Abstract
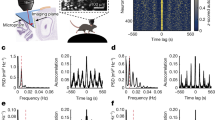
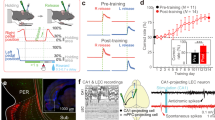
The encoding of time and its binding to events are crucial for episodic memory, but how these processes are carried out in hippocampal–entorhinal circuits is unclear. Here we show in freely foraging rats that temporal information is robustly encoded across time scales from seconds to hours within the overall population state of the lateral entorhinal cortex. Similarly pronounced encoding of time was not present in the medial entorhinal cortex or in hippocampal areas CA3–CA1. When animals’ experiences were constrained by behavioural tasks to become similar across repeated trials, the encoding of temporal flow across trials was reduced, whereas the encoding of time relative to the start of trials was improved. The findings suggest that populations of lateral entorhinal cortex neurons represent time inherently through the encoding of experience. This representation of episodic time may be integrated with spatial inputs from the medial entorhinal cortex in the hippocampus, allowing the hippocampus to store a unified representation of what, where and when.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Tulving, E. Elements of Episodic Memory (Oxford Univ. Press, Oxford, 1985).
Clayton, N. S. & Dickinson, A. Episodic-like memory during cache recovery by scrub jays. Nature 395, 272–274 (1998).
Eichenbaum, H. On the integration of space, time, and memory. Neuron 95, 1007–1018 (2017).
Fortin, N. J., Agster, K. L. & Eichenbaum, H. B. Critical role of the hippocampus in memory for sequences of events. Nat. Neurosci. 5, 458–462 (2002).
Kesner, R. P., Gilbert, P. E. & Barua, L. A. The role of the hippocampus in memory for the temporal order of a sequence of odors. Behav. Neurosci. 116, 286–290 (2002).
Ekstrom, A. D. & Bookheimer, S. Y. Spatial and temporal episodic memory retrieval recruit dissociable functional networks in the human brain. Learn. Mem. 14, 645–654 (2007).
Lehn, H. et al. A specific role of the human hippocampus in recall of temporal sequences. J. Neurosci. 29, 3475–3484 (2009).
Naya, Y. & Suzuki, W. A. Integrating what and when across the primate medial temporal lobe. Science 333, 773–776 (2011).
Ezzyat, Y. & Davachi, L. Similarity breeds proximity: pattern similarity within and across contexts is related to later mnemonic judgments of temporal proximity. Neuron 81, 1179–1189 (2014).
Hsieh, L. T., Gruber, M. J., Jenkins, L. J. & Ranganath, C. Hippocampal activity patterns carry information about objects in temporal context. Neuron 81, 1165–1178 (2014).
Dede, A. J. O., Frascino, J. C., Wixted, J. T. & Squire, L. R. Learning and remembering real-world events after medial temporal lobe damage. Proc. Natl Acad. Sci. USA 113, 13480–13485 (2016).
Buzsáki, G. & Llinás, R. Space and time in the brain. Science 358, 482–485 (2017).
Ferbinteanu, J., Kennedy, P. J. & Shapiro, M. L. Episodic memory—from brain to mind. Hippocampus 16, 691–703 (2006).
Hassabis, D. & Maguire, E. A. Deconstructing episodic memory with construction. Trends Cogn. Sci. 11, 299–306 (2007).
Pastalkova, E., Itskov, V., Amarasingham, A. & Buzsáki, G. Internally generated cell assembly sequences in the rat hippocampus. Science 321, 1322–1327 (2008).
MacDonald, C. J., Lepage, K. Q., Eden, U. T. & Eichenbaum, H. Hippocampal “time cells” bridge the gap in memory for discontiguous events. Neuron 71, 737–749 (2011).
Kraus, B. J. et al. During running in place, grid cells integrate elapsed time and distance run. Neuron 88, 578–589 (2015).
Mau, W. et al. The same hippocampal CA1 population simultaneously codes temporal information over multiple timescales. Curr. Biol. 28, 1499-1508.e4 (2018).
