Abstract
Variants of uncertain significance fundamentally limit the clinical utility of genetic information. The challenge they pose is epitomized by BRCA1, a tumour suppressor gene in which germline loss-of-function variants predispose women to breast and ovarian cancer. Although BRCA1 has been sequenced in millions of women, the risk associated with most newly observed variants cannot be definitively assigned. Here we use saturation genome editing to assay 96.5% of all possible single-nucleotide variants (SNVs) in 13 exons that encode functionally critical domains of BRCA1. Functional effects for nearly 4,000 SNVs are bimodally distributed and almost perfectly concordant with established assessments of pathogenicity. Over 400 non-functional missense SNVs are identified, as well as around 300 SNVs that disrupt expression. We predict that these results will be immediately useful for the clinical interpretation of BRCA1 variants, and that this approach can be extended to overcome the challenge of variants of uncertain significance in additional clinically actionable genes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Rehm, H. L. et al. ClinGen–the Clinical Genome Resource. N. Engl. J. Med. 372, 2235–2242 (2015).
Kuchenbaecker, K. B. et al. Risks of breast, ovarian, and contralateral breast cancer for BRCA1 and BRCA2 mutation carriers. J. Am. Med. Assoc. 317, 2402–2416 (2017).
Hall, J. M. et al. Linkage of early-onset familial breast cancer to chromosome 17q21. Science 250, 1684–1689 (1990).
Olopade, O. I. & Artioli, G. Efficacy of risk-reducing salpingo-oophorectomy in women with BRCA-1 and BRCA-2 mutations. Breast J. 10, S5–S9 (2004).
Rebbeck, T. R. et al. Bilateral prophylactic mastectomy reduces breast cancer risk in BRCA1 and BRCA2 mutation carriers: the PROSE Study Group. J. Clin. Oncol. 22, 1055–1062 (2004).
Easton, D. F. et al. Gene-panel sequencing and the prediction of breast-cancer risk. N. Engl. J. Med. 372, 2243–2257 (2015).
Landrum, M. J. et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 44, D862–D868 (2016).
Millot, G. A. et al. A guide for functional analysis of BRCA1 variants of uncertain significance. Hum. Mutat. 33, 1526–1537 (2012).
Ransburgh, D. J. R., Chiba, N., Ishioka, C., Toland, A. E. & Parvin, J. D. Identification of breast tumor mutations in BRCA1 that abolish its function in homologous DNA recombination. Cancer Res. 70, 988–995 (2010).
Pierce, A. J., Hu, P., Han, M., Ellis, N. & Jasin, M. Ku DNA end-binding protein modulates homologous repair of double-strand breaks in mammalian cells. Genes Dev. 15, 3237–3242 (2001).
Bouwman, P. et al. A high-throughput functional complementation assay for classification of BRCA1 missense variants. Cancer Discov. 3, 1142–1155 (2013).
Woods, N. T. et al. Functional assays provide a robust tool for the clinical annotation of genetic variants of uncertain significance. NPJ Genom. Med. 1, 16001 (2016).
Starita, L. M. et al. Massively parallel functional analysis of BRCA1 RING domain variants. Genetics 200, 413–422 (2015).
Steffensen, A. Y. et al. Functional characterization of BRCA1 gene variants by mini-gene splicing assay. Eur. J. Hum. Genet. 22, 1362–1368 (2014).
de la Hoya, M. et al. Combined genetic and splicing analysis of BRCA1 c.[594-2A>C; 641A>G] highlights the relevance of naturally occurring in-frame transcripts for developing disease gene variant classification algorithms. Hum. Mol. Genet. 25, 2256–2268 (2016).
Ghosh, R., Oak, N. & Plon, S. E. Evaluation of in silico algorithms for use with ACMG/AMP clinical variant interpretation guidelines. Genome Biol. 18, 225 (2017).
Gibson, T. J., Seiler, M. & Veitia, R. A. The transience of transient overexpression. Nat. Methods 10, 715–721 (2013).
Moynahan, M. E., Chiu, J. W., Koller, B. H. & Jasin, M. BRCA1 controls homology-directed DNA repair. Mol. Cell 4, 511–518 (1999).
Drost, R. et al. BRCA1 RING function is essential for tumor suppression but dispensable for therapy resistance. Cancer Cell 20, 797–809 (2011).
