not show cotranslational interactions are regulated by dedicated assembly chaperones5,6,7. Cotranslational assembly often occurs uni-directionally, with one fully synthesized subunit engaging its nascent partner subunit, thereby counteracting its propensity for aggregation. The onset of cotranslational subunit association coincides directly with the full exposure of the nascent interaction domain at the ribosomal tunnel exit. The action of the ribosome-associated Hsp70 chaperone Ssb8 is coordinated with assembly. Ssb transiently engages partially synthesized interaction domains and then dissociates before the onset of partner subunit association, presumably to prevent premature assembly interactions. Our study shows that cotranslational subunit association is a prevalent mechanism for the assembly of hetero-oligomers in yeast and indicates that translation, folding and the assembly of protein complexes are integrated processes in eukaryotes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Balchin, D., Hayer-Hartl, M. & Hartl, F. U. In vivo aspects of protein folding and quality control. Science 353, aac4354 (2016).
Gloge, F., Becker, A. H., Kramer, G. & Bukau, B. Co-translational mechanisms of protein maturation. Curr. Opin. Struct. Biol. 24, 24–33 (2014).
Benschop, J. J. et al. A consensus of core protein complex compositions for Saccharomyces cerevisiae. Mol. Cell 38, 916–928 (2010).
Shieh, Y. W. et al. Operon structure and cotranslational subunit association direct protein assembly in bacteria. Science 350, 678–680 (2015).
Le Tallec, B. et al. 20S proteasome assembly is orchestrated by two distinct pairs of chaperones in yeast and in mammals. Mol. Cell 27, 660–674 (2007).
Smardon, A. M., Tarsio, M. & Kane, P. M. The RAVE complex is essential for stable assembly of the yeast V-ATPase. J. Biol. Chem. 277, 13831–13839 (2002).
Meurisse, J. et al. Hug1 is an intrinsically disordered protein that inhibits ribonucleotide reductase activity by directly binding Rnr2 subunit. Nucleic Acids Res. 42, 13174–13185 (2014).
Pfund, C., Huang, P., Lopez-Hoyo, N. & Craig, E. A. Divergent functional properties of the ribosome-associated molecular chaperone Ssb compared with other Hsp70s. Mol. Biol. Cell 12, 3773–3782 (2001).
Becker, A. H., Oh, E., Weissman, J. S., Kramer, G. & Bukau, B. Selective ribosome profiling as a tool for studying the interaction of chaperones and targeting factors with nascent polypeptide chains and ribosomes. Nat. Protocols 8, 2212–2239 (2013).
Doring, K. et al. Profiling Ssb–nascent chain interactions reveals principles of Hsp70-assisted folding. Cell 170, 298–311.e220 (2017).
Leibundgut, M., Jenni, S., Frick, C. & Ban, N. Structural basis for substrate delivery by acyl carrier protein in the yeast fatty acid synthase. Science 316, 288–290 (2007).
Gipson, P. et al. Direct structural insight into the substrate-shuttling mechanism of yeast fatty acid synthase by electron cryomicroscopy. Proc. Natl Acad. Sci. USA 107, 9164–9169 (2010).
Hansen, W. J., Lingappa, V. R. & Welch, W. J. Complex environment of nascent polypeptide chains. J. Biol. Chem. 269, 26610–26613 (1994).
Cherkasov, V. et al. Systemic control of protein synthesis through sequestration of translation and ribosome biogenesis factors during severe heat stress. FEBS Lett. 589, 3654–3664 (2015).
Duttler, S., Pechmann, S. & Frydman, J. Principles of cotranslational ubiquitination and quality control at the ribosome. Mol. Cell 50, 379–393 (2013).
Schüller, H. J., Förtsch, B., Rautenstrauss, B., Wolf, D. H. & Schweizer, E. Differential proteolytic sensitivity of yeast fatty acid synthetase subunits α and β contributing to a balanced ratio of both fatty acid synthetase components. Eur. J. Biochem. 203, 607–614 (1992).
