Abstract
Sleep and wake have global effects on brain physiology, from molecular changes1,2,3,4 and neuronal activities to synaptic plasticity3,4,5,6,7. Sleep–wake homeostasis is maintained by the generation of a sleep need that accumulates during waking and dissipates during sleep8,9,10,11. Here we investigate the molecular basis of sleep need using quantitative phosphoproteomic analysis of the sleep-deprived and Sleepy mouse models of increased sleep need. Sleep deprivation induces cumulative phosphorylation of the brain proteome, which dissipates during sleep. Sleepy mice, owing to a gain-of-function mutation in the Sik3 gene12, have a constitutively high sleep need despite increased sleep amount. The brain proteome of these mice exhibits hyperphosphorylation, similar to that seen in the brain of sleep-deprived mice. Comparison of the two models identifies 80 mostly synaptic sleep-need-index phosphoproteins (SNIPPs), in which phosphorylation states closely parallel changes of sleep need. SLEEPY, the mutant SIK3 protein, preferentially associates with and phosphorylates SNIPPs. Inhibition of SIK3 activity reduces phosphorylation of SNIPPs and slow wave activity during non-rapid-eye-movement sleep, the best known measurable index of sleep need, in both Sleepy mice and sleep-deprived wild-type mice. Our results suggest that phosphorylation of SNIPPs accumulates and dissipates in relation to sleep need, and therefore SNIPP phosphorylation is a molecular signature of sleep need. Whereas waking encodes memories by potentiating synapses, sleep consolidates memories and restores synaptic homeostasis by globally downscaling excitatory synapses4,5,6. Thus, the phosphorylation–dephosphorylation cycle of SNIPPs may represent a major regulatory mechanism that underlies both synaptic homeostasis and sleep–wake homeostasis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cirelli, C. & Tononi, G. Changes in anti-phosphoserine and anti-phosphothreonine antibody binding during the sleep-waking cycle and after lesions of the locus coeruleus. Sleep Res. Online 1, 11–18 (1998).
Elliott, A. S., Huber, J. D., O’Callaghan, J. P., Rosen, C. L. & Miller, D. B. A review of sleep deprivation studies evaluating the brain transcriptome. Springerplus 3, 728 (2014).
Thompson, C. L. et al. Molecular and anatomical signatures of sleep deprivation in the mouse brain. Front. Neurosci. 4, 165 (2010).
Diering, G. H. et al. Homer1a drives homeostatic scaling-down of excitatory synapses during sleep. Science 355, 511–515 (2017).
Tononi, G. & Cirelli, C. Sleep and the price of plasticity: from synaptic and cellular homeostasis to memory consolidation and integration. Neuron 81, 12–34 (2014).
de Vivo, L. et al. Ultrastructural evidence for synaptic scaling across the wake/sleep cycle. Science 355, 507–510 (2017).
Vyazovskiy, V. V. & Harris, K. D. Sleep and the single neuron: the role of global slow oscillations in individual cell rest. Nat. Rev. Neurosci. 14, 443–451 (2013).
Borbely, A. A. A two process model of sleep regulation. Hum. Neurobiol. 1, 195–204 (1982).
Benington, J. H. Sleep homeostasis and the function of sleep. Sleep 23, 959–966 (2000).
Franken, P., Chollet, D. & Tafti, M. The homeostatic regulation of sleep need is under genetic control. J. Neurosci. 21, 2610–2621 (2001).
Vassalli, A. & Dijk, D. J. Sleep function: current questions and new approaches. Eur. J. Neurosci. 29, 1830–1841 (2009).
Funato, H. et al. Forward-genetics analysis of sleep in randomly mutagenized mice. Nature 539, 378–383 (2016).
Saper, C. B. & Fuller, P. M. Wake-sleep circuitry: an overview. Curr. Opin. Neurobiol. 44, 186–192 (2017).