Shakya, R. et al. BRCA1 tumor suppression depends on BRCT phosphoprotein binding, but not its E3 ligase activity. Science 334, 525–528 (2011).
Vega, A. et al. The R71G BRCA1 is a founder Spanish mutation and leads to aberrant splicing of the transcript. Hum. Mutat. 17, 520–521 (2001).
Findlay, G. M., Boyle, E. A., Hause, R. J., Klein, J. C. & Shendure, J. Saturation editing of genomic regions by multiplex homology-directed repair. Nature 513, 120–123 (2014).
Blomen, V. A. et al. Gene essentiality and synthetic lethality in haploid human cells. Science 350, 1092–1096 (2015).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Beumer, K. J. et al. Efficient gene targeting in Drosophila by direct embryo injection with zinc-finger nucleases. Proc. Natl Acad. Sci. USA 105, 19821–19826 (2008).
Essletzbichler, P. et al. Megabase-scale deletion using CRISPR/Cas9 to generate a fully haploid human cell line. Genome Res. 24, 2059–2065 (2014).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Pollard, K. S., Hubisz, M. J., Rosenbloom, K. R. & Siepel, A. Detection of nonneutral substitution rates on mammalian phylogenies. Genome Res. 20, 110–121 (2010).
Tavtigian, S. V., Byrnes, G. B., Goldgar, D. E. & Thomas, A. Classification of rare missense substitutions, using risk surfaces, with genetic- and molecular-epidemiology applications. Hum. Mutat. 29, 1342–1354 (2008).
Towler, W. I. et al. Analysis of BRCA1 variants in double-strand break repair by homologous recombination and single-strand annealing. Hum. Mutat. 34, 439–445 (2013).
Starita, L. M. et al. A multiplexed homology-directed DNA repair assay reveals the impact of over 1,000 BRCA1 missense substitution variants on protein function. Am. J. Hum. Genet. https://doi.org/10.1016/j.ajhg.2018.07.016 (2018).
Brzovic, P. S., Rajagopal, P., Hoyt, D. W., King, M. C. & Klevit, R. E. Structure of a BRCA1–BARD1 heterodimeric RING–RING complex. Nat. Struct. Biol. 8, 833–837 (2001).
Shiozaki, E. N., Gu, L., Yan, N. & Shi, Y. Structure of the BRCT repeats of BRCA1 bound to a BACH1 phosphopeptide: implications for signaling. Mol. Cell 14, 405–412 (2004).
Wegrzyn, J. L., Drudge, T. M., Valafar, F. & Hook, V. Bioinformatic analyses of mammalian 5′-UTR sequence properties of mRNAs predicts alternative translation initiation sites. BMC Bioinformatics 9, 232 (2008).
Desmet, F.-O. et al. Human Splicing Finder: an online bioinformatics tool to predict splicing signals. Nucleic Acids Res. 37, e67 (2009).
Gasperini, M., Starita, L. & Shendure, J. The power of multiplexed functional analysis of genetic variants. Nat. Protoc. 11, 1782–1787 (2016).
Starita, L. M. et al. Variant interpretation: functional assays to the rescue. Am. J. Hum. Genet. 101, 315–325 (2017).
Plon, S. E. et al. Sequence variant classification and reporting: recommendations for improving the interpretation of cancer susceptibility genetic test results. Hum. Mutat. 29, 1282–1291 (2008).
Lovelock, P. K. et al. Identification of BRCA1 missense substitutions that confer partial functional activity: potential moderate risk variants? Breast Cancer Res. 9, R82 (2007).
Carette, J. E. et al. Ebola virus entry requires the cholesterol transporter Niemann–Pick C1. Nature 477, 340–343 (2011).
Walsh, T. et al. Detection of inherited mutations for breast and ovarian cancer using genomic capture and massively parallel sequencing. Proc. Natl Acad. Sci. USA 107, 12629–12633 (2010).
Hsu, P. D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR–Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Colombo, M. et al. Comprehensive annotation of splice junctions supports pervasive alternative splicing at the BRCA1 locus: a report from the ENIGMA consortium. Hum. Mol. Genet. 23, 3666–3680 (2014).
Romero, A. et al. BRCA1 alternative splicing landscape in breast tissue samples. BMC Cancer 15, 219 (2015).