Scazzari, M., Amm, I. & Wolf, D. H. Quality control of a cytoplasmic protein complex: chaperone motors and the ubiquitin-proteasome system govern the fate of orphan fatty acid synthase subunit Fas2 of yeast. J. Biol. Chem. 290, 4677–4687 (2015).
Karanasios, E., Simader, H., Panayotou, G., Suck, D. & Simos, G. Molecular determinants of the yeast Arc1p–aminoacyl-tRNA synthetase complex assembly. J. Mol. Biol. 374, 1077–1090 (2007).
Koehler, C., Round, A., Simader, H., Suck, D. & Svergun, D. Quaternary structure of the yeast Arc1p–aminoacyl-tRNA synthetase complex in solution and its compaction upon binding of tRNAs. Nucleic Acids Res. 41, 667–676 (2013).
Simader, H. et al. Structural basis of yeast aminoacyl-tRNA synthetase complex formation revealed by crystal structures of two binary sub-complexes. Nucleic Acids Res. 34, 3968–3979 (2006).
Frechin, M. et al. Expression of nuclear and mitochondrial genes encoding ATP synthase is synchronized by disassembly of a multisynthetase complex. Mol. Cell 56, 763–776 (2014).
Hannig, E. M., Cigan, A. M., Freeman, B. A. & Kinzy, T. G. GCD11, a negative regulator of GCN4 expression, encodes the gamma subunit of eIF-2 in Saccharomyces cerevisiae. Mol. Cell. Biol. 13, 506–520 (1993).
Polevoda, B., Cardillo, T. S., Doyle, T. C., Bedi, G. S. & Sherman, F. Nat3p and Mdm20p are required for function of yeast NatB Nα-terminal acetyltransferase and of actin and tropomyosin. J. Biol. Chem. 278, 30686–30697 (2003).
Bhushan, S. et al. α-Helical nascent polypeptide chains visualized within distinct regions of the ribosomal exit tunnel. Nat. Struct. Mol. Biol. 17, 313–317 (2010).
Gautschi, M., Mun, A., Ross, S. & Rospert, S. A functional chaperone triad on the yeast ribosome. Proc. Natl Acad. Sci. USA 99, 4209–4214 (2002).
Gumiero, A. et al. Interaction of the cotranslational Hsp70 Ssb with ribosomal proteins and rRNA depends on its lid domain. Nat. Commun. 7, 13563 (2016).
Willmund, F. et al. The cotranslational function of ribosome-associated Hsp70 in eukaryotic protein homeostasis. Cell 152, 196–209 (2013).
Gavin, A. C. et al. Proteome survey reveals modularity of the yeast cell machinery. Nature 440, 631–636 (2006).
Duncan, C. D. & Mata, J. Widespread cotranslational formation of protein complexes. PLoS Genet. 7, e1002398 (2011).
Halbach, A. et al. Cotranslational assembly of the yeast SET1C histone methyltransferase complex. EMBO J. 28, 2959–2970 (2009).
Sakahira, H. & Nagata, S. Co-translational folding of caspase-activated DNase with Hsp70, Hsp40, and inhibitor of caspase-activated DNase. J. Biol. Chem. 277, 3364–3370 (2002).
Janke, C. et al. A versatile toolbox for PCR-based tagging of yeast genes: new fluorescent proteins, more markers and promoter substitution cassettes. Yeast 21, 947–962 (2004).
Esmaielbeiki, R., Krawczyk, K., Knapp, B., Nebel, J. C. & Deane, C. M. Progress and challenges in predicting protein interfaces. Brief. Bioinform. 17, 117–131 (2016).
Cazals, F., Proust, F., Bahadur, R. P. & Janin, J. Revisiting the Voronoi description of protein–protein interfaces. Protein Sci. 15, 2082–2092 (2006).
Janin, J. et al. CAPRI: a critical assessment of predicted interactions. Proteins 52, 2–9 (2003).
