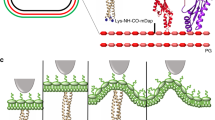
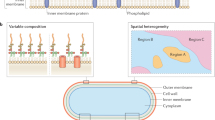
Abstract
Gram-negative bacteria possess a complex cell envelope that consists of a plasma membrane, a peptidoglycan cell wall and an outer membrane. The envelope is a selective chemical barrier1 that defines cell shape2 and allows the cell to sustain large mechanical loads such as turgor pressure3. It is widely believed that the covalently cross-linked cell wall underpins the mechanical properties of the envelope4,5. Here we show that the stiffness and strength of Escherichia coli cells are largely due to the outer membrane. Compromising the outer membrane, either chemically or genetically, greatly increased deformation of the cell envelope in response to stretching, bending and indentation forces, and induced increased levels of cell lysis upon mechanical perturbation and during L-form proliferation. Both lipopolysaccharides and proteins contributed to the stiffness of the outer membrane. These findings overturn the prevailing dogma that the cell wall is the dominant mechanical element within Gram-negative bacteria, instead demonstrating that the outer membrane can be stiffer than the cell wall, and that mechanical loads are often balanced between these structures.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Zgurskaya, H. I., Löpez, C. A. & Gnanakaran, S. Permeability barrier of Gram-negative cell envelopes and approaches to bypass it. ACS Infect. Dis. 1, 512–522 (2015).
Martin, H. & Frank, H. Quantitative bausteinanalyse der stützmembran in der zellwand von Escherichia coli B. Z. Naturforsch. B 17, 190–196 (1962).
Deng, Y., Sun, M. & Shaevitz, J. W. Direct measurement of cell wall stress stiffening and turgor pressure in live bacterial cells. Phys. Rev. Lett. 107, 158101 (2011).
Höltje, J. V. Growth of the stress-bearing and shape-maintaining murein sacculus of Escherichia coli. Microbiol. Mol. Biol. Rev. 62, 181–203 (1998).
Koch, A. L. Biophysics of bacterial walls viewed as stress-bearing fabric. Microbiol. Rev. 52, 337–353 (1988).
Herrmann, M., Schneck, E., Gutsmann, T., Brandenburg, K. & Tanaka, M. Bacterial lipopolysaccharides form physically cross-linked, two-dimensional gels in the presence of divalent cations. Soft Matter 11, 6037–6044 (2015).
Rassam, P. et al. Supramolecular assemblies underpin turnover of outer membrane proteins in bacteria. Nature 523, 333–336 (2015).
Ursell, T. S., Trepagnier, E. H., Huang, K. C. & Theriot, J. A. Analysis of surface protein expression reveals the growth pattern of the Gram-negative outer membrane. PLOS Comput. Biol. 8, e1002680 (2012).
Cayley, D. S., Guttman, H. J. & Record, M. T., Jr. Biophysical characterization of changes in amounts and activity of Escherichia coli cell and compartment water and turgor pressure in response to osmotic stress. Biophys. J. 78, 1748–1764 (2000).
Rojas, E., Theriot, J. A. & Huang, K. C. Response of Escherichia coli growth rate to osmotic shock. Proc. Natl Acad. Sci. USA 111, 7807–7812 (2014).
de Vries, H. Die Plasmolytische Studien über die Wand Vacuolen (G. Bernstein, Berlin, 1885).
Howatson, A. M. Engineering Tables and Data (Springer Science & Business Media, Berlin, 2012).
Tuson, H. H. et al. Measuring the stiffness of bacterial cells from growth rates in hydrogels of tunable elasticity. Mol. Microbiol. 84, 874–891 (2012).
Coughlin, R. T., Peterson, A. A., Haug, A., Pownall, H. J. & McGroarty, E. J. A pH titration study on the ionic bridging within lipopolysaccharide aggregates. Biochim. Biophys. Acta 821, 404–412 (1985).
