Abstract
Reprogramming of mRNA translation has a key role in cancer development and drug resistance1. However, the molecular mechanisms that are involved in this process remain poorly understood. Wobble tRNA modifications are required for specific codon decoding during translation2,3. Here we show, in humans, that the enzymes that catalyse modifications of wobble uridine 34 (U34) tRNA (U34 enzymes) are key players of the protein synthesis rewiring that is induced by the transformation driven by the BRAFV600E oncogene and by resistance to targeted therapy in melanoma. We show that BRAFV600E-expressing melanoma cells are dependent on U34 enzymes for survival, and that concurrent inhibition of MAPK signalling and ELP3 or CTU1 and/or CTU2 synergizes to kill melanoma cells. Activation of the PI3K signalling pathway, one of the most common mechanisms of acquired resistance to MAPK therapeutic agents, markedly increases the expression of U34 enzymes. Mechanistically, U34 enzymes promote glycolysis in melanoma cells through the direct, codon-dependent, regulation of the translation of HIF1A mRNA and the maintenance of high levels of HIF1α protein. Therefore, the acquired resistance to anti-BRAF therapy is associated with high levels of U34 enzymes and HIF1α. Together, these results demonstrate that U34 enzymes promote the survival and resistance to therapy of melanoma cells by regulating specific mRNA translation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
11 November 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41586-021-04074-1
References
Truitt, M. L. & Ruggero, D. New frontiers in translational control of the cancer genome. Nat. Rev. Cancer16, 288–304 (2016).
Novoa, E. M. & Ribas de Pouplana, L. Speeding with control: codon usage, tRNAs, and ribosomes. Trends Genet. 28, 574–581 (2012).
El Yacoubi, B., Bailly, M. & de Crécy-Lagard, V. Biosynthesis and function of posttranscriptional modifications of transfer RNAs. Annu. Rev. Genet. 46, 69–95 (2012).
Patton, E. E. et al. BRAF mutations are sufficient to promote nevi formation and cooperate with p53 in the genesis of melanoma. Curr. Biol. 15, 249–254 (2005).
Davies, H. et al. Mutations of the BRAF gene in human cancer. Nature417, 949–954 (2002).
Selvadurai, K., Wang, P., Seimetz, J. & Huang, R. H. Archaeal Elp3 catalyzes tRNA wobble uridine modification at C5 via a radical mechanism. Nat. Chem. Biol. 10, 810–812 (2014).
Denko, N. C. Hypoxia, HIF1 and glucose metabolism in the solid tumour. Nat. Rev. Cancer8, 705–713 (2008).
Ho, J. et al. Importance of glycolysis and oxidative phosphorylation in advanced melanoma. Mol. Cancer11, 76 (2012).
Bollag, G. et al. Clinical efficacy of a RAF inhibitor needs broad target blockade in BRAF-mutant melanoma. Nature467, 596–599 (2010).
Nedialkova, D. D. & Leidel, S. A. Optimization of codon translation rates via tRNA modifications maintains proteome integrity. Cell161, 1606–1618 (2015).
Zinshteyn, B. & Gilbert, W. V. Loss of a conserved tRNA anticodon modification perturbs cellular signaling. PLoS Genet. 9, e1003675 (2013).
Laguesse, S. et al. A dynamic unfolded protein response contributes to the control of cortical neurogenesis. Dev. Cell35, 553–567 (2015).
Parmenter, T. J. et al. Response of BRAF-mutant melanoma to BRAF inhibition is mediated by a network of transcriptional regulators of glycolysis. Cancer Discov. 4, 423–433 (2014).
Verfaillie, A. et al. Decoding the regulatory landscape of melanoma reveals TEADS as regulators of the invasive cell state. Nat. Commun. 6, 6683 (2015).
Chapman, P. B. et al. Improved survival with vemurafenib in melanoma with BRAF V600E mutation. N. Engl. J. Med. 364, 2507–2516 (2011).
Sosman, J. A. et al. Survival in BRAF V600-mutant advanced melanoma treated with vemurafenib. N. Engl. J. Med. 366, 707–714 (2012).