Liu, S., Liu, Q., Tabuchi, M. & Wu, M. N. Sleep drive is encoded by neural plastic changes in a dedicated circuit. Cell 165, 1347–1360 (2016).
Lizcano, J. M. et al. LKB1 is a master kinase that activates 13 kinases of the AMPK subfamily, including MARK/PAR-1. EMBO J. 23, 833–843 (2004).
Erickson, B. K. et al. Evaluating multiplexed quantitative phosphopeptide analysis on a hybrid quadrupole mass filter/linear ion trap/orbitrap mass spectrometer. Anal. Chem. 87, 1241–1249 (2015).
McAlister, G. C. et al. MultiNotch MS3 enables accurate, sensitive, and multiplexed detection of differential expression across cancer cell line proteomes. Anal. Chem. 86, 7150–7158 (2014).
Weekes, M. P. et al. Quantitative temporal viromics: an approach to investigate host-pathogen interaction. Cell 157, 1460–1472 (2014).
Paulo, J. A. et al. Effects of MEK inhibitors GSK1120212 and PD0325901 in vivo using 10-plex quantitative proteomics and phosphoproteomics. Proteomics 15, 462–473 (2015).
Humphrey, S. J., James, D. E. & Mann, M. Protein phosphorylation: a major switch mechanism for metabolic regulation. Trends Endocrinol. Metab. 26, 676–687 (2015).
Greengard, P., Valtorta, F., Czernik, A. J. & Benfenati, F. Synaptic vesicle phosphoproteins and regulation of synaptic function. Science 259, 780–785 (1993).
Cesca, F., Baldelli, P., Valtorta, F. & Benfenati, F. The synapsins: key actors of synapse function and plasticity. Prog. Neurobiol. 91, 313–348 (2010).
Cantrell, A. R. et al. Molecular mechanism of convergent regulation of brain Na+ channels by protein kinase C and protein kinase A anchored to AKAP-15. Mol. Cell. Neurosci. 21, 63–80 (2002).
Tatsuki, F. et al. Involvement of Ca2+-dependent hyperpolarization in sleep duration in mammals. Neuron 90, 70–85 (2016).
Campbell, I. G. & Feinberg, I. NREM delta stimulation following MK-801 is a response of sleep systems. J. Neurophysiol. 76, 3714–3720 (1996).
Campbell, I. G. & Feinberg, I. Noncompetitive NMDA channel blockade during waking intensely stimulates NREM delta. J. Pharmacol. Exp. Ther. 276, 737–742 (1996).
Schaffer, B. E. et al. Identification of AMPK phosphorylation sites reveals a network of proteins involved in cell invasion and facilitates large-scale substrate prediction. Cell Metab. 22, 907–921 (2015).
Clark, K. et al. Phosphorylation of CRTC3 by the salt-inducible kinases controls the interconversion of classically activated and regulatory macrophages. Proc. Natl Acad. Sci. USA 109, 16986–16991 (2012).
Eng, J. K., McCormack, A. L. & Yates, J. R. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Peng, J., Elias, J. E., Thoreen, C. C., Licklider, L. J. & Gygi, S. P. Evaluation of multidimensional chromatography coupled with tandem mass spectrometry (LC/LC–MS/MS) for large-scale protein analysis: the yeast proteome. J. Proteome Res. 2, 43–50 (2003).
Elias, J. E. & Gygi, S. P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Kall, L., Canterbury, J. D., Weston, J., Noble, W. S. & MacCoss, M. J. Semi-supervised learning for peptide identification from shotgun proteomics datasets. Nat. Methods 4, 923–925 (2007).
Taus, T. et al. Universal and confident phosphorylation site localization using phosphoRS. J. Proteome Res. 10, 5354–5362 (2011).
Wang, X. et al. JUMP: a tag-based database search tool for peptide identification with high sensitivity and accuracy. Mol. Cell. Proteomics 13, 3663–3673 (2014).
Li, Y. et al. JUMPg: an integrative proteogenomics pipeline identifying unannotated proteins in human brain and cancer cells. J. Proteome Res. 15, 2309–2320 (2016).