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
Haslberger, T. et al. Protein disaggregation by the AAA+ chaperone ClpB involves partial threading of looped polypeptide segments. Nat. Struct. Mol. Biol. 15, 641–650 (2008).
Herbert, A. D., Carr, A. M. & Hoffmann, E. FindFoci: a focus detection algorithm with automated parameter training that closely matches human assignments, reduces human inconsistencies and increases speed of analysis. PLoS ONE 9, e114749 (2014).
Acknowledgements
We thank members of the Bukau laboratory for discussions, D. H. Wolf for FAS antisera and the DKFZ Core facility for sequencing. This work was supported by ERC Advanced grant (743118), SFB1036 and an Alexander von Humboldt fellowship to A.S.
Reviewer information
Nature thanks J. Marsh and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
A.S., G.K. and B.B. conceived the study and designed the experiments. A.S., K.D., U.F., K.K., D.M. and M.Z. performed the experiments. A.S., K.D., U.F., K.K., D.M., M.Z., F.T., G.K. and B.B. analysed the data. A.S., G.K. and B.B. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Functionality of GFP-tagged FAS complex subunits, characteristics of co- versus post-translational FAS subunit interactions and the FAS assembly model.
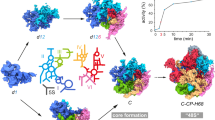
a, GFP tagging of the FAS complex subunits does not affect growth under fatty acid depletion conditions, as compared to wild-type (YPD, right compared to YPD + fatty acids, left). A representative image from three biologically independent experiments is shown. b, Immunoblotting of the FAS complex subunits in input, flow through and immunopurifiation fractions of a typical SeRP experiment analysing samples of strains encoding either GFP-tagged α or β subunits. Data are from three biologically independent experiments. c, Puromycin-induced release of nascent chains (10 μg/ml, 10 min post lysis) decreases the interaction of nascent α with the C-terminally tagged β subunit, analysed by immunopurification followed by RT–qPCR. Data are normalized mean mRNA levels ± s.e.m. with each data point from three biologically independent experiments overlaid. d, Polysome profiles of samples following puromycin (puro) treatment (as in c) or CHX treatment. Data representative of three biologically independent experiments are shown. e, Post-lysis binding control: experimental scheme. Two independent cultures, of two strains, expressing either wild-type α subunit and C-terminally GFP- tagged β subunit; or wild-type β subunit and C-terminally TAP-tagged α subunit, were grown to log phase, OD600 nm 0.5. The cells were then mixed in a 1:1 ratio and subsequently lysed, subjected to GFP immunopurification and SeRP. f, Predicted SeRP engagement of nascent wild-type α subunit or α–TAP ORF, by C-terminally GFP-tagged β subunit. No post-lysis interactions: no detection of ribosome protected footprints of mRNA encoding TAP (top). Post-lysis interactions: detection of ribosome protected footprints of TAP-encoding mRNA at a similar level to wild-type α subunit ORF (bottom). g, Results of post-lysis binding control: engagement of nascent wild-type α subunit or α–TAP by C-terminally GFP-tagged β subunit, analysed by SeRP, as in Fig. 1. Data are from two biologically independent experiments. h, Model of the FAS complex assembly pathway.
Extended Data Fig. 2 Functionality of GFP-tagged multi-aminoacyl-tRNA synthetase complex subunits and the assembly model.
a, GFP tagging of the essential multi-aminoacyl-tRNA synthetase complex subunits does not affect growth, as compared to wild type (YPD). A representative image from three biologically independent experiments is shown. b, Model of the multi-aminoacyl-tRNA synthetase complex assembly pathways.
Extended Data Fig. 3 Cotranslational assembly of the anthranilate synthase complex.
