Abstract
Accurate replication of DNA requires stringent regulation to ensure genome integrity. In human cells, thousands of origins of replication are coordinately activated during S phase, and the velocity of replication forks is adjusted to fully replicate DNA in pace with the cell cycle1. Replication stress induces fork stalling and fuels genome instability2. The mechanistic basis of replication stress remains poorly understood despite its emerging role in promoting cancer2. Here we show that inhibition of poly(ADP-ribose) polymerase (PARP) increases the speed of fork elongation and does not cause fork stalling, which is in contrast to the accepted model in which inhibitors of PARP induce fork stalling and collapse3. Aberrant acceleration of fork progression by 40% above the normal velocity leads to DNA damage. Depletion of the treslin or MTBP proteins, which are involved in origin firing, also increases fork speed above the tolerated threshold, and induces the DNA damage response pathway. Mechanistically, we show that poly(ADP-ribosyl)ation (PARylation) and the PCNA interactor p21Cip1 (p21) are crucial modulators of fork progression. PARylation and p21 act as suppressors of fork speed in a coordinated regulatory network that is orchestrated by the PARP1 and p53 proteins. Moreover, at the fork level, PARylation acts as a sensor of replication stress. During PARP inhibition, DNA lesions that induce fork arrest and are normally resolved or repaired remain unrecognized by the replication machinery. Conceptually, our results show that accelerated replication fork progression represents a general mechanism that triggers replication stress and the DNA damage response. Our findings contribute to a better understanding of the mechanism of fork speed control, with implications for genomic (in)stability and rational cancer treatment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Conti, C. et al. Replication fork velocities at adjacent replication origins are coordinately modified during DNA replication in human cells. Mol. Biol. Cell 18, 3059–3067 (2007).
Halazonetis, T. D., Gorgoulis, V. G. & Bartek, J. An oncogene-induced DNA damage model for cancer development. Science 319, 1352–1355 (2008).
Bryant, H. E. et al. Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature 434, 913–917 (2005).
Luo, X. & Kraus, W. L. On PAR with PARP: cellular stress signaling through poly(ADP-ribose) and PARP-1. Genes Dev. 26, 417–432 (2012).
Schlacher, K., Wu, H. & Jasin, M. A distinct replication fork protection pathway connects Fanconi anemia tumor suppressors to RAD51-BRCA1/2. Cancer Cell 22, 106–116 (2012).
Farmer, H. et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434, 917–921 (2005).
Ledermann, J. et al. Olaparib maintenance therapy in patients with platinum-sensitive relapsed serous ovarian cancer: a preplanned retrospective analysis of outcomes by BRCA status in a randomised phase 2 trial. Lancet Oncol. 15, 852–861 (2014).
Bryant, H. E. et al. PARP is activated at stalled forks to mediate Mre11-dependent replication restart and recombination. EMBO J. 28, 2601–2615 (2009).
Burrell, R. A. et al. Replication stress links structural and numerical cancer chromosomal instability. Nature 494, 492–496 (2013).
Zhong, Y. et al. The level of origin firing inversely affects the rate of replication fork progression. J. Cell Biol. 201, 373–383 (2013).
Toledo, L. I. et al. ATR prohibits replication catastrophe by preventing global exhaustion of RPA. Cell 155, 1088–1103 (2013).
Duxin, J. P. et al. Okazaki fragment processing-independent role for human Dna2 enzyme during DNA replication. J. Biol. Chem. 287, 21980–21991 (2012).
Godon, C. et al. PARP inhibition versus PARP-1 silencing: different outcomes in terms of single-strand break repair and radiation susceptibility. Nucleic Acids Res. 36, 4454–4464 (2008).
el-Deiry, W. S. et al. WAF1, a potential mediator of p53 tumor suppression. Cell 75, 817–825 (1993).
Waga, S., Hannon, G. J., Beach, D. & Stillman, B. The p21 inhibitor of cyclin-dependent kinases controls DNA replication by interaction with PCNA. Nature 369, 574–578 (1994).