Robert, C. et al. Improved overall survival in melanoma with combined dabrafenib and trametinib. N. Engl. J. Med. 372, 30–39 (2015).
Flaherty, K. T. et al. Improved survival with MEK inhibition in BRAF-mutated melanoma. N. Engl. J. Med. 367, 107–114 (2012).
Johannessen, C. M. et al. A melanocyte lineage program confers resistance to MAP kinase pathway inhibition. Nature504, 138–142 (2013).
Karoulia, Z., Gavathiotis, E. & Poulikakos, P. I. New perspectives for targeting RAF kinase in human cancer. Nat. Rev. Cancer17, 676–691 (2017).
Hernandez-Davies, J. E. et al. Vemurafenib resistance reprograms melanoma cells towards glutamine dependence. J. Transl. Med. 13, 210 (2015).
Abdel-Fattah, W. et al. Phosphorylation of Elp1 by Hrr25 is required for elongator-dependent tRNA modification in yeast. PLoS Genet. 11, e1004931 (2015).
Zinzalla, V., Stracka, D., Oppliger, W. & Hall, M. N. Activation of mTORC2 by association with the ribosome. Cell144, 757–768 (2011).
Rapino, F., Naumann, I. & Fulda, S. Bortezomib antagonizes microtubule-interfering drug-induced apoptosis by inhibiting G2/M transition and MCL-1 degradation. Cell Death Dis. 4, e925 (2013).
Close, P. et al. Transcription impairment and cell migration defects in elongator-depleted cells: implication for familial dysautonomia. Mol. Cell22, 521–531 (2006).
Creppe, C. et al. Elongator controls the migration and differentiation of cortical neurons through acetylation of α-tubulin. Cell136, 551–564 (2009).
Ghaye, A. P. et al. Progenitor potential ofnkx6.1-expressing cells throughout zebrafish life and during beta cell regeneration. BMC Biol. 13, 70 (2015).
Delaunay, S. et al. Elp3 links tRNA modification to IRES-dependent translation of LEF1 to sustain metastasis in breast cancer. J. Exp. Med. 213, 2503–2523 (2016).
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature483, 603–607 (2012).
Heiman, M., Kulicke, R., Fenster, R. J., Greengard, P. & Heintz, N. Cell type-specific mRNA purification by translating ribosome affinity purification (TRAP). Nat. Protoc. 9, 1282–1291 (2014).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
R Core Team. R: A Language and Environment for Statistical Computing. http://www.R-project.org/ (R Foundation for Statistical Computing, Vienna, Austria, 2011).
Acknowledgements
The Tg(mitfa:BRAFV600E);tp53−/− fish were a gift from L. Zon. We thank R. Lo and G. Ghanem for providing short-term melanoma cultures; the CHU Liège patients Biobank facility for technical support and K. Scharmann for technical assistance. We are grateful to the GIGA-zebrafish, imaging, genomics and viral vector facilities for their assistance and the ‘Fonds Leon Fredericq’ and the ‘Centre Anticancéreux’ of the CHU Liège for their financial support. This study was supported by an Incentive Grant for Scientific Research (MIS F:4532.13) from the FNRS, grants from the Concerted Research Action Program (Bio-Acet and tRAME) and Special Research Funds (C-15/44) at the University of Liège, the Belgian foundation against Cancer (FAF-F/2016/840), as well as by the Walloon Excellence in Life Sciences and Biotechnology (WELBIO). F.Rap. was supported by a TELEVIE grant (7.4503.14). S.D., F.Rap., A.C. and P.C. are a research fellow, research assistant, research director and research associate at the FNRS, respectively. J.P. and M. Voo. were supported by a grant from the Dutch Cancer Society (KWF) (UM 2015-2018).
