Abstract
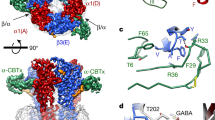
Fast inhibitory neurotransmission in the brain is principally mediated by the neurotransmitter GABA (γ-aminobutyric acid) and its synaptic target, the type A GABA receptor (GABAA receptor). Dysfunction of this receptor results in neurological disorders and mental illnesses including epilepsy, anxiety and insomnia. The GABAA receptor is also a prolific target for therapeutic, illicit and recreational drugs, including benzodiazepines, barbiturates, anaesthetics and ethanol. Here we present high-resolution cryo-electron microscopy structures of the human α1β2γ2 GABAA receptor, the predominant isoform in the adult brain, in complex with GABA and the benzodiazepine site antagonist flumazenil, the first-line clinical treatment for benzodiazepine overdose. The receptor architecture reveals unique heteromeric interactions for this important class of inhibitory neurotransmitter receptor. This work provides a template for understanding receptor modulation by GABA and benzodiazepines, and will assist rational approaches to therapeutic targeting of this receptor for neurological disorders and mental illness.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Nutt, D. J. & Malizia, A. L. New insights into the role of the GABAA-benzodiazepine receptor in psychiatric disorder. Br. J. Psychiatry 179, 390–396 (2001).
Maconochie, D. J., Zempel, J. M. & Steinbach, J. H. How quickly can GABAA receptors open? Neuron 12, 61–71 (1994).
Braat, S. & Kooy, R. F. The GABAA receptor as a therapeutic target for neurodevelopmental disorders. Neuron 86, 1119–1130 (2015).
Chuang, S. H. & Reddy, D. S. Genetic and molecular regulation of extrasynaptic GABA-A Receptors in the brain: therapeutic insights for epilepsy. J. Pharmacol. Exp. Ther. 364, 180–197 (2018).
Jacob, T. C., Moss, S. J. & Jurd, R. GABAA receptor trafficking and its role in the dynamic modulation of neuronal inhibition. Nat. Rev. Neurosci. 9, 331–343 (2008).
Roberts, E. & Frankel, S. γ-Aminobutyric acid in brain: its formation from glutamic acid. J. Biol. Chem. 187, 55–63 (1950).
Awapara, J., Landua, A. J., Fuerst, R. & Seale, B. Free γ-aminobutyric acid in brain. J. Biol. Chem. 187, 35–39 (1950).
Sternbach, L. H. The benzodiazepine story. J. Med. Chem. 22, 1–7 (1979).
Sieghart, W. Allosteric modulation of GABAA receptors via multiple drug-binding sites. Adv. Pharmacol. 72, 53–96 (2015).
Sigel, E. & Buhr, A. The benzodiazepine binding site of GABAA receptors. Trends Pharmacol. Sci. 18, 425–429 (1997).
Votey, S. R., Bosse, G. M., Bayer, M. J. & Hoffman, J. R. Flumazenil: a new benzodiazepine antagonist. Ann. Emerg. Med. 20, 181–188 (1991).
Lobo, I. A. & Harris, R. A. GABAA receptors and alcohol. Pharmacol. Biochem. Behav. 90, 90–94 (2008).
Sieghart, W. Structure and pharmacology of γ-aminobutyric acidA receptor subtypes. Pharmacol. Rev. 47, 181–234 (1995).
Sigel, E. & Steinmann, M. E. Structure, function, and modulation of GABAA receptors. J. Biol. Chem. 287, 40224–40231 (2012).
Nemecz, A., Prevost, M. S., Menny, A. & Corringer, P. J. emerging molecular mechanisms of signal transduction in pentameric ligand-gated ion channels. Neuron 90, 452–470 (2016).
Miller, P. S. & Aricescu, A. R. Crystal structure of a human GABAA receptor. Nature 512, 270–275 (2014).
Whiting, P. J., McKernan, R. M. & Wafford, K. A. Structure and pharmacology of vertebrate GABAA receptor subtypes. Int. Rev. Neurobiol. 38, 95–138 (1995).