Lee, M. H., Na, H., Kim, E. J., Lee, H. W. & Lee, M. O. Poly(ADP-ribosyl)ation of p53 induces gene-specific transcriptional repression of MTA1. Oncogene 31, 5099–5107 (2012).
Frouin, I. et al. Human proliferating cell nuclear antigen, poly(ADP-ribose) polymerase-1, and p21waf1/cip1. A dynamic exchange of partners. J. Biol. Chem. 278, 39265–39268 (2003).
Madison, D. L. & Lundblad, J. R. C-terminal binding protein and poly(ADP)ribose polymerase 1 contribute to repression of the p21waf1/cip1 promoter. Oncogene 29, 6027–6039 (2010).
Yeo, C. Q. X. et al. p53 maintains genomic stability by preventing interference between transcription and replication. Cell Rep. 15, 132–146 (2016).
Chen, J., Jackson, P. K., Kirschner, M. W. & Dutta, A. Separate domains of p21 involved in the inhibition of Cdk kinase and PCNA. Nature 374, 386–388 (1995).
Luo, Y., Hurwitz, J. & Massagué, J. Cell-cycle inhibition by independent CDK and PCNA binding domains in p21Cip1. Nature 375, 159–161 (1995).
Mansilla, S. F. et al. Cyclin Kinase-independent role of p21CDKN1A in the promotion of nascent DNA elongation in unstressed cells. eLife 5, e18020 (2016).
Uhlen, M. et al. Towards a knowledge-based Human Protein Atlas. Nat. Biotechnol. 28, 1248–1250 (2010).
Moudry, P. et al. TOPBP1 regulates RAD51 phosphorylation and chromatin loading and determines PARP inhibitor sensitivity. J. Cell Biol. 212, 281–288 (2016).
Murai, J. et al. Trapping of PARP1 and PARP2 by clinical PARP inhibitors. Cancer Res. 72, 5588–5599 (2012).
Ray Chaudhuri, A. et al. Topoisomerase I poisoning results in PARP-mediated replication fork reversal. Nat. Struct. Mol. Biol. 19, 417–423 (2012).
Zellweger, R. et al. Rad51-mediated replication fork reversal is a global response to genotoxic treatments in human cells. J. Cell Biol. 208, 563–579 (2015).
Kunkel, T. A. DNA replication fidelity. J. Biol. Chem. 279, 16895–16898 (2004).
Flach, J. et al. Replication stress is a potent driver of functional decline in ageing haematopoietic stem cells. Nature 512, 198–202 (2014).
Maya-Mendoza, A., Olivares-Chauvet, P., Kohlmeier, F. & Jackson, D. A. Visualising chromosomal replication sites and replicons in mammalian cells. Methods 57, 140–148 (2012).
Sheu, Y. J., Kinney, J. B., Lengronne, A., Pasero, P. & Stillman, B. Domain within the helicase subunit Mcm4 integrates multiple kinase signals to control DNA replication initiation and fork progression. Proc. Natl Acad. Sci. USA 111, E1899–E1908 (2014).
Kohlmeier, F., Maya-Mendoza, A. & Jackson, D. A. EdU induces DNA damage response and cell death in mESC in culture. Chromosome Res. 21, 87–100 (2013).
Olive, P. L. & Banáth, J. P. The comet assay: a method to measure DNA damage in individual cells. Nat. Protocols 1, 23–29 (2006).
Pines, A., Mullenders, L. H., van Attikum, H. & Luijsterburg, M. S. Touching base with PARPs: moonlighting in the repair of UV lesions and double-strand breaks. Trends Biochem. Sci. 38, 321–330 (2013).
Sims, J. L., Berger, S. J. & Berger, N. A. Poly(ADP-ribose) polymerase inhibitors preserve nicotinamide adenine dinucleotide and adenosine 5′-triphosphate pools in DNA-damaged cells: mechanism of stimulation of unscheduled DNA synthesis. Biochemistry 22, 5188–5194 (1983).
Dutto, I. et al. p21CDKN1A regulates the binding of poly(ADP-ribose) polymerase-1 to DNA repair intermediates. PLoS ONE 11, e0146031 (2016).