Benjamini, Y., Krieger, A. M. & Yekutieli, D. Adaptive linear step-up procedures that control the false discovery rate. Biometrika 93, 491–507 (2006).
Wu, R. et al. Correct interpretation of comprehensive phosphorylation dynamics requires normalization by protein expression changes. Mol. Cell. Proteomics 10, M111 009654 (2011).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
The Gene Ontology Consortium. C. Expansion of the Gene Ontology knowledgebase and resources. Nucleic Acids Res. 45, D331–D338 (2017).
Mi, H. et al. PANTHER version 11: expanded annotation data from Gene Ontology and Reactome pathways, and data analysis tool enhancements. Nucleic Acids Res. 45, D183–D189 (2017).
Beacham, D., Ahn, M., Catterall, W. A. & Scheuer, T. Sites and molecular mechanisms of modulation of Na(v)1.2 channels by Fyn tyrosine kinase. J. Neurosci. 27, 11543–11551 (2007).
James, T. F. et al. The Nav1.2 channel is regulated by GSK3. Biochim. Biophys. Acta 1850, 832–844 (2015).
Siwek, M. E. et al. The CaV2.3 R-type voltage-gated Ca2+ channel in mouse sleep architecture. Sleep 37, 881–892 (2014).
Parker, M. J. et al. De novo, heterozygous, loss-of-function mutations in SYNGAP1 cause a syndromic form of intellectual disability. Am. J. Med. Genet. A. 167A, 2231–2237 (2015).
Carlisle, H. J. et al. Deletion of densin-180 results in abnormal behaviors associated with mental illness and reduces mGluR5 and DISC1 in the postsynaptic density fraction. J. Neurosci. 31, 16194–16207 (2011).
Soorya, L. et al. Prospective investigation of autism and genotype–phenotype correlations in 22q13 deletion syndrome and SHANK3 deficiency. Mol. Autism 4, 18 (2013).
Ahnaou, A., Raeymaekers, L., Steckler, T. & Drinkenbrug, W. H. Relevance of the metabotropic glutamate receptor (mGluR5) in the regulation of NREM–REM sleep cycle and homeostasis: evidence from mGluR5−/−mice. Behav. Brain Res. 282, 218–226 (2015).
Hagebeuk, E. E., van den Bossche, R. A. & de Weerd, A. W. Respiratory and sleep disorders in female children with atypical Rett syndrome caused by mutations in the CDKL5 gene. Dev. Med. Child Neurol. 55, 480–484 (2012).
Lonart, G., Tang, X., Simsek-Duran, F., Machida, M. & Sanford, L. D. The role of active zone protein Rab3 interacting molecule 1 alpha in the regulation of norepinephrine release, response to novelty, and sleep. Neuroscience 154, 821–831 (2008).
Iqbal, Z. et al. Homozygous and heterozygous disruptions of ANK3: at the crossroads of neurodevelopmental and psychiatric disorders. Hum. Mol. Genet. 22, 1960–1970 (2013).
von Stulpnagel, C. et al. SYNGAP1 mutation in focal and generalized epilepsy: a literature overview and a case report with special aspects of the EEG. Neuropediatrics 46, 287–291 (2015).
Mangatt, M. et al. Prevalence and onset of comorbidities in the CDKL5 disorder differ from Rett syndrome. Orphanet J. Rare Dis. 11, 39 (2016).
Fehr, S. et al. The CDKL5 disorder is an independent clinical entity associated with early-onset encephalopathy. Eur. J. Hum. Genet. 21, 266–273 (2013).
Jiang, P. et al. A systems approach identifies networks and genes linking sleep and stress: implications for neuropsychiatric disorders. Cell Reports 11, 835–848 (2015).
Welch, J. M. et al. Cortico-striatal synaptic defects and OCD-like behaviours in Sapap3-mutant mice. Nature 448, 894–900 (2007).