Author information
Authors and Affiliations
Contributions
F.Rap. and P.C. designed the study and analysed experimental data; F.Rap., Z.Z., F.Ram., B.C., M.G. and P.C. performed software and bioinformatic analyses; F.Rap., S.D., D.J., A.F., C.L., L.T., P.D.T., O.S., K.S., S.S., L.K., J.P. and B.G. performed experiments; A.C., A.R., G.J., M.Voo., B.P., R.B., E.L., M.D., M. Voz, S.A.L., M.G., J.-C.M. and P.C. provided resources; F.Rap., J.-C.M. and P.C. wrote the paper. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Loss of U34 enzymes triggers cell death in BRAFV600E melanoma.
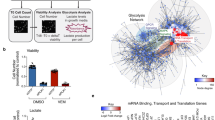
a, b, Melanoma tumours and adjacent skin were extracted from four Tg(mitfa: BRAFV600E);tp53−/− zebrafish. Indicated protein (a) and mRNA (b) levels were assessed by western blot and qPCR, respectively. n = 4 independent experiments, two-sided t-test, data are mean + s.e.m. c, Schematic representation of the wild-type (wt) or mutant Elp3ulg022 (mut) proteins in zebrafish. The radical S-adenosyl methionine (SAM) and lysine acetyl-transferase (KAT) domains are depicted. The coding region of the mutant elp3mut has a 16-bp deletion, leading to a frameshift and the apparition of a premature stop codon (amino acid 446). Relevant nucleotide and protein sequences of the elp3wt and elp3mut fish are shown. d, Representative images of indicated proteins detected in human biopsies using immunohistochemistry. Arrows indicate melanoma structures. e, ELP3 and CTU2 protein levels were assessed by western blot in human melanoma cell lines and in human fibroblasts. n = 2 independent experiments. f, Human melanoma cell lines and fibroblasts were depleted of ELP3. Cell death was measured using FACS. n = 3 independent experiments, two-sided t-test, data are mean + s.d. g, Melanoma cell lines and fibroblasts were depleted of CTU1 or CTU2. Cell death was measured using FACS. n = 3 independent experiments, two-sided t-test, data are mean + s.d. h, B16 mouse melanoma cells were stably transduced with pBABE-control (Ctr) or pBABE-BRAF(V600E) (BRAFV600E) vector. Cell death upon indicated concentrations of vemurafenib (24 h) was assessed using FACS analysis. n = 15 independent experiments, two-sided t-test, data are mean + s.d. i, Cells from h were depleted of Elp3 or Ctu2 by shRNA. Cell death was measured using FACS analysis. n = 6 independent experiments, two-sided t-test, data are mean + s.d. j, Schematic representation of the functional domains of human full-length ELP3 protein (ELP3-FL) and of two inactive mutants generated by deletion of the KAT domain (ELP3-ΔKAT) or by mutation of the essential tyrosine in position 529 in alanine (ELP3-Y529A). k, A2058 cells were stably depleted of ELP3 by shRNA and then complemented with indicated ELP3 plasmids. Cell death was quantified using FACS. n = 6 independent experiments, two-sided t-test, data are mean + s.d. l, Proteins identified by SILAC in B16 melanoma cells stably expressing pBABE-BRAF(V600E) compared to pBABE-control. n = 2,755 genes. m, Upregulated proteins from l were detected by western blot n = 2 independent experiments. Percentage of AAA, CAA, GAA codon content is indicated on the right. n, The proteomic data obtained by comparison of the B16 control- and BRAF(V600E)-expressing cells were analysed using STRING. Most representative upregulated pathways (n = 1,433 genes, database: GO pathways) are listed with P values.
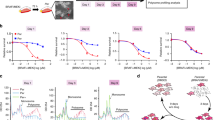
Extended Data Fig. 2 Cells that are dependent on U34 enzymes cells show increased HIF1α-dependent metabolism.