Eichenbaum, H. Time cells in the hippocampus: a new dimension for mapping memories. Nat. Rev. Neurosci. 15, 732–744 (2014).
Cai, D. J. et al. A shared neural ensemble links distinct contextual memories encoded close in time. Nature 534, 115–118 (2016).
Manns, J. R., Howard, M. W. & Eichenbaum, H. Gradual changes in hippocampal activity support remembering the order of events. Neuron 56, 530–540 (2007).
Mankin, E. A. et al. Neuronal code for extended time in the hippocampus. Proc. Natl Acad. Sci. USA 109, 19462–19467 (2012).
Ziv, Y. et al. Long-term dynamics of CA1 hippocampal place codes. Nat. Neurosci. 16, 264–266 (2013).
Mankin, E. A., Diehl, G. W., Sparks, F. T., Leutgeb, S. & Leutgeb, J. K. Hippocampal CA2 activity patterns change over time to a larger extent than between spatial contexts. Neuron 85, 190–201 (2015).
Rubin, A., Geva, N., Sheintuch, L. & Ziv, Y. Hippocampal ensemble dynamics timestamp events in long-term memory. eLife 4, e12247 (2015).
Hargreaves, E. L., Rao, G., Lee, I. & Knierim, J. J. Major dissociation between medial and lateral entorhinal input to dorsal hippocampus. Science 308, 1792–1794 (2005).
Deshmukh, S. S. & Knierim, J. J. Representation of non-spatial and spatial information in the lateral entorhinal cortex. Front. Behav. Neurosci. 5, 69 (2011).
Lu, L. et al. Impaired hippocampal rate coding after lesions of the lateral entorhinal cortex. Nat. Neurosci. 16, 1085–1093 (2013).
Keene, C. S. et al. Complementary functional organization of neuronal activity patterns in the perirhinal, lateral entorhinal, and medial entorhinal cortices. J. Neurosci. 36, 3660–3675 (2016).
Pilkiw, M. et al. Phasic and tonic neuron ensemble codes for stimulus-environment conjunctions in the lateral entorhinal cortex. eLife 6, e28611 (2017).
Howard, M. W. et al. A unified mathematical framework for coding time, space, and sequences in the hippocampal region. J. Neurosci. 34, 4692–4707 (2014).
Rigotti, M. et al. The importance of mixed selectivity in complex cognitive tasks. Nature 497, 585–590 (2013).
Buonomano, D. V. & Maass, W. State-dependent computations: spatiotemporal processing in cortical networks. Nat. Rev. Neurosci. 10, 113–125 (2009).
McEchron, M. D., Tseng, W. & Disterhoft, J. F. Single neurons in CA1 hippocampus encode trace interval duration during trace heart rate (fear) conditioning in rabbit. J. Neurosci. 23, 1535–1547 (2003).
Folkerts, S., Rutishauser, U. & Howard, M. W. Human episodic memory retrieval is accompanied by a neural contiguity effect. J. Neurosci. 38, 4200–4211 (2018).
Finnerty, G. T., Shadlen, M. N., Jazayeri, M., Nobre, A. C. & Buonomano, D. V. Time in cortical circuits. J. Neurosci. 35, 13912–13916 (2015).
Mauk, M. D. & Buonomano, D. V. The neural basis of temporal processing. Annu. Rev. Neurosci. 27, 307–340 (2004).
Merchant, H., Harrington, D. L. & Meck, W. H. Neural basis of the perception and estimation of time. Annu. Rev. Neurosci. 36, 313–336 (2013).
Howard, M. W. & Kahana, M. J. A distributed representation of temporal context. J. Math. Psychol. 46, 269–299 (2002).
Dolorfo, C. L. & Amaral, D. G. Entorhinal cortex of the rat: organization of intrinsic connections. J. Comp. Neurol. 398, 49–82 (1998).