Olsen, R. W. Allosteric ligands and their binding sites define γ-aminobutyric acid (GABA) type A receptor subtypes. Adv. Pharmacol. 73, 167–202 (2015).
Forman, S. A. & Miller, K. W. Anesthetic sites and allosteric mechanisms of action on Cys-loop ligand-gated ion channels. Can. J. Anaesth. 58, 191–205 (2011).
Walsh, R. M. J. et al. Structural principles of distinct assemblies of the human α4β2 nicotinic receptor. Nature 557, 261–265 (2018).
Marangos, P. J., Patel, J., Boulenger, J. P. & Clark-Rosenberg, R. Characterization of peripheral-type benzodiazepine binding sites in brain using [3H]Ro 5-4864. Mol. Pharmacol. 22, 26–32 (1982).
Baumann, S. W., Baur, R. & Sigel, E. Forced subunit assembly in α1β2gγ2 GABAA receptors. Insight into the absolute arrangement. J. Biol. Chem. 277, 46020–46025 (2002).
Baur, R., Minier, F. & Sigel, E. A. GABAA receptor of defined subunit composition and positioning: concatenation of five subunits. FEBS Lett. 580, 1616–1620 (2006).
Tretter, V., Ehya, N., Fuchs, K. & Sieghart, W. Stoichiometry and assembly of a recombinant GABAA receptor subtype. J. Neurosci. 17, 2728–2737 (1997).
Curtis, D. R., Duggan, A. W., Felix, D. & Johnston, G. A. GABA, bicuculline and central inhibition. Nature 226, 1222–1224 (1970).
Ueno, S., Bracamontes, J., Zorumski, C., Weiss, D. S. & Steinbach, J. H. Bicuculline and gabazine are allosteric inhibitors of channel opening of the GABAA receptor. J. Neurosci. 17, 625–634 (1997).
Amin, J. & Weiss, D. S. GABAA receptor needs two homologous domains of the β-subunit for activation by GABA but not by pentobarbital. Nature 366, 565–569 (1993).
Mortensen, M. et al. Photo-antagonism of the GABAA receptor. Nat. Commun. 5, 4454 (2014).
Sigel, E., Baur, R., Kellenberger, S. & Malherbe, P. Point mutations affecting antagonist affinity and agonist dependent gating of GABAA receptor channels. EMBO J. 11, 2017–2023 (1992).
Smith, G. B. & Olsen, R. W. Identification of a [3H]muscimol photoaffinity substrate in the bovine γ-aminobutyric acidA receptor α subunit. J. Biol. Chem. 269, 20380–20387 (1994).
Boileau, A. J., Evers, A. R., Davis, A. F. & Czajkowski, C. Mapping the agonist binding site of the GABAA receptor: evidence for a β-strand. J. Neurosci. 19, 4847–4854 (1999).
Goldschen-Ohm, M. P., Wagner, D. A. & Jones, M. V. Three arginines in the GABAA receptor binding pocket have distinct roles in the formation and stability of agonist- versus antagonist-bound complexes. Mol. Pharmacol. 80, 647–656 (2011).
Buhr, A., Baur, R. & Sigel, E. Subtle changes in residue 77 of the γ subunit of α1β2γ2 GABAA receptors drastically alter the affinity for ligands of the benzodiazepine binding site. J. Biol. Chem. 272, 11799–11804 (1997).
Wingrove, P. B., Thompson, S. A., Wafford, K. A. & Whiting, P. J. Key amino acids in the γ subunit of the γ-aminobutyric acidA receptor that determine ligand binding and modulation at the benzodiazepine site. Mol. Pharmacol. 52, 874–881 (1997).
Wieland, H. A., Luddens, H. & Seeburg, P. H. A single histidine in GABAA receptors is essential for benzodiazepine agonist binding. J. Biol. Chem. 267, 1426–1429 (1992).
Dunn, S. M., Davies, M., Muntoni, A. L. & Lambert, J. J. Mutagenesis of the rat α1 subunit of the γ-aminobutyric acidA receptor reveals the importance of residue 101 in determining the allosteric effects of benzodiazepine site ligands. Mol. Pharmacol. 56, 768–774 (1999).