Bayes, A. et al. Comparative study of human and mouse postsynaptic proteomes finds high compositional conservation and abundance differences for key synaptic proteins. PLoS ONE 7, e46683 (2012).
Li, J. et al. Long-term potentiation modulates synaptic phosphorylation networks and reshapes the structure of the postsynaptic interactome. Sci. Signal. 9, rs8 (2016).
Uezu, A. et al. Identification of an elaborate complex mediating postsynaptic inhibition. Science 353, 1123–1129 (2016).
Gonzalez-Lozano, M. A. et al. Dynamics of the mouse brain cortical synaptic proteome during postnatal brain development. Sci. Rep. 6, 35456 (2016).
Weingarten, J. et al. The proteome of the presynaptic active zone from mouse brain. Mol. Cell. Neurosci. 59, 106–118 (2014).
Boyken, J. et al. Molecular profiling of synaptic vesicle docking sites reveals novel proteins but few differences between glutamatergic and GABAergic synapses. Neuron 78, 285–297 (2013).
Abul-Husn, N. S. et al. Systems approach to explore components and interactions in the presynapse. Proteomics 9, 3303–3315 (2009).
Biesemann, C. et al. Proteomic screening of glutamatergic mouse brain synaptosomes isolated by fluorescence activated sorting. EMBO J. 33, 157–170 (2014).
Distler, U. et al. In-depth protein profiling of the postsynaptic density from mouse hippocampus using data-independent acquisition proteomics. Proteomics 14, 2607–2613 (2014).
Loh, K. H. et al. Proteomic analysis of unbounded cellular compartments: synaptic clefts. Cell 166, 1295-1307 (2016).
Nakamura, Y. et al. Proteomic characterization of inhibitory synapses using a novel pHluorin-tagged γ-aminobutyric acid receptor, type A (GABAA), α2 subunit knock-in mouse. J. Biol. Chem. 291, 12394–12407 (2016).
de Hoon, M. J., Imoto, S., Nolan, J. & Miyano, S. Open source clustering software. Bioinformatics 20, 1453–1454 (2004).
Lee, E. E. et al. A protein kinase C phosphorylation motif in GLUT1 affects glucose transport and is mutated in GLUT1 deficiency syndrome. Mol. Cell 58, 845–853 (2015).
Kinoshita, E., Kinoshita-Kikuta, E., Takiyama, K. & Koike, T. Phosphate-binding tag, a new tool to visualize phosphorylated proteins. Mol. Cell. Proteomics 5, 749–757 (2006).
Vizcaino, J. A. et al. ProteomeXchange provides globally coordinated proteomics data submission and dissemination. Nat. Biotechnol. 32, 223–226 (2014).
Deutsch, E. W. et al. The ProteomeXchange consortium in 2017: supporting the cultural change in proteomics public data deposition. Nucleic Acids Res. 45, D1100–D1106 (2017).
Acknowledgements
We are grateful to M. Dong, S. Chen and H. Mirzaei for mass spectrometry assistance; J. Cohen, R. Greene and F. Shao for comments on the manuscript. Q.L. is a W.A. ‘Tex’ Moncrief Jr. Scholar in Medical Research. Y.Y. is a Virginia Murchison Linthicum Scholar in Medical Research and a CPRIT scholar in Cancer Research. This work was supported by the Welch foundation (I-1608 to Q.L.; I-1800 to Y.Y.), the National Institute of Health (GM111367 to Q.L.; R01AG047928 to J.P.; GM114160 to Y.Y.), JSPS KAKENHI (16K16639 to Z.W.; 17K15592 to J.M.; 26220207, 17H06095 to M.Y., H.F., Q.L.; 17H04023, 16K15187, 15H05942 to H.F.), JST CREST (JPMJCR1655 to M.Y.), FIRST program from JSPS to M.Y., Uehara and Takeda Foundations to M.Y. and the WPI program from Japan’s MEXT.