Bota, M., Sporns, O. & Swanson, L. W. Architecture of the cerebral cortical association connectome underlying cognition. Proc. Natl Acad. Sci. USA 112, E2093–E2101 (2015).
Jin, D. Z. Z., Fujii, N. & Graybiel, A. M. Neural representation of time in cortico-basal ganglia circuits. Proc. Natl Acad. Sci. USA 106, 19156–19161 (2009).
Hyman, J. M., Ma, L., Balaguer-Ballester, E., Durstewitz, D. & Seamans, J. K. Contextual encoding by ensembles of medial prefrontal cortex neurons. Proc. Natl Acad. Sci. USA 109, 5086–5091 (2012).
Igarashi, K. M., Lu, L., Colgin, L. L., Moser, M. B. & Moser, E. I. Coordination of entorhinal-hippocampal ensemble activity during associative learning. Nature 510, 143–147 (2014).
Rowland, D. C., Roudi, Y., Moser, M. B. & Moser, E. I. Ten years of grid cells. Annu. Rev. Neurosci. 39, 19–40 (2016).
Lu, L., Igarashi, K. M., Witter, M. P., Moser, E. I. & Moser, M. B. Topography of place maps along the CA3-to-CA2 axis of the hippocampus. Neuron 87, 1078–1092 (2015).
Tsao, A., Moser, M. B. & Moser, E. I. Traces of experience in the lateral entorhinal cortex. Curr. Biol. 23, 399–405 (2013).
Wood, E. R., Dudchenko, P. A., Robitsek, R. J. & Eichenbaum, H. Hippocampal neurons encode information about different types of memory episodes occurring in the same location. Neuron 27, 623–633 (2000).
Fan, R. E., Chang, K. W., Hsieh, C. J., Wang, X. R. & Lin, C. J. Liblinear: A library for large linear classification. J. Mach. Learn. Res. 9, 1871–1874 (2008).
Aggarwal, C. C., Hinneburg, A. & Keim, D. A. On the surprising behavior of distance metrics in high dimensional space. Lect. Notes Comput. Sci. 1973, 420–434 (2001).
Langston, R. F. et al. Development of the spatial representation system in the rat. Science 328, 1576–1580 (2010).
Kropff, E., Carmichael, J. E., Moser, M. B. & Moser, E. I. Speed cells in the medial entorhinal cortex. Nature 523, 419–424 (2015).
Skaggs, W. E., McNaughton, B. L. & Gothard, K. M. in Advances in Neural Information Processing Systems 5 (NIPS 1992) 1030–1037 (Morgan Kaufmann Publishers Inc., 1993).
Acknowledgements
We thank A. M. Amundsgård, K. Haugen, K. Jenssen, E. Kråkvik, and H. Waade for technical assistance and M. P. Witter for help with determining recording locations in the LEC. The work was supported by a fellowship from the Helen Hay Whitney Foundation (to A.T.), an Advanced Investigator Grant from the European Research Council (GRIDCODE’, grant number 338865), the Centre of Excellence scheme and the National Infrastructure Scheme of the Research Council of Norway (Centre for Neural Computation, grant number 223262; NORBRAIN1, grant number 197467), the Louis Jeantet Prize, the Körber Prize, and the Kavli Foundation.
Reviewer information
Nature thanks M. Shapiro and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
A.T., M.-B.M. and E.I.M. designed experiments and the analytic approach; A.T. developed and performed analyses; A.T., J.S. and L.L. collected data; C.W. and J.J.K. contributed circular track data. A.T. and E.I.M. wrote the paper with input from all authors. A.T. wrote the first draft and had the main role in developing the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Histology for LEC animals.
a–d, Nissl-stained coronal sections showing recording locations for LEC animals used in the BW12 experiments (a), BW4 and figure-eight experiments (b), circular track experiments (c), and object experiments (d). Red arrowheads indicate range for tetrode locations. Activity was recorded from neurons in all layers of the lateral half of the LEC. Dashed lines indicate approximate anatomical borders of the LEC.