Breslin, C. et al. The XRCC1 phosphate-binding pocket binds poly (ADP-ribose) and is required for XRCC1 function. Nucleic Acids Res. 43, 6934–6944 (2015).
Ray Chaudhuri, A., Ahuja, A. K., Herrador, R. & Lopes, M. Poly(ADP-ribosyl) glycohydrolase prevents the accumulation of unusual replication structures during unperturbed S phase. Mol. Cell. Biol. 35, 856–865 (2015).
Strzalka, W. & Ziemienowicz, A. Proliferating cell nuclear antigen (PCNA): a key factor in DNA replication and cell cycle regulation. Ann. Bot. 107, 1127–1140 (2011).
Acknowledgements
We thank W. Dunphy for the treslin antibody, D. Gomez-Cabello for help with Fig. 2j, the Danish Cancer Society, the Novo Nordisk Foundation, the Danish Council for Independent Research, the Swedish Research Council, the Grant Agency of the Czech Republic (17–14743S), the Czech Ministry of Education, Youth and Sports (NPU LO1304; EATRIS-CZ), and the Danish National Research Foundation (project CARD) for grant support.
Reviewer information
Nature thanks A. Vindigni and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
A.M.-M., P.M. and J.B. conceived the study and designed experiments. A.M.-M., P.M., J.M.M.-M., R.S. and M.H.L. performed experiments. A.M.-M., P.M., J.M.M.-M. and J.B. wrote the manuscript. All authors read and accepted the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Fork acceleration is PARPi dose- and time-dependent, and cell-type independent.
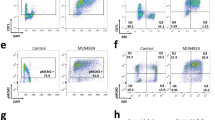
a, Cell cycle profiles of U2-OS cells treated with different concentrations of the PARPi olaparib (0.1, 1 or 10 µM) and BJ cells treated with 10 µM olaparib for 24 h; n = 3 biological replicates. b, Number of U2-OS cells treated with different concentrations of olaparib (0.1, 1 or 10 µM) for 72 h; n = 3 biological replicates. Drug was refreshed every 24 h. Data are mean ± s.d. c, Cell cycle profiles of U2-OS cells treated with increasing concentrations of the olaparib (10, 15 or 30 µM) for 24 h; n = 2 biological replicates. d, DNA fibres from U2-OS cells 24 h after treatment with increasing concentrations of olaparib (0.1, 1, 10, 15 or 30 µM). Scored forks: 0 µM PARPi = 503; 0.1 μM = 606; 1 μM = 406; 10 μM = 244; 15 μM = 372; 30 μM = 217; n = 2 biological replicates. Mean fork speed (kb min−1) is indicated next to each condition. e, CldU/IdU ratios calculated from values in d. Percentage of highly asymmetric forks (CldU/IdU ratios < 0.5 and > 1.5) is indicated next to each condition. f, DNA fibres from U2-OS cells treated with 10 µM olaparib for different periods of time (0.5, 1, 2, 4 or 48 h). Scored forks: 0 h = 744; 0.5 h = 450; 1 h = 379; 2 h = 314; 4 h = 465; 48 h = 589; n = 2 biological replicates. Mean fork speed is indicated. g, CldU/IdU ratios calculated from values in f. Percentage of highly asymmetric fork is indicated. h, DNA fibres from BJ cells treated with 10 µM olaparib for 24 h. Scored forks: 0 µM = 198; 10 μM = 317; n = 2 biological replicates. P values determined by two-tailed Welch’s t -test. i, DNA fibres from HeLa cells treated with 10 µM olaparib for 4 h. Scored forks: 0 µM = 285; 10 μM = 142; n = 2 technical replicates. j, DNA fibres from U2-OS cells 24 h after treatment with increasing concentrations of veliparib (1, 10 or 50 µM). Scored forks: 0 µM = 689; 1 μΜ = 408; 10 μM = 571; 50 μM = 408; n = 2 biological replicates. P value determined by two-tailed Welch’s t-test. k, CldU/IdU ratios calculated from values in j. Percentage of highly asymmetric forks is indicated above each condition. l, Cell cycle profiles of U2-OS cells treated with increasing concentrations of veliparib (1, 10 or 50 µM) for 24h, n = 2 biological replicates. For box plots in d–k, whiskers indicate the fifth and ninety-fifth percentiles, and the centre values depict the median.