a, Hif1α protein levels were detected in B16 cells expressing pBABE-control or pBABE-BRAF(V600E). b, HIF1α protein levels were assessed in a set of human primary melanoma lines by western blot (n = 2 independent experiments). Numbers at the top indicate the fold increase in HIF1α protein levels assessed by densitometry (HIF1α/GAPDH), percentages on the right indicate combined AAA, CAA, GAA codon content. c, Cells as in a were tested for expression of the indicated mRNAs by qPCR. n = 2 independent experiments, data are mean + s.d. d, Patient-derived short-term cultured melanoma lines were tested for expression of the indicated mRNAs by qPCR. n = 3 independent experiments, two-sided t-test, data are mean + s.d. e, Measurement of the uptake of 2-DG using FACS (n = 2 independent experiments, data are median + s.d). f, Lactate produced in the medium obtained from the indicated lines was quantified after 24 h by luminescence assay. n = 2 independent experiments, data are mean + s.d. g, Kinetic ECAR response of indicated patient-derived short-term cultured melanoma lines to glucose (10 mM), oligomycin (1 μM) and 2-DG (100 mM). n = 3 independent experiments, data are mean + s.e.m. h, Quantification of glycolysis, glycolysis capacity, glycolytic reserve and non-glycolytic acidification of patient-derived short-term cultured melanoma lines. n = 3 independent experiments, two-sided t-test, data are mean + s.d. i, NRASQ61*-mutated short-term melanoma lines MM011 and MM047 were tested for expression of the indicated mRNAs by qPCR. n = 2 independent experiments, data are mean + s.d. j, Cells as in i, the lactate produced in the medium of the indicated lines was quantified after 24 h by luminescence assay. n = 2 independent experiments, data are mean + s.d. k, l, Glucose consumption (k) and lactate produced in the medium (l) of the indicated primary lines was quantified by NMR over time. n = 3 independent experiments, two-sided t-test, data are mean + s.d. m, Cells as in i were depleted of either ELP3 or CTU1 and cell death was quantified by FACS analysis. n = 3 independent experiments, two-sided t-test, data are mean + s.d.
Extended Data Fig. 3 Loss of U34 enzymes correlates with decreased HIF1α protein levels and functionality.
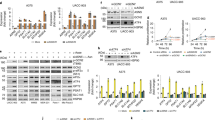
a–c, B16 melanoma cells were depleted of Elp3 by shRNA. RNA and RPF libraries were prepared and deep sequencing was performed. Differential expression analysis between control (shCtrl) and Elp3 shRNA (shElp3) RNA and RPF libraries was performed. Results of one-sided Mann–Whitney U-tests assessing the difference between the distribution of residuals between groups of genes. Top 1%, n = 137; top 5%, n = 683; top 10%, n = 1,366; top 20%, n = 2,732. a, For none of the sets of randomly selected mRNAs is the average as high or as low as the one observed for the top 5% richest or poorest mRNAs, respectively. b, Volcano plot of control and Elp3 shRNA RNA libraries is shown. n = 6 independent samples, Wald tests assessing whether the log2 fold-change coefficient (control versus Elp3 shRNA versus; paired design) fitted for each gene in a negative binomial generalized linear model comparing control (shCtrl) and Elp3 shRNA (paired design) are significantly different from zero. Correction for multiple tests was performed using the Benjamini–Hochberg procedure. c, GSEA analysis was used to analyse for significant protein signatures of genes. Datasets of the target genes of transcription factors were used. n = 17,128 genes. d–f, Indicated melanoma lines were depleted of ELP3 and CTU1 by shRNA. mRNA expression (e) and protein levels (d and f) of indicated genes were assessed by qPCR (data are mean + s.d.) and western blot, respectively. n = 2 independent experiments. g, h, B16 melanoma cells were depleted of Elp3 or Ctu2 by shRNA. Indicated protein (g) and mRNA (h) levels were quantified by western blot and qPCR, respectively. n = 2 independent experiments, data are mean + s.d. i, j, A2058 melanoma cells were depleted of ELP3 or CTU1 by shRNA. Indicated protein (i) and mRNA (j) levels were quantified by western blot and qPCR, respectively. n = 2 independent experiments, data are mean + s.d.