Broady, K. W., Rietschel, E. T. & Lüderitz, O. The chemical structure of the lipid A component of lipopolysaccharides from Vibrio cholerae. Eur. J. Biochem. 115, 463–469 (1981).
Leive, L., Shovlin, V. K. & Mergenhagen, S. E. Physical, chemical, and immunological properties of lipopolysaccharide released from Escherichia coli by ethylenediaminetetraacetate. J. Biol. Chem. 243, 6384–6391 (1968).
Amro, N. A. et al. High-resolution atomic force microscopy studies of the Escherichia coli outer membrane: structural basis for permeability. Langmuir 16, 2789–2796 (2000).
Ruiz, N., Falcone, B., Kahne, D. & Silhavy, T. J. Chemical conditionality: a genetic strategy to probe organelle assembly. Cell 121, 307–317 (2005).
Sampson, B. A., Misra, R. & Benson, S. A. Identification and characterization of a new gene of Escherichia coli K-12 involved in outer membrane permeability. Genetics 122, 491–501 (1989).
Amir, A., Babaeipour, F., McIntosh, D. B., Nelson, D. R. & Jun, S. Bending forces plastically deform growing bacterial cell walls. Proc. Natl Acad. Sci. USA 111, 5778–5783 (2014).
Auer, G. K. et al. Mechanical genomics identifies diverse modulators of bacterial cell stiffness. Cell Syst. 2, 402–411 (2016).
Billings, G. et al. De novo morphogenesis in L-forms via geometric control of cell growth. Mol. Microbiol. 93, 883–896 (2014).
Yao, Z., Kahne, D. & Kishony, R. Distinct single-cell morphological dynamics under beta-lactam antibiotics. Mol. Cell 48, 705–712 (2012).
Kawai, Y., Mickiewicz, K. & Errington, J. Lysozyme counteracts β-lactam antibiotics by promoting the emergence of L-form bacteria. Cell 172, 1038–1049.e10 (2018).
Rick, P. D., Hubbard, G. L. & Barr, K. Role of the rfe gene in the synthesis of the O8 antigen in Escherichia coli K-12. J. Bacteriol. 176, 2877–2884 (1994).
Edelstein, A., Amodaj, N., Hoover, K., Vale, R. & Stuurman, N. Computer control of microscopes using μManager. Curr. Protoc. Mol. Biol. 92, 14.20.1–14.20.17 (2010).
Desmarais, S. M. et al. High-throughput, highly sensitive analyses of bacterial morphogenesis using ultra performance liquid chromatography. J. Biol. Chem. 290, 31090–31100 (2015).
Glauner, B., Höltje, J. V. & Schwarz, U. The composition of the murein of Escherichia coli. J. Biol. Chem. 263, 10088–10095 (1988).
Glauner, B. Separation and quantification of muropeptides with high-performance liquid chromatography. Anal. Biochem. 172, 451–464 (1988).
Hutter, J. L. & Bechhoefer, J. Calibration of atomic-force microscope tips. Rev. Sci. Instrum. 64, 1868–1873 (1993).
Acknowledgements
The authors thank members of the Huang, Theriot, and Weibel laboratories for discussions, K. Amberg-Johnson for assistance with immunoblotting, and T. Silhavy, T. Bernhardt, A. Maurelli, B. Hammer and H. Arjes for feedback, antibodies and strains. This work was supported by National Institutes of Health (NIH) Director’s New Innovator Awards DP2OD006466 (to K.C.H.) and DP2OD008735 (to D.B.W.), National Science Foundation (NSF) CAREER Award MCB-1149328 (to K.C.H.), the Stanford Systems Biology Center funded by NIH grant P50 GM107615 (to K.C.H. and J.A.T.), NIH Grant R37-AI036929 (to J.A.T.), the Howard Hughes Medical Institute (to J.A.T.), and NSF Grant DMR-1121288 (to D.B.W.). K.C.H. is a Chan Zuckerberg Biohub Investigator. E.R.R. was supported by a postdoctoral fellowship from the Simbios Center for Physics Based Computation at Stanford University under NIH Grant U54 GM072970. P.D.O. was supported by a postdoctoral fellowship from the Swiss National Science Foundation under Grant P2ELP3_172318. This work was also supported in part by the NSF under Grant PHYS-1066293, the hospitality of the Aspen Center for Physics and by the Allen Discovery Center program through The Paul G. Allen Frontiers Group.