Extended Data Fig. 2 Single-cell responses in the LEC related to time.
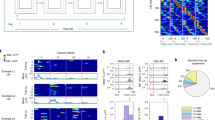
a, Schematic of the GLM used for determining cell selectivity. Learned weights for the relevant predictors, as determined by the stepwise selection process, were put through an exponential nonlinearity that returned the mean rate of a Poisson process from which spikes were drawn (for LEC, 14.0% selective for session time, 3.8% selective for trial time, 2.7% selective for a mixture of trial and session time, percentages averaged across all animals). b, Explained variance for all LEC cells fit by the GLM for BW12 experiment (n = 186 cells). Average explained variance was 0.05. c, Distribution of time constants for trial-time or session-time selective cells (n = 80 cells). Time constants were estimated for each cell classified as trial-time or session-time selective by fitting a single-term exponential. d, Left, five additional example cells exhibiting ramping activity across the session. The firing rate of each cell is shown in grey, with the model-predicted firing rate in blue. The R2 value is shown in for each cell. Right, average waveforms across four recording channels for the first (green line) and last (black line) quarter of the session. e, Pearson correlation between waveform distances (Euclidean distance between pairs of consecutive spike waveforms) and firing rate for all LEC cells used in the BW12 experiment (n = 451 cells). f, Comparison of the correlation between waveform distance and firing rate, as in e, for time-selective LEC cells (n = 92 cells, grey) and all other LEC cells used in the BW12 experiment (n = 359 cells, black). g, Six example cells demonstrating that notable fluctuations in firing rate can occur during stable recordings. For each cell, the top left row shows activity during trial periods, bottom left row shows activity during intertrial periods, and right panel shows average waveforms, as in d.
Extended Data Fig. 3 Time-cell-like activity.
a–h, Temporal specificity examined as a function of peak firing times of individual cells. a, b, Mean activity from example animals for the LEC (top matrix), CA3 (middle matrix) and MEC (bottom matrix) during trial (a) and intertrial (b) periods. For each matrix, rows show mean firing rates for individual cells ordered by the time of peak firing rate. Actual data shown in left column, shuffled data shown in right column. Cell identities were not maintained across actual and shuffled data. c, Fraction of cells with significant temporally specific activity for trial (top) and intertrial (bottom) periods (n = 3, 3 and 2 animals for LEC, CA3 and MEC, respectively). d, Time of peak activity for cells with significant temporally specific activity during trial periods. e, As in d, but for intertrial periods. f–h, Temporal specificity examined by calculating temporal information. f, Fraction of cells with significant temporal information for trial (top) and intertrial (bottom) periods (n = 3, 3 and 2 animals for LEC, CA3 and MEC, respectively). g, Distribution of significant temporal information scores for trial periods. h, As in g, but for intertrial periods. i, Predictors added for expanded GLM: symmetrical ramps and single-trial ramps. Coloured lines highlight the predictors for the respective trial indicated on top, grey lines are the rest of the available predictors. j, Explained variance for all LEC cells fit by expanded GLM for the BW12 experiment (n = 350 cells). Average explained variance was 0.03. k, Distribution of selectivity for time, wall colour and position for single LEC cells, determined using the expanded GLM (n = 3 animals). Shade indicates the same individual across the different variables. l, Left, examples of expanded GLM fit results for four cells with selectivity for different features. The firing rate of each cell is shown in grey, with the model-predicted firing rate in blue. The R2 value is shown for each cell. Right, average waveforms across four recording channels for the first quarter (green line) and last quarter (black line) quarter of the session. Circles indicate individual animals, solid lines indicate mean fraction of cells ± s.e.m.