Tavtigian, S. V. et al. Comprehensive statistical study of 452 BRCA1 missense substitutions with classification of eight recurrent substitutions as neutral. J. Med. Genet. 43, 295–305 (2006).
Kumar, P., Henikoff, S. & Ng, P. C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 4, 1073–1081 (2009).
Adzhubei, I. & Jordan, D. M. Predicting functional effect of human missense mutations using PolyPhen-2. Curr. Protoc. Hum. Gen. 76, 7.20.1–7.20.41 (2013).
Acknowledgements
We thank M. Spielmann, D. Witten, A. McKenna, M. Kircher, M. Dougherty, J. Lazar, Y. Yin, and B. Shirts for insights on data analysis and/or comments on the manuscript, J. Kitzman for sharing reagents and protocols, R. Acuña-Hidalgo, J. Milbank, and E. van Veen for experimental assistance, and the Feng Zhang laboratory for sharing Cas9/gRNA plasmids. This work was supported by the Brotman Baty Institute for Precision Medicine, an NIH Director’s Pioneer Award (DP1HG007811 to J.S.) and a training award from the National Cancer Institute (F30CA213728 to GMF). J.S. is an Investigator of the Howard Hughes Medical Institute.
Reviewer information
Nature thanks H. Rehm, J. Weissman and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
G.M.F., J.S. and L.M.S. conceived the project. G.M.F. designed experiments. G.M.F. and R.M.D. performed experiments with assistance from B.M., M.D.Z., A.P.L., L.M.S. and M.G. G.M.F. performed analysis with assistance from L.M.S., J.D.J., X.H. and R.M.D. G.M.F, J.S. and L.M.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 CRISPR targeting of HDR pathway genes to confirm essentiality in HAP1 cells.
a, Schematic, HAP1 cells are transfected with a plasmid expressing a gRNA and a Cas9-2A-puromycin cassette24. Owing to low transfection rates for HAP1 cells, puromycin selection reduces viable cells in all transfections. Over time, however, CRISPR targeting of non-essential genes leads to increased cell growth compared to CRISPR targeting of essential genes. b, HAP1 cell populations were transfected with a Cas9/gRNA plasmid either targeting the non-essential gene HPRT1 (control) or exon 17 of BRCA1 on day 0. Successfully transfected cells were selected with puromycin (days 1–4) and cultured until imaging on day 7, at which point cells were imaged. Images are representative of two transfection replicates. c, Cell viability of HAP1 cells transfected with Cas9/gRNA constructs targeting different HDR genes and controls (HPRT1, TP53) was measured using the CellTiterGlow assay. Luminescence is proportional to the number of living cells in each well when the assay is performed. Triplicate wells for each gRNA at each time point were processed, quantified on a plate reader and averaged. Error bars show the standard error of the mean. gRNA sequences are included in Supplementary Table 3. d, The targeted BRCA1 exon 17 locus was deeply sequenced from a population of transfected cells sampled on day 5 and day 11. The fold-change from day 5 to day 11 for each editing outcome observed at a frequency over 0.001 in day 5 sequencing reads is plotted.
Extended Data Fig. 2 Analysis of Cas9-induced indels observed in BRCA1 SGE experiments.
Variants observed in gDNA sequencing were included in this analysis if (i) they aligned to the reference with either a single insertion or deletion within 15 bp of the predicted Cas9 cleavage site and (ii) were observed at a frequency greater than 1 in 10,000 reads in both replicates. a, Histograms show the number of unique indels observed of each size, with negative sizes corresponding to deletions. More unique indels were observed in wild-type HAP1 cells compared to HAP1-LIG4KO cells for exons compared (wild-type data for exon 22 was excluded). b, Day 11 over day 5 indel frequencies were normalized to the median synonymous SNV in each replicate and then averaged across replicates to measure selection on each indel. The distribution of selective effects is shown for each experiment as a histogram, in which indels are coloured by whether their size was divisible by 3 (that is, ‘in-frame’ versus ‘frameshifting’). Whereas frameshifting variants were consistently depleted, some exons were tolerant to in-frame indels.