Extended Data Fig. 2 PARPi induces accumulation of cells in mid-late S phase.
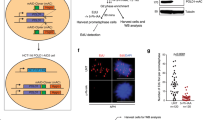
a, Representative images of DNA replication patterns in control U2-OS cells. Cells were pulse-labelled with 10 µM BrdU for 30 min. Scale bars, 5 µm. b, Outline of the experimental design of detailed DNA replication pattern analysis. U2-OS cells were labelled with BrdU (green) for 30 min, washed, chased for 4 h in fresh medium and labelled with EdU (red) for 30 min. Transition between replication patterns was classified as early–early (cells that did not leave early S phase during the experiment time), early–mid and mid–late (cells that progressed to the consecutive part of S phase) and early–late (cells that progressed fast through S phase). Scale bars, 10 µm. c, Representative images of double-labelled DNA replication patterns in U2-OS cells treated with DMSO (CT) or 10 µM olaparib (PARPi) for 24 h. Scale bars, 10 µm. d, Representative images of BrdU-positive nuclei from control and PARPi-treated U2-OS cells in early, mid and late S phase. Images were acquired using high-throughput microscopy. e, Percentage of U2-OS cells in early, mid and late S phase after treatment with DMSO or 10 µM olaparib for 24 h was quantified using high-throughput microscopy based on BrdU intensity versus DNA content. S phase patterns were gated as indicated. f, Distribution of S phase patterns in U2-OS cells treated with DMSO or 10 µM olaparib for 24 h (CT: early = 29%; mid = 60%, late = 11%; PARPi: early = 16%; mid = 40%, late = 44%; n = 2 biological replicates) (see Source Data).
Extended Data Fig. 3 PARPi induces DNA damage response.
a, Representative images of DDR markers analysed by high-throughput microscopy in U2-OS cells treated with DMSO or 10 µM olaparib (PARPi) for 24 h. b, Percentage of U2-OS cells with more than five RPA32, RAD51, γH2AX or 53BP1 foci after 24 h of PARPi treatment. Data are mean ± s.d., n = 3 biological replicates (see Source Data). c, Immunoblots of DDR proteins in U2-OS cells after 24 h of PARPi treatment. IMP-B is a loading control; n = 2 biological replicates. d, Representative images of the alkaline comet assay performed using U2-OS cells treated with DMSO, 10 µM olaparib, 2 mM hydroxyurea or 10 µM VP-16 for 24 h. Scale bars, 10 µm. e, Representative images of the TUNEL assay performed using pre-extracted BJ cells treated with DMSO, DNase I or 10 µM olaparib (left) along with U2-OS cells treated with DMSO or 10 µM olaparib (right). Scale bars, 10 µm.
Extended Data Fig. 4 Fork speed and origin activation.