Extended Data Fig. 4 Decoding of temporal epochs.
a, 2D projections of neural population responses during the BW12 experiment for trial periods only. Axes correspond to the first two linear discriminants (LD1 and LD2; arbitrary units). Left column shows LEC population responses, middle column shows CA3 population responses, right column shows MEC population responses, each from an example animal. The wall colour of each trial is indicated by a shade of green (black walls) or purple (white walls), with progression of shade from dark to light indicating the progression of trials. b, Fraction of variance explained by the first 20 principal components for each area. Principal components were computed using PCA on raw data. Lines indicate variance explained for individual animals (n = 3, 3 and 2 animals for LEC, CA3 and MEC, respectively). c, Regressing the first two principal components from PCA results for individual animals against time leads to significant fits for all areas, but substantially higher explained variance for LEC. P < 0.001 (LEC versus CA3, t(4) = 9.79), P < 0.01 (LEC versus MEC, t(3) = 6.13), unpaired t-test. Top row, example fits for individual animals are shown with black lines indicating time, coloured lines indicating regression fit and R2 values indicated for the example fit. Bottom two rows, first two principal components for example fits. d, Illustration of cross-validation procedure: fivefold cross-validation is shown for data containing four temporal epochs, with five time bins in each epoch. A different subset of time bins is used as test data for each iteration of the cross-validation procedure. Actual data consisted of 24 epochs (trial and intertrial periods) with 25 or 14 time bins in each epoch, and tenfold cross-validation was used. e, Z-scored decoding accuracy using cells recorded in a single day (left, P = 0.68, one-sided binomial test, n = 46 days), pairs of consecutive days (middle, P = 0.29, one-sided binomial test, n = 72 pairs), pairs of days separated by half the total number of recording days (right, P = 0.37, one-sided binomial test, n = 44 pairs). f, Decoding accuracy for temporal epoch using behaviour tracking data in place of neural activity (n = 3 animals). ‘All’ tracking data consisted of the animal’s position, velocity, acceleration and head direction. g, Left, decoding accuracy for temporal epoch using BW4 data, compared to decoding accuracy using BW12 data. P = 0.32 (t(5) = 1.09), unpaired t-test; matched population size = 47 cells, n = 4 and 3 animals for BW4 and BW12 respectively. Right, decoding accuracy for wall colour from the BW4 experiment, compared to decoding accuracy using matched data from BW12 experiment. P = 0.35 (t(5) = 1.03), unpaired t-test; matched population size = 47 cells, n = 4 and 3 animals for BW4 and BW12 respectively. h, Decoding accuracy for temporal epoch using BW4 data from the LEC, MEC, CA3, CA2 and CA1. One-way ANOVA, F(4) = 20.78, P < 1 × 10−5, post hoc Bonferroni multiple comparisons test, P < 0.005 (for each comparison against LEC); matched population size = 24 cells, n = 7, 2, 7, 3 and 3 animals for LEC, MEC, CA3, CA2 and CA1 respectively. i, Relation between population size and decoding accuracy for the LEC, CA3 and MEC. Left, decoding accuracy for varying population sizes; each line indicates the curve fit to data (shown as points) from one animal, with colours indicating recording area (n = 3, 3 and 2 animals for LEC, CA3 and MEC, respectively). Right, relation between population size and decoding accuracy for the LEC and CA3, pooled across animals from BW12 and BW4 experiments (data pooled from n = 7 animals for both LEC and CA3). j, Decoding accuracy for wall colour from trial period activity alone for the LEC, CA3 and MEC. P = 0.47 (LEC versus CA3, t(4) = 0.81) unpaired t-test; matched population size = 28 cells, n = 3, 3 and 2 animals for LEC, CA3 and MEC, respectively. k, Decoding accuracy for wall colour using data that was shuffled in time. P = 0.74 (t(2) = 0.38), two-tailed paired t-test; n = 3 animals. l, Decoding accuracy for wall colour using a subpopulation with all wall-colour-selective cells removed, compared to size-matched populations that were randomly drawn from full population. P = 0.12 (t(2) = 2.66), paired t-test; n = 3 animals with 77, 126 and 195 cells not selective for wall colour. For f, g, j–l, circles indicate individual animals, solid lines indicate mean decoding accuracy ± s.e.m., dashed lines indicate chance levels.