Reviewer information
Nature thanks J.-F. Collet, W. Margolin, N. Minc and A. Rutenberg for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
E.R.R. conceptualized the study. E.R.R., G.B., P.D.O., G.K.A., A.M., F.C., D.B.W., J.A.T. and K.C.H. designed the experiments. E.R.R. and G.B. performed detergent treatments, oscillatory osmotic shock assays and protein release assays. G.B. performed cloning. E.R.R. and P.D.O. performed AFM experiments. G.K.A. performed microfluidic bending assays. E.R.R., G.B. and L.Z. performed L-form experiments. A.M. performed ultra performance liquid chromatography experiments. E.R.R., G.B., P.D.O., A.M. and G.K.A. analysed the data. E.R.R., G.B., G.K.A., P.D.O., A.M., J.A.T. and K.C.H. wrote the manuscript. All authors reviewed the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Cell-wall deformation is approximately linear with respect to hyperosmotic shock over a large range.
Population-averaged contraction of cell-wall lengths versus hyperosmotic shock magnitude (n = 92, 73, 53, 71, 58, 11, 31, 47 cells). Data are mean ± s.d. The dotted line is the linear best fit for experimental data for shocks with magnitude ≤ 800 mM. The plateau after 800 mM demonstrates that the cell envelope has reached its minimum length upon large, 3 M hyperosmotic shocks.
Extended Data Fig. 2 Detergent or EDTA treatment causes dissolution of the outer membrane.
a, Description of the outer-membrane release assay used to measure the effect of detergent and EDTA on the outer membrane. See Methods section ‘Outer membrane release assay’ for a more detailed description. b, Absorbance of 410-nm light (left axis) in the supernatant from detergent-treated cell suspensions after performing the LAL LPS detection assay. Absorbance is correlated with the amount of LPS in the sample, and is linear below 1,000 EU ml−1 (right axis). This experiment was performed once. c, Immunoblot with an antibody cross-reactive with several outer-membrane proteins (for example, LamB and OmpA) of the supernatant of cells treated with various concentrations of N-lauroyl sarcosine (detergent). This experiment was performed once. d, Absorbance of 410-nm light (left axis) in the supernatant from EDTA-treated cell suspensions after performing the LAL LPS detection assay. Absorbance is correlated with the amount of LPS in the sample, and is linear below 1,000 EU ml−1 (right axis). This experiment was performed once. e, Montage of a representative plasmolysed cell expressing cytosolic GFP lysing upon detergent treatment. Cell expansion corresponds to the time of GFP disappearance. Data are mean ± s.d. from 32 cells in one experiment. f, Duration of swelling (left), of release of cytosolic GFP (centre), and of release of ribosomes upon lysis of plasmolysed cells (n = 22, 32, 12 cells, respectively; one experiment each). Error bars, ±1 s.d. The duration of swelling is approximately equal to the duration of the release of cytoplasmic contents. g, Ultra performance liquid chromatography of peptidoglycan composition of cell walls purified from untreated (red), sorbitol-treated (yellow), detergent-treated (green), and EDTA-treated (blue) cells. Mean glycan strain lengths (i) and percentage of various peptidoglycan subunit eluents (ii) are the same across all treatments. Dap–Dap, Dimers formed with double diaminopimelic acid bonds as opposed to diaminopimelic acid–d-alanine bonds; lipoprotein, subunits bound to Lpp. n = 4, 2, 2 and 1 sacculi preparations for untreated, sorbitol-treated, detergent-treated and EDTA-treated cells, respectively.