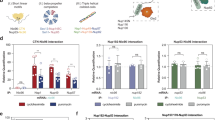
a, Domain organization of the anthranilate synthase subunits. b, Engagement of nascent Trp2p (tryptophan 2) and Trp3p (tryptophan 3) by C-terminally-tagged Trp2p subunit (top) compared to engagement of nascent Trp2p and Trp3p by C-terminally-tagged Trp3p subunit (bottom), analysed by SeRP. Data are from two biologically independent experiments. Coloured numbers indicate ribosome positions where the enrichment stably crosses the twofold threshold. The area between replicates is shaded, indicating the degree of experimental variation. c, Crystal structure of the homologous anthranilate synthase complex from the archaea Sulfolobus solfataricus (~60% sequence similarity, PDB: 1QDL1). d, GFP tagging of the complex subunits does not affect cell growth under tryptophan depletion conditions (YPD, right panel compared to SD lacking tryptophan, left). A representative image from three biologically independent experiments is shown. e, Model of the anthranilate synthase assembly pathway.
Extended Data Fig. 4 Cotranslational assembly of the phosphofructokinase complex.
a, Domain organization of the phosphofructokinase (PFK) subunits. b, Engagement of nascent α and β subunits by C-terminally tagged α subunit (top) compared to engagement of nascent α and β by C-terminally tagged β subunit (bottom), analysed by SeRP. Data are from two biologically independent experiments. Coloured numbers indicate ribosome positions when the enrichment stably crosses the twofold threshold. The area between replicates is shaded, indicating the degree of experimental variation. c, Top, crystal structure of the S. cerevisiae PFK complex (PDB: 3O8O2). Bottom, crystal structure of the highly homologous (~75% sequence similarities) Pichia pastoris (also known as Komagataella pastoris) PFK complex, (PDB: 3OPY3). The N′-terminal glyoxalase I-like interface domains of α and β are outlined. This domain is missing in the S. cerevisiae structure, as the first 200 amino acids of each subunit, containing this domain were cleaved before crystallization. d, GFP tagging of the complex subunits does not affect cell growth with glucose as carbon source (YPD). A representative image from three biologically independent experiments is shown. e, Model of PFK assembly pathways.
Extended Data Fig. 5 Aggregation and degradation propensity of individual complex subunits.
a, Stability of individual complex subunits, tagged by GFP, determined by CHX chase, in wild-type and deletion strains expressing orphan complex subunit. Cells with GFP fluorescence were analysed by FACS. Mean GFP fluorescence ± s.e.m. are presented with each data point from three biologically independent experiments overlaid. In each experiment, 20,000 events were recorded. **P = 0.0253, two tailed t-test. b, Solubility of individual complex subunits, tagged by GFP, determined by localization patterns changes, in wild-type and in deletion strains expressing orphan complex subunit. Log-phase cells (30 °C) were fixed and analysed by confocal microscopy (left). A representative image is shown. Scale bar, 4 μm. The fraction of cells displaying foci of GFP-tagged subunit per cell was quantified (right) (n = 155 cells per sample; from three biologically independent experiments). Data are mean ± s.e.m. overlaid with each data point. c, Subunit aggregation is complex-specific. Solubility of the Naa15–GFP subunit of the NatA complex in trp2∆ mutant cells deleted for the Trp2 subunit of the TRP complex, analysed as in b. (n = 155 cells/sample; from three biologically independent experiments). Data are mean ± s.e.m. overlaid with each data point. **P = 1.367248 × 10−11 (middle) and P = 7.850135 × 10−10 of a (lower panel) of a two tailed t-test. d, Characteristics of cotranslational complex assembly interactions. Left, zoom-in on the first 400 codons, displaying the onset and persistence of cotranslational interaction of each subunit with its partner subunit or subunits, for all 14 subunits identified as cotranslationally engaged. Right, the corresponding normalized length of each ORF at the onset of cotranslational interactions with partner subunits, demonstrating the length variability at the onset position.