Extended Data Fig. 5 Evolution of LEC population dynamics.
Structure and evolution of LEC (purple, n = 3 animals), CA3 (blue, n = 3 animals) or MEC (gold, n = 2 animals) activity, quantified by applying several distance measurements to population activity states (see Supplementary Information). The distance being measured is illustrated in cartoon form above each plot (d1: distance between time bin 1 and time bin 2, d2: distance between time bin 2 and time bin 3, and so on). a, Manhattan distance between the population state for the middle time bin of a trial or intertrial period and all other time bins within that period. b, Manhattan distance between the population state for the last time bin of the preceding trial/intertrial period and all time bins within the current intertrial/trial period. c, Left, Manhattan distance between the population state for the first time bin of a trial or intertrial period and all subsequent time bins within that period. Right, fitting distance using an autoregressive (AR) model. Red line indicates model fit. d, Manhattan distance between consecutive pairs of population states across trial or intertrial periods. e, Pairwise angles, measured across consecutive points in time along neural trajectories during trial or intertrial periods as the angle between two vectors, each defined as the difference of consecutive population states. f, Manhattan distance between the overall mean population state for the first trial or intertrial period and the overall mean population states of all subsequent trial or intertrial periods. Solid lines indicate mean distance across all animals, shaded area indicates s.e.m., distance measures were all z-scored.
Extended Data Fig. 6 Decoding shortened temporal epochs.
a, Decoding accuracies for different temporal epoch lengths across the LEC, MEC, CA3, CA2 and CA1 using BW4 data (n = 7, 2, 7, 3 and 3 animals for LEC, MEC, CA3, CA2 and CA1, respectively). Decoding accuracy for 20-s epochs using LEC data was significantly better than all hippocampal areas. P < 1 × 10−5 (F(17) = 20.69) one-way ANOVA, post hoc Bonferroni multiple comparisons test; P < 0.05 (LEC versus MEC), P < 0.001 (LEC versus CA3, CA2 or CA1); matched population size = 24 cells. Decoding accuracy was higher for the LEC and MEC than for the CA3 for 10-s epochs. P < 0.001 (LEC versus CA3, all other comparisons were not significant, F(17) = 8.00), one-way ANOVA, post hoc Bonferroni multiple comparisons test; matched population size = 24 cells. Decoding accuracy was similar across all areas for 1-s epochs. P = 0.05 (F(17) = 2.95), one-way ANOVA. b, Confusion matrix from an example LEC animal for 10-s epochs. The matrix contains 468 epochs (each trial period of 250 s divided into 25 10-s epochs, each intertrial period of 140 s divided into 14 10-s epochs, 39 epochs per trial/intertrial pair, 12 total pairs across the session, giving 468 epochs). Left, confusion matrix for the entire session. Right, sum across trial/intertrial sections of the whole session matrix (outlined by dashed line in whole session matrix). c, Decoding accuracy for re-binned confusion matrices, see Supplementary Information (circles and solid lines with black indicates mean, shade indicates the same individual; P < 1 × 10−4 (comparing across epoch lengths, F(3) = 32.28) one-way ANOVA; P = 0.12 (trial versus 20 s), P < 0.001 (trial versus 10 s, and trial versus 1 s)) compared to decoding accuracy for re-binned confusion matrices following shuffling along columns, with diagonals preserved (triangles and dashed lines, with black indicating mean and shade indicating individual animals; comparing decoding accuracies from shuffled and unshuffled confusion matrices: two sided paired t-tests, for 20 s: t(2) = 7.42, P < 0.05; for 10 s: t(2) = 14.31, P < 0.01; for 1 s: t(2) = 4.46, P < 0.05). Grey dashed line indicates chance at 4.2%, shade indicates individual animals. d, Decoding accuracy for different temporal epoch lengths for the LEC (matched population size = 90 cells, n = 3 animals), based only on activity within individual trial or intertrial periods (top or bottom, respectively). Decoding accuracy was measured for each trial or intertrial period individually, and then the average was taken across all trials or intertrial periods, respectively, for each animal. Temporal bin size was: 1 s for 20-s-long epochs, 500 ms for 10-s-long epochs, and 50 ms for 1-s-long epochs. Chance levels for trial periods were 8.3%, 4.0% and 0.4%, and 14.3%, 7.1% and 0.7% for intertrial periods. e, Confusion matrices from an example LEC animal for single-trial or intertrial periods as in d, using 20-s (top) and 10-s (bottom) epochs, with intertrial results on the left and trial results on the right. For a and d, circles indicate individual animals, solid lines indicate mean decoding accuracies ± s.e.m., dashed lines indicate chance levels.