Reviewer information
Nature thanks D. Kirkpatrick and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Z.W., J.M. and Q.L. designed experiments with inputs from M.Y., H.F. and L.C. Z.W. received mass spectrometric training from Y.Y. Z.W., Y.L. and C.L. performed bioinformatics analysis with advice from Y.O. and J.P. J.M., Z.W., C.Ma and X.Y. performed biochemical studies. C.Mi., M.K., A.I., N.H.-H., S.K., X.G., J.M., Z.W. collected tissue samples for mass spectrometry. J.M., Z.W., T.L., X.G., S.Z. and M.S. completed EEG/EMG data analysis. D.K.-N., T.F. and S.T. produced genetically modified mice. J.M. and Z.W. made the figures. Q.L. and Z.W. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
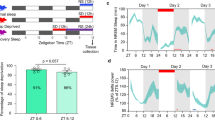
Extended Data Fig. 1 Sleep phenotype analysis of the sleep-deprived and Sleepy models.
a–e, Analysis of circadian (a) and mean (b) absolute NREMS delta power, absolute EEG power spectra (c), relative EEG power spectra (d) and duration (e) of NREMS, REMS and wake states of wild-type mice (n = 24) without (WT-basal) and with 6 h of sleep deprivation (WT-SD). f–i, Analysis of circadian (f) and mean (g) relative NREMS delta power, relative EEG power spectra (h) and duration (i) of NREMS, REMS and wake states of Sik3+/+ (WT, n = 24) and Sik3Slp/+ (Slp, n = 24) mice. Mean ± s.e.m., two-way ANOVA with Sidak’s test (a, c–f, h, i); Paired t-test, two-tailed (b); Mean, unpaired t-test, two-tailed (g). *(black), P < 0.05; *(cyan), P < 0.01; *(red), P < 0.001.
Extended Data Fig. 2 Analysis of global signalling changes in two models of increased sleep need.
a–m, Representative immunoblots using antibodies specific for 13 phosphorylation motifs to assess global signalling changes in whole brain lysates of two models. Blots represent three (sleep-deprived) or two (Sleepy) independent experiments. Quantitative analysis of immunoblots of all 14 phosphorylation-motif antibodies is shown in Fig. 1c. n = 12 (S6), 9 (SD6, RS3), 6 (wild-type, Sleepy).
Extended Data Fig. 3 Analysis of sleep phenotype and signalling changes after food-and-water deprivation in the baseline and sleep deprivation conditions.
a–c, Analysis of circadian (a) and mean (b) absolute NREMS delta power, and duration (c) of NREMS, REMS and wake states, of wild-type mice (n = 8) without (sham) or with 6 h of food-and-water deprivation (FD 6 h). d, Quantitative analysis of immunoblots with six phosphorylation-motif antibodies using whole brain lysates of sham and 6-h food-and-water deprived mice (n = 8) collected at ZT6. e–g, Analysis of circadian (e) and mean (f) absolute NREMS delta power, and duration (g) of NREMS, REMS and wake states, of wild-type mice (n = 11) without (SD + sham) or with 6-h food-and-water deprivation during 6-h sleep deprivation (SD + FD 6 h). h, Quantitative analysis of immunoblots with six phosphorylation-motif antibodies using whole brain lysates of SD + sham and SD + FD mice (n = 6) collected at ZT6. Mean ± s.e.m., two-way ANOVA, Sidak’s test (a, c, e, g); Paired t-test, two-tailed (b, f); Mean ± s.d., two-way ANOVA, Fisher’s LSD test (d, h). *(black), P < 0.05; ns, P > 0.05.