Extended Data Fig. 6 Proteome wide bioinformatics analysis of Ssb1 interplay with putative onset of cotranslational assembly interactions.
a, Metagene analysis of Ssb1–GFP interaction profiles with the nascent chains of 116 yeast proteins identified as putative cotranslationally assembling subunits (putative assembly identification algorithm and parameters are detailed in the Supplementary Information). The dark grey line indicates Ssb interaction profiles4, aligned to the subunits putative onset of cotranslational subunit association positions depicted as 0 (onset position alignment). A zoomed-in view of the nascent-chain segments at assembly onset position ±75 amino acids is shown. The orange line indicates Ssb binding profiles for nascent chains aligned to random positions along the ORFs. Data are from two biologically independent experiments. The area between replicates is shaded, indicating the degree of experimental variation. There is no correlation detected between the random and onset position alignment (Pearson correlation r2 = 0.2911), thus Ssb depletion at positions of onset is significant. b, Average Kyte–Doolittle hydrophobicity plot (7-amino-acid window) of the 116 nascent-chain segments. A zoomed-in view of the nascent-chain segments at assembly onset position ±75 amino acids is shown, as in a.
Extended Data Fig. 7 Cotranslational interactions networks of FAS β, Cpa2 and PFK β metabolic enzymes subunits, analysed by SeRP.
a, Fatty acid synthesis metabolic pathway: nascent Faa1 is not engaged by C-terminally-tagged FAS complex β subunit, whereas nascent Acc1 shows a transient interaction, crossing the twofold enrichment threshold, at position approximately 250 codons/amino acids (indicated by an arrow). b, Arginine biosynthetic pathway: nascent Arg4 (argininosuccinate lyase) is not engaged by C-terminally-tagged Cpa2 subunit, whereas nascent Arg1 shows several transient interactions crossing the twofold enrichment threshold, at positions indicated by arrows. c, Glycolysis pathway: nascent Fba1 (fructose 1,6-bisphosphate aldolase) is not engaged by C-terminally tagged PFK complex β subunit, whereas Pyc2 (pyruvate carboxylase isoform) shows several transient interactions crossing the twofold enrichment threshold, at positions indicated by arrows. a–c, Data are from two biologically independent experiments. The area between replicates is shaded, indicating the degree of experimental variation.
Extended Data Fig. 8 Model of cotranslational folding and assembly of complex subunits.
a, Nascent chains emerging from the ribosome exit tunnel are first engaged by ribosome-associated chaperones. Upon emergence of a complete interaction domain the nascent chain is engaged by its complex partner subunit. This engagement remains stable throughout the rest of the ORF translation. b, The nascent-chain amino acid composition at the ribosome exit tunnel may direct the interplay between Ssb and partner-subunit association. High hydrophobicity and positively charged amino acids (aa) are engaged by Ssb; low hydrophobicity disfavours binding of Ssb at the onset of subunit association, allowing for folding of the interaction domain and subunit joining. c, Modes of cotranslational assembly: most complexes are assembled in a unidirectional manner, in which one dedicated, fully synthesized subunit engages its nascent partner. d, Diverging misfolding propensities of complex subunits: subunits engaged as nascent chains are prone to misfolding, whereas their partner subunits are generally more stable.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-2 and Supplementary Figure 1, the uncropped blots with size marker indications.
Rights and permissions
About this article
Cite this article
Shiber, A., Döring, K., Friedrich, U. et al. Cotranslational assembly of protein complexes in eukaryotes revealed by ribosome profiling. Nature 561, 268–272 (2018). https://doi.org/10.1038/s41586-018-0462-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0462-y
Keywords
This article is cited by
-
Direct structural analysis of a single acyl carrier protein domain in fatty acid synthase from the fungus Saccharomyces cerevisiae
Communications Biology (2024)
-
Diverging co-translational protein complex assembly pathways are governed by interface energy distribution
Nature Communications (2024)
-
Mutational biases favor complexity increases in protein interaction networks after gene duplication
Molecular Systems Biology (2024)
-
The ribosome lowers the entropic penalty of protein folding
Nature (2024)
-
Hierarchical TAF1-dependent co-translational assembly of the basal transcription factor TFIID
Nature Structural & Molecular Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.