Extended Data Fig. 4 Loss of U34 enzymes impairs HIF1α-dependent metabolism.
a, b, MM117 (BRAFWT) primary line was depleted of ELP3 (a) or CTU2 (b). Lactate and 13C-glucose-derived lactate produced in the medium was quantified by NMR over time. n = 3 independent experiments, two-sided t-test, data are mean + s.d. c, d, MM029 (BRAFV600E) primary line was depleted of ELP3 (c) or CTU2 (d). Lactate and 13C-glucose derived lactate produced in the medium was quantified by NMR over time. n = 3 independent experiments, two-sided t-test, data are mean + s.d. e–g, Human short-term primary melanoma lines were depleted of HIF1A by esiRNA. e, Indicated proteins were detected by western blot (e, n = 1 independent experiment). f, Death of cells in e was quantified by FACS. n = 2 independent experiments, data are mean + s.d. g, Indicated mRNA levels of cells in e were quantified by qPCR. n = 2 independent experiments, data are mean + s.d. h, i, B16 melanoma cells stably expressing pBABE-control or pBABE-BRAF(V600E) were depleted of Hif1a by esiRNA. h, Indicated protein levels were detected by western blot. n = 1 independent experiment. i, Cell death was quantified by FACS. n = 2 independent experiments, data are mean + s.d. j–l, A2058 cells were depleted of HIF1A by esiRNA. j, Indicated protein levels were detected by western blot. n = 1 independent experiment. k, Cell death was quantified by FACS. n = 2 independent experiments, data are mean + s.d. l, The expression of the indicated mRNAs was detected by qPCR. n = 2 independent experiments, data are mean + s.d.
Extended Data Fig. 5 HIF1α requires U34 enzymes for proper translation.
a, qPCR products from input and ribosome immunoprecipitation were run in agarose gel. n = 3 independent experiments. b, c, A2058 cells expressing HIF1AWT–Flag (HIF1α-WT) or HIF1AMut–Flag (HIF1α-Mut) constructs were depleted of ELP3 or CTU1 by shRNA. b, The indicated protein levels were detected by western blot. n = 2 independent experiments. c, Cell death was quantified by FACS analysis. n = 4 independent experiments, two-sided t-test, data are mean + s.d. d, e, MM029 short-term primary melanoma cell line (d) and A2058 establish melanoma line (e) expressing the empty vector (Empty), the HIF1AWT–Flag or HIF1AMut–Flag constructs were depleted of ELP3 or CTU1 by shRNA. Measurement of the uptake of 2-deoxy-2-DG by FACS analysis. Data are expressed as fold increase of median fluorescence compared to the corresponding control shRNA condition. n = 3 independent experiments, two-sided t-test, data are mean + s.d. f, g, MM029 short-term primary melanoma cell line (f) and A2058 establish melanoma line (g) expressing the empty vector, the HIF1AWT–Flag or HIF1AMut–Flag constructs were depleted of ELP3 or CTU1 by shRNA. Lactate release in the medium was quantified after 24 h by luminescence assay. Data are expressed as fold increase over the corresponding control shRNA condition. n = 2 independent experiments, data are mean + s.d. h, Protein aggregates were detected in ELP3 and CTU2-depleted primary melanoma cultures after PROTEOSTAT staining using FACS. n = 3 independent experiments, two-sided t-test, data are mean + s.d. Representative plots are shown on the left. i, j, Protein aggregates were isolated in MM029 cells depleted of ELP3 or CTU2 (i) and in B16 cells depleted of Elp3 (j). Indicated protein levels in the total and aggregate compartment were detected by western blot. n = 2 independent experiments. k, B16 cells depleted of Elp3 or Ctu2 by shRNA were treated for 4 h with the indicated concentration of proteasome inhibitor MG132 and lysosome inhibitor chloroquine. The levels of indicated proteins were detected by western blot. n = 2 independent experiments.