Extended Data Fig. 3 Detergent treatment causes contraction of the cell wall across a range of conditions and organisms.
a, Length of the cell wall versus time during hyperosmotic shock (3 M sorbitol, solid arrow) and subsequent treatment with detergent (5% N-lauroyl sarcosine, dotted arrow), and washout of the detergent (switching back to 3 M sorbitol, dashed arrow) for four representative cells (n = 260 cells). The cells briefly swelled upon washout, then relaxed to the pre-washout rest length. We speculate that the brief swelling is caused by a hypoosmotic shock during washout (lowering the concentration of detergent). That is, we propose that the detergent has a slow diffusion constant through the cell wall, and therefore washing it out imposes a transient hypoosmotic shock, before detergent within the cell diffuses out of it. When it does, the cell wall relaxes again to its rest state, demonstrating that cell-wall contraction upon detergent treatment is not due to compaction by the detergent. b, Lengths of the cell wall versus time during hyperosmotic shock (3 M sorbitol, solid arrow) and subsequent addition of detergent (5% N-lauroyl sarcosine, dotted arrow) for four representative B. subtilis cell chains (n = 112 cell chains). Rather than contract upon detergent treatment, the cell walls re-extend. We hypothesize that the cell wall is in a highly compressed state after 3-M hyperosmotic shock because the cytoplasm is exerting an inward force via connections between the plasma membrane and cell wall. When the plasma membrane is dissolved with detergent, this force is relieved and the cell wall extends to its rest length. c, Lengths of the cell wall of B. subtilis cell chains versus time during a 1-M oscillatory osmotic shock, which caused cell lysis (for example, red arrows), and treatment with detergent (5% N-lauroyl sarcosine, dotted arrow; n = 93 cell chains). The cell walls expanded only slightly upon detergent addition, indicating that it is lysis rather than detergent treatment that causes the re-extension in (b). d, Contraction upon plasmolysis, subsequent lysis and total contraction for three environmental conditions: i) LB (n = 79, 56 and 56 cells, respectively), ii) M9 (n = 46, 52 and 38 cells, respectively) and iii) cultured in LB and transferred to PBS before plasmolysis (n = 95, 86 and 94 cells, respectively). Error bars indicate ±1 s.d. e, Contraction upon plasmolysis, subsequent lysis and total contraction for three wild-type E. coli strains: i) MG1655 (n = 79, 56 and 56 cells, respectively), ii) MC4100 (n = 48, 48 and 48, respectively) and iii) AB1133 (n = 56, 56 and 56 cells, respectively). Error bars indicate ±1 s.d. f, Contraction upon plasmolysis, subsequent lysis, and total contraction for i) P. aeruginosa (n = 12, 12 and 12 cells, respectively) and ii) V. cholerae (n = 36, 36 and 36 cells, respectively). Error bars indicate ±1 s.d. g, The mean ratio between the rest length of the outer membrane, lom, and the length of the cell envelope in the fully turgid state, l1, for three wild-type E. coli strains (n = 12, 10 and 10 cells for MG155, MC4100 and AB1133, respectively). The rest length of the outer membrane is approximately equal to the length of the fully turgid cell for all three strains. Error bars indicate ±1 s.d. The P value was calculated using a Student’s two-sided t-test. h, Mean contraction upon plasmolysis, subsequent lysis, and total contraction for wild-type AB1133 (n = 56 cells) and an isogenic strain expressing the O8 antigen (n = 55 cells). The ratio for cells expressing the O8 antigen is very large because the contraction upon lysis for the parental wild-type strain (AB1133) was close to zero, and adding the antigen markedly increased contraction. Error bars indicate ±1 s.d. P values were calculated using a Student’s two-sided t-test.