Davies, M., Bateson, A. N. & Dunn, S. M. Structural requirements for ligand interactions at the benzodiazepine recognition site of the GABAA receptor. J. Neurochem. 70, 2188–2194 (1998).
Baur, R. & Sigel, E. Replacement of histidine in position 105 in the α5 subunit by cysteine stimulates zolpidem sensitivity of α5β2γ2 GABAA receptors. J. Neurochem. 103, 2556–2564 (2007).
Ramerstorfer, J. et al. The GABAA receptor α+β− interface: a novel target for subtype selective drugs. J. Neurosci. 31, 870–877 (2011).
Unwin, N. Nicotinic acetylcholine receptor and the structural basis of neuromuscular transmission: insights from Torpedo postsynaptic membranes. Q. Rev. Biophys. 46, 283–322 (2013).
Wongsamitkul, N. et al. alpha subunits in GABAA receptors are dispensable for GABA and diazepam action. Sci. Rep. 7, 15498 (2017).
Hansen, S. B., Wang, H. L., Taylor, P. & Sine, S. M. An ion selectivity filter in the extracellular domain of Cys-loop receptors reveals determinants for ion conductance. J. Biol. Chem. 283, 36066–36070 (2008).
Buller, A. L., Hastings, G. A., Kirkness, E. F. & Fraser, C. M. Site-directed mutagenesis of N-linked glycosylation sites on the gamma-aminobutyric acid type A receptor alpha 1 subunit. Mol. Pharmacol. 46, 858–865 (1994).
Dwyer, T. M., Adams, D. J. & Hille, B. The permeability of the endplate channel to organic cations in frog muscle. J. Gen. Physiol. 75, 469–492 (1980).
Jensen, M. L., Schousboe, A. & Ahring, P. K. Charge selectivity of the Cys-loop family of ligand-gated ion channels. J. Neurochem. 92, 217–225 (2005).
Johnston, G. A. GABAA receptor pharmacology. Pharmacol. Ther. 69, 173–198 (1996).
Penmatsa, A., Wang, K. H. & Gouaux, E. X-ray structure of dopamine transporter elucidates antidepressant mechanism. Nature 503, 85–90 (2013).
Baur, R. et al. Molecular analysis of the site for 2-arachidonylglycerol (2-AG) on the β2 subunit of GABAA receptors. J. Neurochem. 126, 29–36 (2013).
Miller, P. S. et al. Structural basis for GABAA receptor potentiation by neurosteroids. Nat. Struct. Mol. Biol. 24, 986–992 (2017).
Laverty, D. et al. Crystal structures of a GABAA-receptor chimera reveal new endogenous neurosteroid-binding sites. Nat. Struct. Mol. Biol. 24, 977–985 (2017).
Morales-Perez, C. L., Noviello, C. M. & Hibbs, R. E. Manipulation of subunit stoichiometry in heteromeric membrane proteins. Structure 24, 797–805 (2016).
Kawate, T. & Gouaux, E. Fluorescence-detection size-exclusion chromatography for precrystallization screening of integral membrane proteins. Structure 14, 673–681 (2006).
Nielsen, H. Predicting secretory proteins with SignalP. Methods Mol. Biol. 1611, 59–73 (2017).
Reeves, P. J., Callewaert, N., Contreras, R. & Khorana, H. G. Structure and function in rhodopsin: high-level expression of rhodopsin with restricted and homogeneous N-glycosylation by a tetracycline-inducible N-acetylglucosaminyltransferase I-negative HEK293S stable mammalian cell line. Proc. Natl Acad. Sci. USA 99, 13419–13424 (2002).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Scheres, S. H. Processing of structurally heterogeneous cryo-EM data in RELION. Methods Enzymol. 579, 125–157 (2016).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Schwede, T., Kopp, J., Guex, N. & Peitsch, M. C. SWISS-MODEL: An automated protein homology-modeling server. Nucleic Acids Res. 31, 3381–3385 (2003).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Emsley, P. & Crispin, M. Structural analysis of glycoproteins: building N-linked glycans with Coot. Acta Crystallogr D Struct Biol 74, 256–263 (2018).