Extended Data Fig. 6 HIF1α and U34 enzymes levels are upregulated in samples from patients with metastatic and drug-resistant melanoma.
a, Representative images of indicated proteins detected by immunohistochemistry are shown in human melanoma biopsies classified by Clark index. b, Immunohistochemistry staining of indicated proteins were performed in primary melanoma samples (n = 25) and in metastasis (n = 12). Quantification is shown on the right and representative images are shown on the left. Two-sided t-test, data are mean + s.d. c, Correlation index (Pearson coefficient) of samples in b. d, e, HIF1α signature (d, invasive (INV), n = 95 independent samples; proliferative (PRO), n = 82 independent samples) and U34 enzymes (e, invasive, n = 95 independent samples; proliferative, n = 82 independent samples) are elevated in short-term cultured melanoma lines of the invasive compared to proliferative phenotype (classification from Chapman et al.15). d, Two-tailed Wilcoxon matched-pairs signed-rank test. e, Two-tailed Mann–Whitney U-test. d, e, Data are median, minimum and maximum. f, Schematic representing the selection of samples from patients with BRAFV600E melanoma. g, Elevated MITFlow signature expression is correlated with resistance to BRAF inhibition. BRAFV600E melanoma cell lines (n = 28) from the CCLE cohort were used to correlate PLX4720 (BRAF inhibition) IC50 concentrations with the expression of the MITFlow signature (Pearson correlation coefficient R; statistical test: non-FDR corrected, two-tailed t-test). h, The expression of the U34-enzyme signature (that is, ELP1–ELP3–ALKBH8–CTU1–CTU2) and the HIF1α signature (HIF1A–GLUT1-GLUT3–VEGFA–LDHA–CXCR4–MCT4–PDK1) was evaluated in MITFlow and MITFhigh populations. MITFlow, n = 59 independent samples; MITFhighn = 128 independent samples; two-tailed Mann–Whitney U-test; data are the median and 25th to 75th percentile (see ‘Statistical analysis’).
Extended Data Fig. 7 Melanoma cells that are resistant to BRAF-inhibitors show increased levels of U34 enzymes and high HIF1α-dependent metabolism.
a, Indicated mRNA levels of isogenic primary melanoma cells sensitive (SENS) or resistant (RES) to vemurafenib (1 μM) were assessed by qPCR. n = 2 independent experiments, data are mean + s.d. b, Indicated mRNA levels of HIF1α targets of cells in a were assessed by qPCR. n = 2 independent experiments, data are mean + s.d. c, Cells as in a. Lactate production in the medium was quantified by NMR. n = 3 independent experiments, two-sided t-test, data are mean + s.d. d, Lactate production in the medium was assessed in cells as in a after 24 h by luminescence assay. n = 2 independent experiments, data are mean + s.d. e, f, Melanoma cell lines, which differ in BRAF status and in sensitivity to MAPK inhibitors (MeWo is BRAFWT, insensitive; SkMel5 is BRAFV600E, sensitive; A2058 is BRAFV600E, resistant) were depleted of ELP3 by shRNA and treated for 24 h with the indicated concentrations of vemurafenib or the U0126 MEK1 inhibitor (top and bottom panels of e, respectively). e, Cell death was quantified by FACS. n = 9 independent experiments, two-sided t-test, data are mean ± s.d. f, Synergy between specific inhibitors and ELP3 depletion was calculated by the Calcusyn software. C.I. index, in red the synergistic C.I. value. g, h, As in e and f, but with cells depleted of CTU2. n = 9 independent experiments, two-sided t-test, data are mean ± s.d.
Extended Data Fig. 8 Depletion of ELP3 or CTU2 strongly reduces tumour growth of melanoma tumours with acquired drug resistance in a HIF1α-dependent manner.