a, Immunoblots of treslin knockdown efficiency in U2-OS cells, 72 h after transfection with three different siRNAs (si1–si3). n = 2 biological replicates. b, DNA fibres from non-targeting or treslin-knockdown U2-OS cells, 72 h after transfection with three different siRNA. Mean fork speed (kb min−1): NT = 0.96; si1 = 1.54; si2 = 1.58; si3 = 1.54. Scored forks: NT = 316; si1 = 367; si2 = 368; si3 = 358; n = 2 biological replicates. Data are mean ± s.d. P values determined by Welch’s two-tailed t-test (see Source Data). c, Immunoblots of MTBP knockdown efficiency in U2-OS cells, 72 h after transfection with two different siRNAs. Tubulin is a loading control; n = 2 biological replicates. d, Percentage of non-targeting, MTBP- or treslin-knockdown U2-OS cells with more than γH2AX foci. Data are mean ± s.d., n = 2 biological replicates. e, Cell cycle profiles of non-targeting or MTBP-knockdown U2-OS cells. Indicated cells were treated with 10 µM PARPi for 24 h; n = 2 biological replicates. f, DNA fibres from non-targeting or treslin-knockdown U2-OS cells. Indicated cells were treated as in e. Mean fork speed (kb min−1): NT = 1.0; PARPi = 1.74; treslin KD–PARPi = 1.72. Scored forks: NT = 410; PARPi = 395; treslin KD–PARPi = 424; n = 2 biological replicates. Data are mean ± s.d. P values determined by Welch’s two-tailed t-test. g, Immunoblots of the chromatin-associated fraction of treslin after 24 h of treatment with 10 µM olaparib. Histone H3 is a loading control; n = 2 biological replicates. h, Cell cycle profiles of non-targeting or treslin-knockdown U2-OS cells. Indicated cells were treated as in e; n = 2 biological replicates. i, Representative images of the origin-to-origin distance measurements in double-labelled DNA fibres from non-targeting or treslin-knockdown/ATR-inhibited U2-OS cells. Scale bars, 10 µm. j, Representative images of fork density from non-targeting or treslin-knockdown U2-OS cells. Indicated cells were treated with 10 µM PARPi for 24 h or 1 µM ATRi for 1 h before a 20-min 10 μM BrdU pulse. DNA (grey) and BrdU (red) were detected by immunofluorescence. Scale bars, 10 µm. k, Number of forks in a well-spread single DNA fibre, counted from multiple fibres per each condition and converted into number of forks per Mb: NT = 17; PARPi = 14; treslin-KD = 15; ATRi = 58; PARPi–ATRi = 59; treslin KD–ATRi = 22; n = 3 biological replicates. Whiskers indicate fifth and ninety-fifth percentiles and centre values depict the median. P values determined by Welch’s two-tailed t-test. l, Representative images of double-labelled DNA fibres from non-targeting or treslin-knockdown U2-OS cells. Indicated cells were treated with 10 µM PARPi for 24 h and/or 1 µM ATRi for 1 h, before pulse-labelling with CldU (red) for 20 min and IdU (green) for another 20 min. Scale bars, 10 µm. m, Left, cell cycle profiles of non-targeting, LIG1- or FEN1-knockdown U2-OS cells. Right, the percentage of U2-OS cells with more than five DDR foci (representative experiment from n = 2 biological replicates).
Extended Data Fig. 5 Low-dosage of olaparib did not induce strong DDR.
a, Percentage of U2-OS cells with more than 5 DDR foci after treatment with 1 µM or 10 μM olaparib for 24 h, or 10 µM olaparib for 1 h. Data are mean ± s.d., n = 3 biological replicates (see Source Data). b, Cell cycle profiles of U2-OS cells treated with 1 µM olaparib for 24 h, or 10 µM olaparib for 1 h. HU (2 mM, 24 h) was included as a positive control for inhibition of S phase progression, and VP-16 (10 µM, 24 h) for G2/M phase arrest (n = 2 biological replicates). c, d, Alkaline (c) or neutral (d) comet assays from U2-OS cells treated as in b. HU is a positive control for ssDNA; VP-16 is a positive control for dsDNA. Whiskers indicate fifth and ninety-fifth percentiles, and centre value depicts the median. P values determined by two-sided Kolmogorov–Smirnov test and two-tailed t-test; n = 2 biological replicates. e, Number of U2-OS cells treated as in b relative to control cells (n = 2 biological replicates).
Extended Data Fig. 6 Olparib did not induce global changes in chromatin structure.
a, Analysis of chromatin sensitivity to MNase digestion in control and HDAC-inhibited cells (n = 3 biological replicates; representative experiment is shown). Gel densitometries at different time points are presented next to the agarose gel. The number of detected bands is shown on densitometry plots. The smaller number of bands, the more sensitive chromatin is. b, Analysis of chromatin sensitivity to MNase digestion in control and PARP-inhibited cells (n = 3 biological replicates; representative experiment is shown).