Extended Data Fig. 3 HAP1 cell line optimizations for saturation genome editing to assay essential genes.
a, A gRNA targeting Cas9 to the coding sequence of LIG4, a gene integral to the non-homologous end-joining pathway, was cloned into a vector co-expressing Cas9-2A-GFP24. Wild-type HAP1 cells were transfected, and single GFP-expressing cells were sorted into wells of a 96-well plate. Eight monoclonal lines were grown out over a period of three weeks and screened using Sanger sequencing for frameshifting indels in LIG4. The Sanger trace shows the frameshifting deletion present in the clonal line chosen for subsequent experiments, referred to as HAP1-LIG4KO. b, To purify HAP1 cells for haploid cells, live cells were stained for DNA content with Hoechst 34580 and sorted using a gate to select cells with the lowest DNA content, corresponding to 1n cells in G1. c, The fraction of all possible SNVs scored is shown for each exon. SNVs were excluded mainly due to proximity to the HDR marker and/or poor sampling (Methods). d, e, Measurements across replicates are plotted for exon 17 SNVs assayed in HAP1-LIG4KO cells to show correlations of day 5 frequencies (d) and day 11 over library ratios (e). f–h, Plots comparing SNV function scores across replicate experiments for exon 17 saturation genome editing experiments performed in unsorted wild-type HAP1 cells (f), HAP1-LIG4KO cells (g), and wild-type HAP1 cells sorted on 1n ploidy (h). i, Function scores (averaged across replicates) are plotted to compare results for exon 17 experiments performed in wild-type 1n-sorted HAP1 cells and HAP1-LIG4KO cells. The number of SNVs plotted and the Spearman correlation is displayed for each plot (d–i).
Extended Data Fig. 4 Correlations for SNV measurements within single experiments, across transfection replicates, and to CADD scores for all SGE experiments.
Heat maps indicate Spearman correlation coefficients for SNV measurements from experiments in wild-type HAP1 cells (a) and in HAP1-LIG4KO cells (b). Grey boxes indicate absent RNA data from wild-type HAP1 cells. The four leftmost columns show how SNV frequencies correlate between samples from within a single replicate experiment. The unusually high correlations between exon 22 SNV frequencies in the plasmid library and in day 5 gDNA samples from wild-type HAP1 cells suggests plasmid contamination in gDNA. Indeed, primer homology to a repetitive element in the exon 22 library was identified. Consequently, the wild-type HAP1 exon 22 data was removed from analysis and a different primer specific to gDNA was used to prepare exon 22 sequencing amplicons from HAP1-LIG4KO cells. The low HAP1-LIG4KO correlations between exon 18 SNV frequencies in day 5 gDNA and RNA and between RNA replicates suggests RNA sample bottlenecking consequential to low RNA yields. Therefore, exon 18 RNA was also excluded from analysis. Consistent with the higher rates of HDR-mediated genome editing (Fig. 2a), replicate correlations (middle columns) were generally higher in HAP1-LIG4KO cells than wild-type HAP1 cells. CADD scores predict the deleteriousness of each SNV, and are therefore negatively correlated with function scores (rightmost columns).
Extended Data Fig. 5 Models of SNV editing rates across BRCA1 exons to account for positional biases.
Gene conversion tracts arising during HDR in human cells are short such that library SNVs are introduced to the genome more frequently near the CRISPR target site. We modelled this positional effect in our data for n = 4,002 SNVs (pre-filtering) using a LOESS regression fit on day 5 over library SNV ratios. a, Plots shown here are of the average of n = 2 replicates per exon, with the black line indicating the LOESS regression. By day 5, selective effects on gene function are evidenced by nonsense SNVs (red) appearing at lower frequencies compared to neighbouring SNVs. Therefore, to best approximate the SNV editing rate as a function of position alone (that is, the ‘baseline’), the regression excluded SNVs that were selected against between day 11 and day 5 (see Methods). b, c, Day 11 over library SNV ratios were adjusted by the positional fit for each experiment in calculating function scores. This adjustment is illustrated here for an exon 3 replicate by plotting the day 11 over library ratio as a function of position before (b) and after (c) adjustment for (n = 298 SNVs). The elevated day 11 over library ratios for SNVs near the CRISPR cleavage site (indicated with an arrow) are corrected to achieve a more uniform baseline across the mutagenized region. d, e, The distributions of SNV day 11 over library ratios before and after accounting for positional effects are shown, coloured by mutational consequence (n = 4,002 SNVs, averaged across n = 2 replicates).