Codding, P. W. & Muir, A. K. Molecular structure of Ro15-1788 and a model for the binding of benzodiazepine receptor ligands. Structural identification of common features in antagonists. Mol. Pharmacol. 28, 178–184 (1985).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Amunts, A. et al. Structure of the yeast mitochondrial large ribosomal subunit. Science 343, 1485–1489 (2014).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Wallace, A. C., Laskowski, R. A. & Thornton, J. M. LIGPLOT: a program to generate schematic diagrams of protein–ligand interactions. Protein Eng. 8, 127–134 (1995).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Morin, A. et al. Collaboration gets the most out of software. eLife 2, e01456 (2013).
Hadingham, K. L. et al. Role of the beta subunit in determining the pharmacology of human gamma-aminobutyric acid type A receptors. Mol. Pharmacol. 44, 1211–1218 (1993).
Karim, N. et al. Potency of GABA at human recombinant GABAA receptors expressed in Xenopus oocytes: a mini review. Amino Acids 44, 1139–1149 (2013).
Mortensen, M., Patel, B. & Smart, T. G. GABA potency at GABAA receptors found in synaptic and extrasynaptic zones. Front. Cell. Neurosci. 6, 1 (2011).
Baur, R. & Sigel, E. On high- and low-affinity agonist sites in GABAA receptors. J. Neurochem. 87, 325–332 (2003).
Richter, L. et al. Diazepam-bound GABAA receptor models identify new benzodiazepine binding-site ligands. Nat. Chem. Biol. 8, 455–464 (2012).
Middendorp, S. J., Puthenkalam, R., Baur, R., Ernst, M. & Sigel, E. Accelerated discovery of novel benzodiazepine ligands by experiment-guided virtual screening. ACS Chem. Biol. 9, 1854–1859 (2014).
Du, J., Lu, W., Wu, S., Cheng, Y. & Gouaux, E. Glycine receptor mechanism elucidated by electron cryo-microscopy. Nature 526, 224–229 (2015).
Hibbs, R. E. & Gouaux, E. Principles of activation and permeation in an anion-selective Cys-loop receptor. Nature 474, 54–60 (2011).
Jansen, M., Bali, M. & Akabas, M. H. Modular design of Cys-loop ligand-gated ion channels: functional 5-HT3 and GABA rho1 receptors lacking the large cytoplasmic M3M4 loop. J. Gen. Physiol. 131, 137–146 (2008).
Acknowledgements
We thank D. Cawley for antibody production, X. Bai for electron microscopy discussion, P. Emsley for guidance on glycosylation tools in Coot, W. Costello for initial construct screening and all members of the Hibbs laboratory for discussion. Single-particle cryo-EM data were collected at the University of Texas Southwestern Medical Center Cryo-Electron Microscopy Facility, which is supported by the CPRIT Core Facility Support Award RP170644. We thank D. Nicastro and D. Stoddard for support in facility access and data acquisition. R.W. acknowledges support from the Sara and Frank McKnight Fund for Biochemical Research and the NIH (T32GM008203). R.E.H. is supported by a McKnight Scholar Award, The Welch Foundation (I-1812) and the NIH (DA037492, DA042072, and NS095899).
Reviewer information
Nature thanks G. Akk, P. Corringer, A. Evers and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
S.Z. performed sample preparation, data collection, model building and writing of the manuscript. C.M.N. and R.M.W. collected microscopy data and edited the manuscript. J.T. performed electrophysiology experiments. J.J.K. assisted in biochemistry and edited the manuscript. R.E.H. oversaw all aspects of the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Alignment of GABAA and other Cys-loop receptor subunits.