Extended Data Fig. 4 Impermeable molecules within the cell wall after lysis cause residual turgor pressure.
a, Increasing detergent concentration caused an approximately proportional increase in the mean cell-wall contraction upon lysis and the mean total contraction in E. coli MG1655 cells. Each point represents one experiment. The number of cells for each experiment is given in Supplementary Table 2. b, Representative DAPI-stained E. coli cells before (left) and after (right) lysis shown in phase contrast and epifluorescence, demonstrating that DNA is not retained in the lysed cells. The experiment was performed once. c, Representative E. coli cells expressing a fluorescent protein fusion to the S2 ribosomal protein before (left) and after (right) lysis shown in phase contrast and epifluorescence, demonstrating that ribosomes are not retained in lysed cells. The experiment was performed once. d, Representative E. coli cells expressing cytosolic GFP before (left) and after (right) lysis shown in phase contrast and epifluorescence, demonstrating that GFP is not retained in lysed cells. The experiment was performed once. e, Representative E. coli cells expressing a fluorescently tagged version of MreB after lysis shown in phase contrast (left), epifluorescence (centre) and in overlay (right), demonstrating that MreB is retained within most lysed cells, but that cells with weak phase density after lysis retain low levels of MreB (arrows). f, Cumulative fluorescence intensity of MreB–sfGFP versus the average phase-contrast intensity within the cell after lysis (n = 162 cells). Cells with higher phase density have lower intensity. g, Cell-wall contraction upon lysis versus average phase-contrast intensity within the cell after lysis (n = 46 cells). h, Total contraction during plasmolysis and lysis versus average phase-contrast intensity within the cell after lysis (n = 46 cells).
Extended Data Fig. 5 EDTA weakens the E. coli cell envelope.
a, Length of the cell wall versus time for 7 representative B. subtilis cell chains during treatment with detergent with treatment using detergent and 10 mM EDTA (n = 68 cell chains). Although detergent caused lysis, subsequent addition of EDTA did not affect cell-wall rest length. b, Lengths of the cell wall of B. subtilis cell chains versus time during 1-M oscillatory osmotic shocks, which caused cell lysis (for example, red arrows), with treatment using 10 mM EDTA (dashed arrow; n = 127 cell chains). EDTA did not affect the rest length of the cell walls. c, Top, length of the B. subtilis cell wall versus time through a hyperosmotic shock (3 M sorbitol, solid arrow) and subsequent treatment with 10 mM EDTA (dotted arrow) for four representative cell chains (n = 61 cell chains). Cell-wall length did not decrease after detergent, as it did for E. coli. Bottom, micrograph of a two-cell chain expressing cytosolic (strain HA405, left) and a kymograph showing fluorescence intensity along the dotted red line during the experiment in the top graph. The cell chain in the kymograph corresponds to the bottom-most length trace in the top graph. Red arrows demonstrate that the discrete increases in length observed after EDTA treatment correspond to cell-lysis events, when fluorescence within the cells begin to decrease. d, Population-averaged E. coli cell-wall length contraction upon EDTA application after plasmolysis increased with increasing concentration of EDTA (n = 131, 193, 225, 138, 81, 94, 36, 28, 72 cells, respectively). Error bars indicate ±1 s.d. e, Length of the cell wall versus time during hyperosmotic shock (3 M sorbitol, solid arrow) and subsequent treatment with 10 mM EDTA and 50 mM MgCl2 (dotted arrow) for representative E. coli cells (n = 91 cells). f, Length of the cell walls of representative E. coli cells during 100-mM oscillatory shocks with 2-min period (n = 243 cells). g, Length of the cell walls of representative E. coli cells during a 100-mM oscillatory shock with 2-min period and 10 mM EDTA (n = 284 cells). h, Population-averaged elongation rate of the E. coli cell wall during 100-mM oscillatory shocks with 2-min period for untreated (black line) and 10 mM EDTA-treated cells (n = 284 cells). i, Effective population-averaged cell length (leff), calculated by integrating the population averaged elongation rate in h during 100-mM oscillatory shocks with 2-min period for untreated (black line) and 10 mM EDTA-treated cells (n = 284 cells). Dotted lines are the respective time-averaged leff using a rolling-window averaging filter with a 2-min window (equal to the period of oscillations). j, Deviation of the effective population-averaged length in i from the respective time-averaged trace. k, The mean amplitude of oscillation was found by averaging the peak-to-peak amplitude in j over cycles (n = 10 cycles). Error bars indicate ±1 s.d. The P value was calculated using a Student’s two-sided t-test. l, Amplitude of cell-wall length oscillations (ratio with respect to the untreated wild type; Fig. 2j) versus cell-wall stiffness calculated from plasmolysis–lysis experiments (ratio with respect to the untreated wild type; Fig. 2g). Solid line, linear best fit for only perturbations to the outer membrane (red circles; linear regression: R2 = 0.71, F = 9.7, P = 0.0356). Dashed line, best fit when additionally considering perturbations to protein linkages between the outer membrane and cell wall (dashed circles; linear regression: R2 = 0.25, F = 1.4, not significantly different from horizontal). For the O8-expressing strain, we conservatively used a stiffness ratio of 1.5 for the fits.