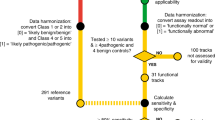
Extended Data Fig. 6 SNV filtering to prevent erroneous functional classification.
a, The flow chart describes filters used to produce the final SNV dataset and shows how many SNVs were removed at each step. b, Raw day 5 over library SNV ratios are shown for a portion of exon 15 to illustrate how re-editing biases necessitate filtering. The three depleted SNVs marked with asterisks create alternative PAM sequences that probably allow the Cas9–gRNA complex to re-cut the locus and cause their removal. For other SNVs, the fixed PAM edit (a GGG to GCG synonymous change) minimalizes re-editing. Alternative PAM sequences created by each indicated SNV are shown in magenta. The LOESS regression curve in shown in black. c, d, Plots show the relationship between day 5 over library and day 11 over day 5 ratios before (c) and after (d) filtering steps 1 and 2. Filtering removes outliers because editing biases primarily affect the day 5 over library ratio. e–g, Histograms show the distributions of function scores for SNVs deemed ‘pathogenic’ or ‘benign’ in ClinVar at different stages of filtering. Scores in e are derived before normalization across exons.
Extended Data Fig. 7 Mixture modelling of scores to classify SNVs by functional effect.
a, Distributions of ‘non-functional’ and ‘functional’ SNVs plotted here were defined respectively as all nonsense SNVs and all synonymous SNVs with RNA scores within 1 standard deviation of the median synonymous SNV. b, An ROC curve was generated using SGE function scores to distinguish the 634 ‘functional’ and ‘non-functional’ SNVs defined in a. c, A two-component Gaussian mixture model was used to produce point estimates of the probability that each SNV was ‘non-functional’, Pnf, given its average function score across replicates. These P values are plotted in d against function scores for a subset of the data. Thresholds were set such that Pnf < 0.01 corresponds to ‘functional’, and Pnf > 0.99 corresponds to ‘non-functional’, and 0.01 < Pnf < 0.99 corresponds to ‘intermediate’ classification. Functional classification thresholds are drawn as dashed lines; black denotes the non-functional threshold and grey the intermediate threshold. e, f, SNV function scores across replicates are plotted for each exon with SNVs coloured by mutational consequence (e), and for each type of mutational consequence with SNVs coloured by ClinVar status (f). Using the optimal function score cutoff for all SNVs tested (Fig. 3b), sensitivities and specificities for distinguishing ‘Pathogenic’/’Likely pathogenic’ from ‘Benign’/’Likely benign’ ClinVar annotations for each type of mutation are as follows: 92.7% and 92.9% for missense SNVs (n = 55), 100% and 100% for splice region SNVs (n = 23), and 95.2% sensitivity for canonical splice site SNVs (n = 83; specificity not calculable).
Extended Data Fig. 8 BRCA1 SNVs observed more frequently in large-scale population sequencing are more likely to score as functional.
a–c, SNV function scores are plotted against gnomAD (a), Bravo (b), and FLOSSIES (c) allele frequencies. a, Among the 302 SNVs assayed also present in gnomAD, higher allele frequencies associate with higher function scores (Wilcoxon signed-rank test, P = 3.7 × 10−12). b, Bravo is a collection of whole-genome sequences ascertained from 62,784 individuals through the NHLBI TOPMed program. Similarly to SNVs present in gnomAD, higher allele frequencies in Bravo correlate with higher function scores. c, FLOSSIES is a database of variants seen in targeted sequencing of breast cancer genes sampled from approximately 10,000 cancer-free women who are at least 70 years old. Only 1 of 39 assayed SNVs present in FLOSSIES scored as non-functional. c, d, Missense SNVs in ClinVar are separated by whether they have (c) or have not (d) been seen in either gnomAD or Bravo and function scores across replicates are plotted, with dashed lines demarcating functional classes. A higher proportion of ClinVar missense SNVs absent from gnomAD and Bravo score as non-functional (50.6% versus 15.7%; Fisher’s exact test, P = 1.80 × 10−17).