a, Eight-week-old NOD/SCID mice were injected with A375 drug-resistant cells (RES; vemurafenib 0.3 μM) depleted of ELP3 using an inducible shRNA against ELP3 or control. After 10 days, all mice were treated daily with IPTG to induce depletion for two weeks. Mice were also daily treated or not with vemurafenib (25 mg kg−1) for two weeks. Indicated protein levels were assessed in tumours after death by western blot. n = 3 independent animals. b, Eight-week-old NOD/SCID mice were injected with A375 drug-resistant cells (vemurafenib 0.3 μM) depleted of CTU2 expression using an inducible shRNA against CTU2 or control. After 10 days, mice were treated daily with IPTG to induce depletion for two weeks. Mice were also daily treated or not with vemurafenib (25 mg kg−1) for two weeks. Indicated protein levels were assessed in tumours after death by western blot. n = 3 independent animals. c, Indicated primary melanoma lines were depleted of HIF1α by esiRNA and treated or not with 15 μM vemurafenib for 24 h. Cell death was quantified by FACS. n = 2 independent experiments, data are mean + s.d. d, A2058 cells were depleted of HIF1A and treated for 24 h as indicated. Cell death was quantified by FACS. n = 2 independent experiments, data are mean + s.d. e, f, MM029 (e) or A2058 (f) cells expressing the empty vector (Empty), HIF1AWT–Flag or HIF1AMut–Flag constructs were depleted of ELP3 or CTU1 using shRNA and treated as indicated for 24 h. Cell death was quantified by FACS. n = 2 independent experiments, data are mean + s.d. g, NOD/SCID mice were injected with A2058 cells expressing the HIF1AWT–Flag or HIF1AMut–Flag constructs and infected with inducible shRNA control or ELP3 shRNA (1 × 106 cells per condition). After 10 days, mice received daily IPTG (100 μl; 25 mM) and vemurafenib (25 mg kg−1) treatment for 26 days. ELP3 and Flag–HIF1α protein levels were detected after death in three different tumours by western blot.
Extended Data Fig. 9 The PI3K–mTORC2 pathway regulates elongator protein levels.
a, B16 melanoma cells were treated for 48 h with DMEM (C), 100 nM MK2206 (M), 100 nM NVP-BEZ235 (B), 100 nM rapamycin (R), 100 nM vemurafenib (V) or 100 nM erlotinib (E) and the indicated proteins were detected by western blot. n = 3 independent experiments. b, The indicated melanoma cell lines were treated with DMEM, 100 nM NVP-BEZ235, 100 nM rapamycin or 100 nM MK2206 (M) for 48 h. Levels of indicated proteins were assessed by western blot. n = 2 independent experiments. c, B16 cells were treated for 48 h with the mTOR catalytic inhibitor PP242 (10 nM). Indicated protein levels were detected by western blot. n = 2 independent experiments. d, B16 cells were treated with insulin (250 nM) for the indicated times. Protein levels were assessed by western blot. n = 2 independent experiments. e, B16 cells were treated or not with 250 nM insulin for 2 h. Elp1 phosphorylation was assessed after endogenous Elp1 immunoprecipitation and treatment or not with phosphatase λ. Phosphorylation was assessed by western blot with a pan p-serine antibody. n = 1 independent experiment. f, B16 melanoma cells were transfected with Flag–ELP1 or Flag–ELP1(S1174A) phospho-mutant constructs and stimulated with 250 nM insulin for 2 h. Elp1 phosphorylation at S1174 was assessed after anti-Flag immunoprecipitation and using the anti-phosphorylated ELP1 (S1174) specific antibody for blotting. n = 2 independent experiments. g, Schematic representation of human ELP1 protein, its functional domains and the localization of S1174 (top). A2058 cells were depleted of ELP1 by shRNA and complemented with the empty vector (Flag–EV), the Flag–ELP1(WT) or Flag–ELP1(S1174A) (phospho-mutant) constructs. Cell death was quantified by FACS analysis. n = 6 independent experiments, two-sided t-test, data are mean + s.e.m. h, B16 cells were depleted of indicated genes by esiRNA. Indicated mRNA levels were assessed by qPCR. n = 2 independent experiments, data are mean + s.d. i, j, B16 cells were treated with 250 nM insulin for 2 h. Elp1 interaction with the mTORC2 or mTORC1 complexes was assessed by Rictor (i) or Raptor (j) immunoprecipitation, respectively. Indicated proteins were detected by western blot. n = 2 independent experiments.