Extended Data Fig. 7 p21 and fork speed regulation.
a, Distribution of S phase patterns by BrdU incorporation in non-targeting or PARP1-knockdown U2-OS cells. Indicated cells were treated with 10 µM PARPi for 24 h; n = 3 biological replicates (see Source Data). b, Representative images of PAR and PARP1 in non-targeting or PARP1-knockdown U2-OS cells. Indicated cells were treated as in a. c, Immunoblots of PARP1 knockdown efficiency in U2-OS cells 72 h after transfection with siRNA. Lamin B1 is a loading control; n = 2 biological replicates. d, Mean intensity of p21 and p53 in non-targeting or PARP1-knockdown U2-OS cells. Indicated cells were treated as in a. Data are mean ± s.d. P values were determined by a two-tailed t-test; n = 3 biological replicates. e, Immunoblots of γH2AX, PARP1 and p21 in the p21-knockdown U2-OS stable cell line. Actin is a loading control; n = 2 biological replicates. shNT, U2-OS cell line with non-targeting shRNA. f, DNA fibres from the p21-knockdown U2-OS stable cell line. Mean fork speed (kb min−1): shNT = 1.2; shP21 = 1.68. Scored forks: shNT = 305; shP21 = 207; n = 2 biological replicates. Data are mean ± s.d. P values determined by two-tailed Welch’s t-test. g, CldU/IdU ratios calculated from values in f. Percentage of highly asymmetric forks (CldU/IdU ratios < 0.5 and > 1.5) is indicated above each condition. h, Mean intensity of PAR in non-targeting or p21-knockdown U2-OS cells. Indicated cells were treated as in a. Data are mean ± s.d. P values determined by two-tailed Welch’s t-test; n = 3 biological replicates. i, Representative images of PAR in non-targeting, PARP1- or p53-knockdown U2-OS cells. Mean intensity of PAR relative to non-targeting control in U2-OS cells (representative results from n = 2 biological replicates). j, U2-OS cells, 72 h after transfection with non-targeting or p21 siRNA were pulse-labelled for 10 min with CldU (red), washed and pulse-labelled with IdU (green) for 30 min. Fork length (µm) of the first (CldU) pulse. k, Fork length (µm) of the second (IdU) pulse from the experiment in j. Mean ± s.d. of separate forks is indicated above each condition. Scored forks: NT = 388; p21 KD = 272; n = 2 biological replicates.
Extended Data Fig. 8 Effect of PARP1 knockdown in HeLa cells.
a, Immunoblots of p21 in HeLa cells and in non-targeting or p21-knockdown U2-OS cells (lines 2, 4; 10 µM PARPi, 24 h). n = 2 biological replicates. b, Immunoblots of PARP1 in non-targeting or PARP1-knockdown HeLa cells (lines 2, 4; 10 µM PARPi, 24 h). n = 2 biological replicates. c, Mean intensity of p21 in non-targeting or PARP1-knockdown HeLa cells. Indicated cells were treated with PARPi (10 µM, 24h), representative experiment from n = 2 biological replicates (see Source Data). d, Representative images of PAR and PARP1 in non-targeting or PARP1-knockdown HeLa cells. Indicated cells were treated as in c. e, Mean intensity of PAR and PARP1 in non-targeting or PARP1-knockdown HeLa cells. Indicated cells were treated as in c; representative experiment from n = 2 biological replicates.
Extended Data Fig. 9 Fork speed in double-knockdown PARP1/2.
a, DNA fibres from U2-OS cells 72 h after transfection with different siRNAs. Indicated cells were treated with 10 µM PARPi for 24 h. Mean fork speed (kb min−1) is indicated. Scored forks: NT = 586; PARPi = 263; PARP1 KD = 327; PARP1 KD–PARPi = 451; PARP2 KD = 794; PARP2 KD–PARPi = 597; PARP1/2 KD = 831; PARP1/2 KD–PARPi = 962; n = 2 biological replicates. Data are mean ± s.d. P values were determined by two-tailed Welch’s t-test (see Source Data). b, Cell cycle profiles of U2-OS cells 72 h after transfection with different siRNA and treated as in a. Percentage of cells in different phases of the cell cycle analysed using FlowJo software are indicated next to the histograms; n = 2 biological replicates. c, Immunoblots of PARP1, PARP2, p21 and p53 from experimental conditions described in a. n = 2 biological replicates.