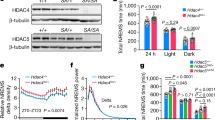
Extended Data Fig. 4 Quality assessment of proteomic and phosphoproteomic analysis.
a, Representative TMT quantification spectrum for the pS551-containing phosphopeptide from the skipped Sik3 exon-13 among phosphoproteomic data of the Sleepy model (two independent experiments). b–e, Quality assessment of one proteomic dataset (EX4, SlpWTpa2) by two search pipelines. Global distribution of protein quantification using Proteome Discoverer (PD v.2.1; n = 8,273) (b) and JUMP (v.1.12.1; n = 8,473) (c). Boxes correspond to the 25th, 50th and 75th percentiles of the data, whiskers extend to 1.5-fold of the interquartile range. A similar number of accepted proteins (1% FDR) were identified by two pipelines (d). Pearson correlation between the two pipelines was calculated for each PSM from quantified proteins by both pipelines (e). The vast majority (99.88%) of PSMs (n = 73,454) have R2 values larger than 0.9 (red dashed line). f, A Venn diagram showing overlaps of quantified proteins between whole brain proteomes of Sleepy and sleep-deprived models. g, Volcano plots showing comparative analysis of Sleepy/wild-type, SD6/RS3 and SD6/S6 proteomes. Multiple unpaired t-test (P value) followed by FDR (Q value) analysis. x axis, log2(fold change) in abundance; y axis, −log(Q value) of abundance change. The numbers of total (n), increased (in: Q < 0.2, red) and decreased (de: Q < 0.2, blue) subjects are shown. Orange dotted lines indicate Q = 0.2. h, Pearson correlation between normalized and unnormalized phosphopeptides in Sleepy/wild-type, SD6/RS3, SD6/S6 groups. The numbers of phosphopeptides that can be normalized are shown. i, Immunoblots were performed with phosphorylation-site specific antibodies to verify hyper-phosphorylation of several proteins in two models. Three or two independent experiments for sleep-deprived or Sleepy models, respectively. j, Quantitative analysis of immunoblots in i, normalized with whole protein abundance, for Sleepy (n = 6) and sleep-deprived (n = 9) models. Mean ± s.d., two-way ANOVA with Fisher’s LSD test. *(black), P < 0.05; *(cyan), P < 0.01; *(red), P < 0.001; ns, P > 0.05.
Extended Data Fig. 5 Liver phosphoproteome analysis of the sleep-deprived model.
a, Quantitative analysis of immunoblots with six phosphorylation-motif antibodies using whole liver lysates from the sleep-deprived model. n = 8 (S6), 10 (SD6), 7 (RS3). Mean ± s.d., two-way ANOVA with Fisher’s LSD test. *(black), P < 0.05; ns, P > 0.05. b, c, Volcano plots showing comparative analysis of liver phosphoproteomes in the SD6/RS3 (b) and SD6/S6 (c) groups. Multiple unpaired t-test (P value) followed by FDR (Q value) analysis. x axis, log2(fold change) in abundance; y axis, −log(Q value) of abundance change. The numbers of total (n), increased (in: Q < 0.2, red) and decreased (de: Q < 0.2, blue) subjects are shown. Orange dotted lines indicate Q = 0.2. d, A Venn diagram showing overlaps of significantly changed (Q < 0.2) phosphopeptides among the SD6/RS3 and SD6/S6 groups. e, f, Global ∆Ps analysis of all phosphoproteins identified in the SD6/RS3 (e) and SD6/S6 (f) groups of liver phosphoproteomes. Dotted lines, ∆Ps = ±2.4.
Extended Data Fig. 6 Examples of cumulative phosphorylation of SNIPPs and synaptic phosphoproteomic analysis of normal sleep–wake model.