Extended Data Fig. 7 Temporal coding within a fixed environmental context.
a, Experimental design: 10 min without object, 10 min with object, and 10 min without object. b, Predictors used for object GLM: object presence, trial time and session time (24.0% selective for session time, 4.8% selective for trial time, 4.3% selective for a mixture of trial time and session time, percentages averaged across all animals). c, Explained variance for all LEC cells fit by GLM for object experiments (n = 150 cells). Average explained variance was 0.05. d, Distribution of selectivity for time, object and mixtures of time and object for single LEC cells (n = 3 animals with 56, 57 and 150 cells). Shade indicates the same individual across the different variables. e, Examples of GLM fit results for 12 cells from the object experiment with selectivity for different features. The firing rate of each cell is shown in grey, with the model-predicted firing rate in blue. The R2 value is shown for each cell. f, 2D projection of LEC neural population responses during object experiment from example animal. g, Decoding accuracy for trial identity or object presence (n = 3 animals). h, Decoding accuracies for temporal epochs of shortened length (n = 3 animals, matched population size = 56 cells). i, Decoding accuracy for trial identity (left) or object presence (right), with cells selective for decoded variable removed, compared to size-matched populations randomly drawn from full population. Trial identity: P < 0.05 (t(2) = 4.95), paired t-test; n = 29, 26 and 79 cells not selective for time; object presence: P < 0.05 (t(2) = 7.17), paired t-test; n = 39, 47, and 110 cells not selective for object presence. Circles indicate individual animals, solid lines indicate mean ± s.e.m. of described measurement, dashed lines indicate chance levels.
Extended Data Fig. 8 Explicit versus inherent mechanisms for temporal coding.
a, Top, a series of experiences occurs, each containing different event content and spanning different amounts of time. Bottom, two different ways in which temporal information within this series of experiences may be encoded. For both cases, a population code is used, but this may just as easily be replaced by a rate code within single cells. An explicit mechanism (left) purposely represents the passage of time, such that each chunk of time is represented equally. Thus, two experiences with the same temporal length but differing numbers of events would correspond to the same change in activity. An inherent mechanism (right) encodes temporal information entirely by representing the events within each experience. Thus, two experiences with the same temporal length but differing numbers of events would correspond to differing changes in activity. In either case, the high dimensionality of the representations would allow temporal information to be read out easily by downstream readout neurons, for example, cells in the hippocampus. b, As in a, but in this example, instead of a series of different experiences, the same experience is repeated three times (analogous to performing a learned task three times). Here, an explicit mechanism for temporal coding would exhibit the same amount of change in activity as in a, whereas an inherent mechanism would exhibit considerably reduced differences in activity across the experiences.