Extended Data Fig. 9 SGE function scores correlate with computational metrics and perform favourably at predicting ClinVar annotations.
a, SNV function scores are plotted against mammalian phyloP scores, with colours indicative of ClinVar status (Spearman’s correlation shown). b, c, ROC curves show the performance of CADD scores and phyloP scores for discriminating ClinVar ‘pathogenic’ and ‘benign’ SNVs (including ‘likely’), as described in Fig. 3b for SGE data. d–g Plots as in a, but for missense SNVs only, showing correlations between SGE function scores and CADD28 scores, phyloP scores29, Grantham differences (Grantham amino acid variation minus Grantham amino acid deviation; GV − GD), and align-GVGD classifications47. Missense SNV function scores also correlate with SIFT scores48 (ρ = 0.363) and PolyPhen-2 scores49 (ρ = −0.277). (Spearman’s correlation, P < 1 × 10–37 for all correlations). h–l, ROC curves assess the performance of SGE function scores and each indicated metric at distinguishing firmly ‘pathogenic’ and ‘benign’ missense SNVs (not including ‘likely’). m, n, SGE scores for missense variants are plotted against results from homology-directed repair assays9,31 (m) and results from transcriptional activation assays12 (n). In cases where multiple SNVs assayed lead to same amino acid substitution, function scores were averaged and coloured red if either SNV had an RNA score less than −2. Box plots depict the sample median (line) and the interquartile range (box).
Extended Data Fig. 10 Evidence supporting SNV scores in discordance with ClinVar classifications.
a, b, Complete maps of RNA scores for exons 16 (a) and exon 19 (b) reveal highly variable sensitivity to RNA depletion. The location of the strongest predicted exonic splice enhancer in exon 16 is indicated by the orange line36. c, Function scores (means from two replicates) are plotted to compare results from preliminary experiments in wild-type HAP1 to those in HAP1-LIG4KO. Data are shown only for experiments with Spearman’s correlations between replicates greater than 0.50 in wild-type HAP1 cells (n = 2,096 SNVs; exons 3, 4, 5, 16, 17, 19, 21). Discordantly classified SNVs are indicated with arrows. c.19–2A>G was the only firmly discordant SNV for which the function score could not be corroborated in wild-type HAP1, consequent to low reproducibility of exon 2 wild-type function scores. Indeed, c.19–2A>G scored highly variably between wild-type replicates. d, The sequence-function map of exon 21 is shown with the function scores for the two ‘pathogenic’ SNVs observed in linkage indicated. Dashed lines demarcate functional classifications. c, Function scores are plotted against CADD scores for all canonical splice SNVs assayed, coloured by ClinVar status. The six possible exon 2 splice acceptor SNVs (circled) have the lowest CADD scores among all canonical splice SNVs assayed, and none score as ‘non-functional’. e, A USCS Genome Browser shot shows the PhyloP conservation track and selected mammalian sequence alignments for the exon 2 acceptor region, with the canonical acceptor site nucleotides highlighted in light blue (hg19 chr17:41,276,108–41,276,139). Multiple mammalian species are identified that have a G at position c.19–2 of the human transcript (corresponding to a C in the plus-strand orientation shown).
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1-3, Supplementary References and a Supplementary Table Guide.
Supplementary Table 1
Saturation genome editing scores for 3,893 BRCA1 SNVs – see Supplementary Information document for full description.
Supplementary Table 2
Analysis of SNVs with conflicting interpretations in ClinVar – see Supplementary Information document for full description.
Supplementary Table 3
DNA sequences used in BRCA1 saturation genome editing experiments – see Supplementary Information document for full description.
Rights and permissions
About this article
Cite this article
Findlay, G.M., Daza, R.M., Martin, B. et al. Accurate classification of BRCA1 variants with saturation genome editing. Nature 562, 217–222 (2018). https://doi.org/10.1038/s41586-018-0461-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0461-z
Keywords
This article is cited by
-
Minimum information and guidelines for reporting a multiplexed assay of variant effect
Genome Biology (2024)
-
A loss-of-function variant in ZCWPW1 causes human male infertility with sperm head defect and high DNA fragmentation
Reproductive Health (2024)
-
scSNV-seq: high-throughput phenotyping of single nucleotide variants by coupled single-cell genotyping and transcriptomics
Genome Biology (2024)
-
Whole genome sequencing in clinical practice
BMC Medical Genomics (2024)
-
Combining full-length gene assay and SpliceAI to interpret the splicing impact of all possible SPINK1 coding variants
Human Genomics (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.