Extended Data Fig. 10 mTORC2–ELP1–HIF1α axis.
a, b, B16 (a) or A2058 (b) cells were treated or not with 250 nM insulin for 2 h and the interaction of Elp1, Rictor, Raptor and Rpl7 with the ribosomal protein Rpl26 was assessed by Rpl26 immunoprecipitation. Indicated proteins were detected by western blot. n = 2 independent experiments. c, The ribosome-containing fraction (Ribo) was separated from the whole-cell extract (WCE) by ultracentrifugation of A2058 melanoma cells. Levels of the indicated proteins were detected by western blot. n = 2 independent experiments. d, e, Indicated short-term melanoma lines were depleted of RICTOR by esiRNA; the protein (d, n = 1 independent experiment) and the mRNA (e, n = 2 independent experiments, data are mean + s.d.) levels of indicated genes were assessed by western blot and qPCR, respectively. f–i, RICTOR was depleted in A2058 cells using esiRNA. f, The expression of the indicated proteins was assessed by western blot. n = 1 independent experiment. g, The expression of HIF1A, VEGF and GLUT1 was assessed by qPCR. n = 2 independent experiments, data are mean + s.d. h, i, Cell death in untreated (h) or 24-h vemurafenib-treated (i) control or RICTOR-depleted resistant BRAFV600E A2058 cells was quantified by FACS analysis. n = 2 independent experiments, data are mean + s.d. j, k, RICTOR association with RPL26 in the sensitive BRAFV600E SkMel5 (j) and in the resistant BRAFV600E A375 (k) melanoma cell lines was assessed by RPL26 immunoprecipitation. Indicated proteins were detected by western blot n = 1 independent experiment. l, Plot showing the correlation between drug sensitivity and mTORC2 activity. Drug sensitivity was quantified as the fold induced cell death after 24 h of 10 μM vemurafenib treatment as measured by FACS analysis. n = 3 independent experiments. mTORC2 activity was defined as Rictor recruitment to the ribosome and quantified by its binding to the ribosomal protein Rpl26. n = 1 independent experiment. The correlation coefficient (R, Pearson coefficient) and P value (two-sided t-test) were calculated. m, High HIF1α expression is linked with poor survival in a PanCancer analysis. One-sided Mantel–Cox test. n, o, Elevated HIF1α expression in poor survivors is concomitant with elevated RICTOR (n) and ELP1 (o) expression. n = 2,752 independent samples; two-sided Mann–Whitney U-test; data are mean ± 95% confidence interval, dots indicate the individual measurements of the n = 2,752 independent samples.
Supplementary Information
Supplementary Information
This file contains Supplementary Figure 1 (gel source data), the gating strategy and Supplementary Tables 2 and 4.
Supplementary Table 1
Analysis of SILAC data. SILAC row data file, GSEA analysis of SILAC upregulated (n=1337) and downregulated (n=1430) proteins, list of hypoxia genes, U34 sensitive codons (i.e. AAA, AAG, AAC) enrichment analysis and significance of HIF1α codon enrichment are displayed in the different sheets of the table (χ2 test).
Supplementary Table 3
Melanoma resistant signature. MITF anticorrelated gene signature was identified and associated with resistance towards BRAFi21.
Supplementary Table 5
A list of antibodies used in this study. Antibodies used in western blot, immunoprecipitation (IP) and immunohistochemistry (IHC) are listed.
Supplementary Table 6
A list of primers and RNA interference constructs used in this study. qPCR primers, shRNA sequences and esiRNA sequences are listed.
Rights and permissions
About this article
Cite this article
Rapino, F., Delaunay, S., Rambow, F. et al. Codon-specific translation reprogramming promotes resistance to targeted therapy. Nature 558, 605–609 (2018). https://doi.org/10.1038/s41586-018-0243-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0243-7
This article is cited by
-
Ribosome profiling: a powerful tool in oncological research
Biomarker Research (2024)
-
tRNA modification reprogramming contributes to artemisinin resistance in Plasmodium falciparum
Nature Microbiology (2024)
-
Quantitative analysis of tRNA abundance and modifications by nanopore RNA sequencing
Nature Biotechnology (2024)
-
RNA modifications in physiology and disease: towards clinical applications
Nature Reviews Genetics (2024)
-
Activation of CTU2 expression by LXR promotes the development of hepatocellular carcinoma
Cell Biology and Toxicology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.