a, b, A schematic of the domain structure of synapsin-122 (a) and Nav1.223,41,42 (b) that summarizes known phosphorylation sites, kinases and physiological functions. Synapsin-1 can be divided into five domains (domains A–E). Nav1.2 can be divided into cytoplasmic N-terminal (NT), C-terminal (CT), four homologous transmembrane domains (DI–DIV) and intracellular loops (DI–II, DII–III, DIII–IV). Amino acid numbers refer to the sequence of the mouse proteins. Sites 1–9 of synapsin-1 are designated according to the consensus in the literature. Phosphorylation sites that are undetected or unchanged in our experiments are labelled in grey, whereas those that exhibit significantly increased phosphorylation with sleep deprivation are shown in red. Dashed arrows indicate the presence of contrasting data for biological functions in the literature. c, Published forebrain PSD phosphoproteome results4 were used for comparative analysis between normal wake (W4) and sleep (S4) brains. d, Global ∆Ps analysis of all identified phosphoproteins in the W4/S4 group. Dotted lines (∆Ps = ±2.4). e, Quantitative ∆Ps analysis of SD1/SD0, SD3/SD0 and SD6/SD0 groups. Mean; one-way ANOVA, Tukey’s test (total, SNIPPs); unpaired t-test, two-tailed (total versus SNIPPs). *(red), P < 0.001.
Extended Data Fig. 7 Physiological functions of 80 SNIPPs.
a, A Venn diagram showing overlaps of the set of hyperphosphorylated proteins (∆Ps >2.4) between sleep-deprived and Sleepy models. b, A summary of 80 SNIPPs and their physiological functions. Stars mark the 13 SWA-SNIPPs (Fig. 3f). Gene names for annotated synaptic proteins are shown in bold.
Extended Data Fig. 8 Phosphorylation-state changes of SNIPPs correspond to changes of sleep need in NMDAR inhibition model.
a, Representative 8-s EEG and EMG from ZT0–ZT3 for NREMS, REMS and wake for vehicle or MK801-treated mice. b, Mean absolute NREMS delta power analysis of vehicle or MK801-injected mice (n = 14). Paired t-test, two-tailed. c–e, Analysis of absolute EEG power spectra (c), relative EEG power spectra (d) and duration (e) for vehicle or MK801-injected wild-type mice (n = 14). Mean ± s.e.m., two-way ANOVA with Sidak’s test. f, Volcano plot showing comparison between phosphoproteomes of MK801 and vehicle treated mice. Orange dotted line, Q = 0.2. Multiple unpaired t-test (P value) followed by FDR (Q value) analysis. g, Phosphorylation state of synapsin-1 was assessed by SDS–PAGE followed by phospho-tag (top) and immunoblotting with anti-synapsin-1 antibody (bottom). The Rf value of 1.0 is defined as the position of bromphenol blue dye (two independent experiments). h, Quantitative ∆Ps analysis of MK801/vehicle group. Mean, unpaired t-test, two-tailed. i, Percentage of synaptic proteins among the total, hypophosphorylated and hyperphosphorylated proteins in the MK801/vehicle group. χ2 test, two-sided. j, Venn diagram showing overlaps of hyperphosphorylated proteins (∆Ps >2.4) among all three (Sleepy, SD and MK801) models. *(black), P < 0.05; *(cyan), P < 0.01; *(red), P < 0.001.
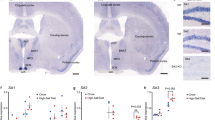
Extended Data Fig. 9 SLEEPY causes constitutively high sleep need by preferentially associating with and phosphorylating SNIPPs.