Extended Data Fig. 6 FM 4-64 softens E. coli cells.
a, Cantilever force versus indentation distance during successively increasing EDTA concentrations, as in Fig. 3a. Lines indicate the average force–distance curves taken during the period in which the cell was treated with the given concentration of EDTA. Shaded areas indicate ±1 s.d (n = 4 cells). Stiffness was measured every minute during the 5–10 min periods when the cell was treated with each EDTA concentration. Average-force curves were registered with respect to the onset of force increase as the cantilever was lowered. b, Stiffness of a representative cell versus time, as measured with AFM. At t = 10 min the cell was treated with 2 μg ml−1 FM 4-64 (n = 3 cells). c, The ratio of the cell stiffness computed with AFM (Fig. 3c) versus the ratio of envelope stiffness computed from plasmolysis–lysis experiments (Fig. 2g) across chemical and genetic perturbations. The solid line is the linear best fit (linear regression: R2 = 0.27, F = 1.4, not significantly different from horizontal). The numbers of cells used for each measurement are the same as given for Fig. 3c (AFM measurements of cell stiffness) and Fig. 2d–f (envelope stiffness). For the O8-expressing strain, we conservatively used a stiffness ratio of 1.5 for the fit. d, Calibration of AFM measurements using a polydimethylsiloxane sample with known Young’s modulus of 3.5 MPa. Top, distribution of measurements of Young’s modulus across the calibration sample. Red curve is the Gaussian fit to the data. Dashed line is the mean. Top inset, Young’s modulus measurements were spatially uniform. Bottom, box plot of the distribution of Young’s modulus measurements showing the median stiffness (red line), 25% and 75% percentiles (edges of box), extreme bounds (whiskers), and outliers (red points).
Extended Data Fig. 7 Genetic perturbations to the outer membrane render cells vulnerable to mechanical perturbation.
a, Cell lysis versus time and cycle number during application of 5% detergent (N-lauroyl sarcosine, n = 112 cells) and 5% detergent with 400-mM oscillatory osmotic shock (n = 100 cells). Cell-lysis rate increased notably during 400-mM oscillatory osmotic shocks. b, Cell lysis versus time and cycle number during 400-mM oscillatory osmotic shocks with 2-min period for mutant E. coli strains with the MC4100 wild-type background (n = 476, 98, 187, 99, 280 cells for the wild-type, imp4213, ∆ompA, ∆lpp and ∆pal groups, respectively). c, Cell lysis versus time and cycle number during 400-mM oscillatory osmotic shock with 2-min period for E. coli wild-type AB1133 (n = 273 cells) and ATM378 (AB1133+O8, n = 123 cells). d, Time at which 25% of cells had lysed (ratio to the untreated wild type, Fig. 4b) versus the ratio of amplitudes during 100-mM oscillatory osmotic shocks (Fig. 2j). The line is the linear best fit (linear regression; R2 = 0.84, F = 19.4, P = 0.0045). e, Serial dilutions of overnight E. coli L-form cultures spotted onto solid media permissive for L-form growth. For chemical treatments, 10 mM EDTA and 2 μg ml−1 FM 4-64 were included in the liquid media used to culture L-forms overnight, but not in the solid media onto which L-forms were spotted. Mutants (ΔompA, +O8, Δlpp, Δpal) formed very small colonies, and were thus viewed using an inverted microscope to ensure accurate counting.