Extended Data Fig. 9 Decoding trial identity with additional matched data.
a, Euclidean distance between mean LEC population states for pairs of adjacent trials with different or same wall colour. P < 10−9 (t(31) = 8.81), unpaired t-test; n = 12 and 21 for same colour (BB/WW) and different colour (BW/WB) transitions, respectively, pooled from three animals. b, Decoding accuracy for trial identity during the figure-eight experiment, compared to decoding accuracy for temporal epoch using matched data from BW experiments that did not account for intertrial intervals or trial type. P < 0.01 (t(8) = 4.91), unpaired t-test; matched population size = 31 cells, n = 3 and 7 animals for figure-eight and BW, respectively. c, Circular-track task, in which animals alternated between clockwise and anticlockwise runs for 15 consecutive back and forth laps. Black circle indicates midpoint of the track. d, Left, 2D projection of the LEC neural population response during circular track experiment from example animal. Right, 2D projection of the LEC neural population response during matched periods from BW experiments. e, Left, decoding accuracy for trial identity during circular track experiment compared to decoding accuracy for temporal epoch using matched data from BW experiments. P < 0.0001 (t(7) = 6.21), unpaired t-test; matched population size = 47 cells, n = 2 and 7 animals for circular track and BW, respectively. Right, as in the left panel, but using temporally consecutive data that did not account for intertrial intervals or running direction. P < 0.05 (t(7) = 3.17), unpaired t-test; matched population size = 47 cells, n = 2 and 7 animals for circular track and BW, respectively. f, Confusion matrix for decoding trial identity in the circular track experiment from example animal. Circles indicate individual animals, solid lines indicate mean ± s.e.m. of described measurement, dashed lines indicate chance levels.
Extended Data Fig. 10 Additional characterization of LEC activity during figure-eight task.
a, Manhattan distance between consecutive pairs of population states (binned in 500-ms time bins) across single trials, averaged across all animals. b, Predictors used for the figure-eight GLM: trial type, trial time and session time (1.3% selective for session time, 9.3% selective for trial time, 6.5% selective for a mixture of trial time and session time, percentages averaged across all animals). c, Explained variance for all LEC cells fit by GLM for the figure-eight experiment (n = 149 cells). Average explained variance was 0.10. d, Distribution of selectivity for time, trial type and a mixture of time and trial type for single LEC cells, determined using a GLM (n = 3 animals with 72, 76 and 31 cells). Circles indicate individual animals, solid lines indicate mean fraction of cells ± s.e.m., shade indicates the same individual across the different variables. e, Examples of GLM fit results for six cells from the figure-eight experiment with selectivity for different features. The firing rate of each cell is shown in grey, with the model-predicted firing rate in blue. R2 value is shown for each cell. f, LEC activity for 12 example cells during the figure-eight task. Each plot shows the mean firing rate (top, 95% percentile confidence interval shaded), and peristimulus time histograms for left-turn (middle) and right-turn trials (bottom), with time centred on the point at which the animal reaches the base of the central stem on each trial. Cells 7–12 exhibited similar firing patterns for both left- and right-turn trials, including during the first 3 s of the trial, in which the animal occupied a different spatial location for left- versus right-turn trials. Such activity may be used for temporal information. Cells 13–18 exhibited highly divergent firing patterns for left- and right-turn trials, which may reflect the animal’s spatial location, the behavioural context of the trial, or a combination of the two variables. All example cells exhibited relatively stable firing across trials, a common feature observed during the figure-eight task.
Supplementary information
Supplementary Information
This file contains Supplemental materials and methods 1-6.
Rights and permissions
About this article
Cite this article
Tsao, A., Sugar, J., Lu, L. et al. Integrating time from experience in the lateral entorhinal cortex. Nature 561, 57–62 (2018). https://doi.org/10.1038/s41586-018-0459-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0459-6
Keywords
This article is cited by
-
Sleep restores an optimal computational regime in cortical networks
Nature Neuroscience (2024)
-
A generative model of memory construction and consolidation
Nature Human Behaviour (2024)
-
Lateral entorhinal cortex subpopulations represent experiential epochs surrounding reward
Nature Neuroscience (2024)
-
Spontaneous persistent activity and inactivity in vivo reveals differential cortico-entorhinal functional connectivity
Nature Communications (2024)
-
Minute-scale oscillatory sequences in medial entorhinal cortex
Nature (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.