Extended Data Fig. 10 The FSRN.
a, Number of non-targeting or BRCA1-knockdown U2-OS cells. Indicated cells were treated with 10 µM PARPi for 24 h. Data are mean ± s.d. n = 4 biological replicates (see Source Data). b, Number of MDA-MB-436 BRCA1-deficient cells 24 h after olaparib treatment. Data are mean ± s.d., n = 4 biological replicates. c, Number of OVCAR-5 ovarian cancer cells 24 h after olaparib treatment. Data are mean ± s.d., n = 4 biological replicates. d, Cell cycle profiles of OVCAR-5 ovarian cancer cells 24 h after olaparib treatment. e, The fork speed regulatory network (FSRN) model. (1) During unperturbed S phase, inactive PARP1 inhibits transcription of p21. Induction of PARP enzymatic activity is necessary for p21 promoter activation, by relief of repression, for both p53-dependent and -independent pathways (our data and shown previously18). PARP1 has high affinity to DNA nicks and ssDNA. Binding of PARP1 to DNA nicks stimulates its activity34. Moreover, a steady-state level of PARylation is necessary for the normal cell physiology, as excess of PARP activity after DNA damage reduces the amount of NAD+, affecting the ATP level35. PARP1 can bind directly to p21 and the PARP inhibitor olaparib reduces this interaction36. In our model, levels of p53 (pink hexagon), p21 (red rectangle), p21–PARP1 complex, free PARP1 (yellow trapezoid) and a low level of PARylation (small empty pentagon) are maintained at a steady state during normal S phase. PCNA (blue circle) is associated with replication forks and is bound by polymerase (Pol) δ on the lagging strand and Polε on the leading strand. In replication factories, PARP1 can be associated directly to DNA (that is, at the nicks of the lagging DNA strand) and to PCNA17. The balance between these players enables the normal speed of fork progression to be maintained. (2) Any break in DNA is promptly recognized by PARP1, which triggers its activity. PARylation can promote the recruitment of important DDR proteins37 or can directly inhibit fork progression. Excess of PARylation needs to be removed by PARG enzymes, allowing the fork to resume38. (3) When DNA is severely damaged, PARP1 is strongly activated. PARylated PARP1 releases p21 from the p21–PARP1 complexes. PARylated PARP1 is also bound by p53, which transactivates p21. After prolonged fork arrest, processive DNA polymerases dissociate from modified PCNA39 and are replaced by p21. p21 can inhibit PCNA-dependent DNA replication in the absence of cyclin/CDK. Furthermore, p21 blocks the ability of PCNA to activate DNA Polδ15. Therefore, PARylation and p21 act as suppressors of DNA replication. (4) PARP inhibitors disrupt FSRN.
Supplementary Information
Supplementary Information
This file contains the uncropped western blots and FACS gating examples.
Source Data
Rights and permissions
About this article
Cite this article
Maya-Mendoza, A., Moudry, P., Merchut-Maya, J.M. et al. High speed of fork progression induces DNA replication stress and genomic instability. Nature 559, 279–284 (2018). https://doi.org/10.1038/s41586-018-0261-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0261-5
This article is cited by
-
Quantity and quality of minichromosome maintenance protein complexes couple replication licensing to genome integrity
Communications Biology (2024)
-
MBD1 protects replication fork stability by recruiting PARP1 and controlling transcription-replication conflicts
Cancer Gene Therapy (2024)
-
Transcription–replication conflicts underlie sensitivity to PARP inhibitors
Nature (2024)
-
FANCJ promotes PARP1 activity during DNA replication that is essential in BRCA1 deficient cells
Nature Communications (2024)
-
Accelerated DNA replication fork speed due to loss of R-loops in myelodysplastic syndromes with SF3B1 mutation
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.