Supplementary information
Supplementary Information
This file contains Supplementary Text which contains additional information regarding E. coli envelope behaviour after detergent treatment, effects of detergent and EDTA on Bacillus subtilis cells, and correlations between outer membrane stiffness and other variables. Supplemental Tables S1 and S2 show the bacterial strains used in this study and number of cells for experiments in Extended Data Fig. 4a, respectively.
Video 1: Detergent treatment after plasmolysis causes further contraction of the cell wall.
Phase contrast (left) and epifluorescence (525 nm emission, center; 630 nm, right) of E. coli cells whose cell walls were stained with fluorescent WGA (center) and whose outer membranes were stained with FM 4-64 (right). The cells, which were growing in LB medium, were first subjected to a large hyperosmotic shock by exposing them to LB + 3 M sorbitol, after which their plasma and outer membranes were removed by exposing them to LB + 3 M sorbitol + 5% N-lauroyl sarcosine (detergent). Cell-wall lengths shrank substantially after detergent treatment.
Video 2: Detergent dissolves the outer membrane.
E. coli cells that had been stained with FM 4-64 (outer membrane) and fluorescent WGA (cell wall) were subjected to a 3-M hyperosmotic shock, then treated with lysozyme, causing spheroplasting in many cells. The spheroplasts were then treated with detergent (5% N-lauroyl sarcosine), whereupon the outer membrane completely dissolved.
Video 3: Oscillatory osmotic shocks probe outer membrane stiffness.
E. coli cells in LB under oscillatory 100-mM hyperosmotic shocks with 2-min period.
Video 4: Outer membrane perturbation causes accelerated lysis of E. coli cells under oscillatory osmotic shock.
E. coli cells under oscillatory 400-mM shocks (between LB and LB + 400 mM sorbitol) with 2-min period. Before the shocks were initiated, the cells were first imaged in LB for 5 min. In the left panel, the media were cycled between PBS and PBS + 200 mM sorbitol. In the right panel, 10 mM EDTA was included in both media.
Video 5: imp4213 cells lyse during L-form generation.
E. coli MG1655 imp4213 cells grown on LFLB agarose pads lyse immediately after the protoplast blebs out of the cell wall (arrows, n = 37 cells). Compare to Video S1 in Ref. 22, which shows wild-type L-form generation under similar conditions.
Rights and permissions
About this article
Cite this article
Rojas, E.R., Billings, G., Odermatt, P.D. et al. The outer membrane is an essential load-bearing element in Gram-negative bacteria. Nature 559, 617–621 (2018). https://doi.org/10.1038/s41586-018-0344-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0344-3
This article is cited by
-
Engineering Escherichia coli for increased Und-P availability leads to material improvements in glycan expression technology
Microbial Cell Factories (2024)
-
RNA-seq reveals multifaceted gene expression response to Fab production in Escherichia coli fed-batch processes with particular focus on ribosome stalling
Microbial Cell Factories (2024)
-
Prolonged versus intermittent β-lactam infusion in sepsis: a systematic review and meta-analysis of randomized controlled trials
Annals of Intensive Care (2024)
-
SlyB encapsulates outer membrane proteins in stress-induced lipid nanodomains
Nature (2024)
-
A genetic screen identifies a role for oprF in Pseudomonas aeruginosa biofilm stimulation by subinhibitory antibiotics
npj Biofilms and Microbiomes (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.