Cryo-EM constructs (γ2 affinity tag not shown) are numbered starting with the first residue of the mature protein. Sequences aligned (UniProt or PDB accession codes): Homo sapiens α1 GABAA (HS, P14867), H. sapiens β2 GABAA (P47870), H. sapiens γ2 GABAA (P18507), H. sapiens GABAA β3 (4COF), H. sapiens glycine α3 (5CFB), Danio rerio glycine α1 (DR, 3JAE), Caenorhabditis elegans α (CE, 3RHW), H. sapiens α4 nAChR (5KXI), H. sapiens β2 nAChR (5KXI) and Mus musculus 5-HT3 receptor (MM, 4PIR). α-helices (cylinders), β-strands (arrows), and inserted linker (cyan) are indicated.
Extended Data Fig. 2 Biochemistry and binding assay.
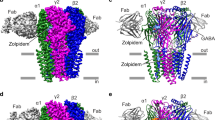
a, FSEC of GABAA receptor with and without Fab bound, and SDS–PAGE analysis of a representative purification (from n > 10 purifications). b, Saturation binding assay with [3H]-flumazenil. Single-site binding fits for receptor alone and receptor plus Fab both exhibited a Hill slope of ~1 (0.97 and 0.89, respectively). Plotted results are from a representative experiment performed in triplicate. n = 3 independent experiments. Data are shown as mean ± s.d. for a representative triplicate measurement. c, Competition of 10 nM [3H]-flumazenil with diazepam. Calculated Ki for diazepam assumes a Kd for [3H]-flumazenil of 7.7 nM. n = 2 independent experiments in triplicate. Data are shown as mean ± s.d. for a representative triplicate measurement. d, Dose–response experiments in the presence or absence of Fab. HEK cells were transfected with cryo-EM constructs and patch-clamped with or without pretreatment with 1 µM Fab for 1 min. Hill slopes are 1.7 and 1.4 with and without Fab, respectively. Published EC50 values for GABA range from 6.6 µM–107 µM71,72,73,74. n = 3 experiments from different cells. Data are plotted as mean ± s.d. e, Whole-cell patch-clamp recording of long application of cryo-EM ligands at concentrations used in cryo-EM samples to assess conformational state at equilibrium. The two traces shown are from one continuous recording; in between the two responses, Fab was added to 1 µM for one minute to saturate all receptor sites before second application of GABA and flumazenil (including Fab). n = 3 independent experiments. f–g, Docking of diazepam at the benzodiazepine-binding site based on superposition of benzodiazepine rings. The phenyl ring of diazepam would orient towards the membrane, possibly forming π–π-stacking interactions with Y58 on the complementary subunit. Similar to flumazenil, the halogen of diazepam could interact with H102, suggesting that this contact is conserved broadly among benzodiazepines and flumazenil. This orientation is largely consistent with predictions from a modelling and docking study75, and distinct from that suggested by affinity labelling76. In this latter prediction, the diazepam phenyl group orients away from the membrane and would require local reorganization of side chains to avoid atomic clashes. h–j, Structural details of Fab–α1 interaction. Labelled residues are on the α-subunit. i, Top view. j, Side view.
Extended Data Fig. 3 Cryo-EM image processing procedure.
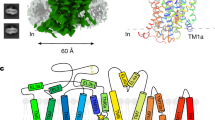
a, Representative cryo-electron micrograph of the GABAA receptor–Fab complex. n = 5,594 images. b, Images of selected 2D classes from reference-free 2D classification by Relion. c, Overview of the image processing procedure (see Methods).
Extended Data Fig. 4 Three-dimensional reconstructions of the two GABAA receptor conformations.
a, Angular distribution histogram of GABAA receptor conformation A particle images. b, Fourier shell correlation (FSC) of conformation A maps before (black) and after (blue) masking. c, Local resolution of the GABAA receptor estimated by ResMap. d–f, as in a–c but for GABAA receptor conformation B.
Extended Data Fig. 5 GABAA receptor model map validation.
a, Data collection and refinement statistics. b, c, FSC curves for cross-validation between the maps and models of both conformation A (b) and conformation B (c). FSC curves for final model versus summed map (whole) in black, for model versus half map in green (work), and for model versus half map not used for refinement in blue (free).
Extended Data Fig. 6 Cryo-EM density of the GABAA receptor in conformation A.
a–e, Cryo-EM density map of the GABAA receptor conformation A for a representative of each subunit. f–h, Cryo-EM density segments of loop C in α1, β2 and γ2-subunits. i–k, Cryo-EM density segments of M2 helix in α1, β2 and γ2-subunits. l–n, Cryo-EM density maps of ligand binding sites: flumazenil (l), two GABA binding sites (m, n).
Extended Data Fig. 7 Cryo-EM density of the GABAA receptor in conformation B.
a–e, Cryo-EM density map of the GABAA receptor conformation B for a representative of each subunit; chain IDs are in parentheses. f–h, Cryo-EM density segments of loop C in α1, β2 and γ2-subunits. i–k, Cryo-EM density segments of M2 helix in α1, β2 and γ2-subunits. l–n, Cryo-EM density maps of ligand-binding sites: flumazenil (l), two GABA binding sites (m, n).
Extended Data Fig. 8 Superposition of subunits.
a–e, Subunits of conformation A are compared to the corresponding subunit from conformation B. f–j, Superposition of subunits within conformation A. k–o, Superposition of subunits within conformation B. Chimera MatchMaker was used to generate alignments; r.m.s.d. values in Å are for Cα atoms over entire subunit. Chain IDs are in parentheses.
Extended Data Fig. 9 Permeation pathway and subunit interfaces.
a, Cartoon of permeation pathway for conformation A. A single β2-subunit is removed for clarity. Purple spheres indicate pore diameters >5.6 Å; yellow is >2.8 Å and <5.6 Å; red is <2.8 Å. b, Same as a but for conformation B. c, Pore diameters for conformation A (red) and conformation B (black). The zero value along the y axis of the plot is aligned with the α-carbon of the −2′ position of conformation B. d–m, Side view of two adjacent subunits in conformations A (d–h) and B (i–m). The view is from the periphery of the receptor towards the pore axis. Cholesterol at the interface is shown in yellow in d, i and k. Cartoon pentagons (bottom) are coloured to illustrate all subunits composing the displayed interface; subunits not participating in the displayed interface are grey. Principal (+) and complementary (−) sides of the displayed interface are labelled on each pentagon. n, Analysis of the subunit interfaces of both conformations using PDBePISA server69.
Extended Data Fig. 10 Transmembrane domain flexibility and comparison with reference structures.
a, b, Top and side view of the TMD of conformation A with density for the γ2-subunit shown. c, d, As in a, but for conformation B. e, Transmembrane domain superposition of conformation A (subunits in colour) over conformation B (grey). α-Helices are represented as cylinders. f–j, Superposition of single subunit TMD in conformation A (coloured) with its corresponding subunit in conformation B (grey). k–r, Superpositions of the four non-γ-subunits. Top and bottom rows contain the same superpositions in different representations. Conformation B is shown in all panels with α-subunits in green and β-subunits in blue. Reference structures include the glycine receptor with ivermectin bound (3JAF)77, glutamate-gated chloride channel with ivermectin bound (3RHW)78 and the GABAA β3 homopentamer (4COF)16.
Supplementary information
Supplementary Figure
This file contains Supplementary Figure 1, the uncropped gel shown in Extended Data Fig. 2.
Rights and permissions
About this article
Cite this article
Zhu, S., Noviello, C.M., Teng, J. et al. Structure of a human synaptic GABAA receptor. Nature 559, 67–72 (2018). https://doi.org/10.1038/s41586-018-0255-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0255-3
This article is cited by
-
Modulators of GABAA receptor-mediated inhibition in the treatment of neuropsychiatric disorders: past, present, and future
Neuropsychopharmacology (2024)
-
The modes of action of ion-channel-targeting neurotoxic insecticides: lessons from structural biology
Nature Structural & Molecular Biology (2023)
-
Cryo-EM structures reveal native GABAA receptor assemblies and pharmacology
Nature (2023)
-
Highly efficient and robust π-FISH rainbow for multiplexed in situ detection of diverse biomolecules
Nature Communications (2023)
-
Structural insights into opposing actions of neurosteroids on GABAA receptors
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.