a, Experimental design for comparing the interactomes of SIK3 and SLEEPY from whole brain lysates. b, Summary of SIK3 and SLEEPY interacting proteins (ip) and preferential interacting proteins (pip). c, Gene-annotation enrichment analysis of 289 SLEEPY-preferential interacting proteins (SLEEPY-pip). GO cellular component enrichment analysis using all 22,262 genes of Mus musculus as reference (Ref). Fisher’s exact with FDR multiple test correction was used to determine statistical significance. Top 10 GO terms of fold enrichment (FDR <0.0001), the gene number of SLEEPY-pip and Ref in each term are shown. d, e, Volcano plots showing phosphorylation changes of all putative AMPK substrates in the Sleepy/wild-type group (d) or from the 28 SLEEPY-pip SNIPPs (e). Orange dotted lines, Q = 0.2. f, In vitro kinase assay of recombinant SLEEPY and SIK3, and immunoblotting with AMPK phosphorylation motif antibody (two independent experiments). g–i, Volcano plot showing comparative analysis of whole brain phosphoproteomes (g), all putative AMPK substrates (h) or from 28 SLEEPY-pip SNIPPs (i) in the HG/vehicle (Slp) group. Orange dotted lines, Q = 0.2. j, Quantitative ∆Ps analysis of 190 hyperphosphorylated proteins and 52 hypophosphorylated proteins in the HG/vehicle (Slp) group. Dotted lines, ∆Ps = ±2.4. k–m, Analysis of absolute EEG power spectra (k), relative EEG power spectra (l) and duration (m) of NREMS, REMS and wake states of Sik3Slp/+ (Slp, n = 14) mice injected with vehicle (Veh) or 8 mg/kg HG at ZT6 and ZT9. Multiple unpaired t-test (P value) followed by FDR (Q value) analysis (d, e, g–i). Mean, one-way ANOVA with Dunnett’s test (j). Mean ± s.e.m., two-way ANOVA with Sidak’s test (k–m). *(black), P < 0.05; *(cyan), P < 0.01; *(red), P < 0.001; ns, P > 0.05.
Extended Data Fig. 10 Inhibition of SIK3 kinase activity reduces phosphorylation of AMPK substrates in sleep-deprived wild-type brains.
a–c, Volcano plots showing phosphorylation changes of all putative AMPK substrates in the SD6/RS3 (a), SD6/S6 (b) and time-course sleep-deprivation groups (c). Orange dotted lines, Q = 0.2. d, e, Volcano plots showing comparative analysis of whole brain phosphoproteome (d) and phosphorylation changes of all putative AMPK substrates (e) in the HG/vehicle (WT-SD) group. Orange dotted lines, Q = 0.2. f–h, Analysis of absolute EEG power spectra (f), relative EEG power spectra (g) and duration (h) of NREMS, REMS and wake states of sleep-deprived (ZT0–ZT6) wild-type (n = 16) mice injected with vehicle (Veh) or 8 mg/kg HG at ZT0 and ZT3. Multiple unpaired t-test (P value) followed by FDR (Q value) analysis (a–e). Mean ± s.e.m., two-way ANOVA with Sidak’s test (f–h). *(black), P < 0.05; *(cyan), P < 0.01; *(red), P < 0.001; ns, P > 0.05.
Supplementary information
Supplementary Figure 1
This file contains the uncropped scans with size marker indications.
Supplementary Information
This file contains Supplementary Discussions 1-4.
Supplementary Table 1
Supplementary Table 1 (data lists of quantitative proteomic analysis.) can be downloaded from: ftp://massive.ucsd.edu/MSV000081865/updates/2018-04-27_wang_zhiqiang_99_8af04504/other/. The full description and datasets for all four proteomic experiments were listed and statistical analysis and n numbers for all comparisons were shown.
Supplementary Table 2
Supplementary Table 2 (Data lists of quantitative phosphoproteomic analysis) can be downloaded from: ftp://massive.ucsd.edu/MSV000081865/updates/2018-04-27_wang_zhiqiang_99_8af04504/other/. The full description and datasets for all phosphoproteomic experiments were listed and statistical analysis and n numbers for all comparisons were shown.
Supplementary Table 3
Data lists of protein phosphorylation state analysis. The full protein phospho-state analysis description and datasets for all comparisons were listed.
Supplementary Table 4
Databases for functional analysis. The sleep phenotypes, molecular and neuronal functions of 80 SNIPPs; an integrated synaptic protein database; complete data for AMPK substrate prediction were listed.
Supplementary Table 5
Data lists of PSD phosphoproteomic analysis of normal wake and sleep mouse brains. Raw data and protein phosphorylation state analysis of normal sleep-wake model were listed.
Supplementary Table 6
Data lists of SIK3 and SLEEPY interactomes. The full description and datasets for SIK3 and SLEEPY interactome analysis were listed. Criteria used to define the interacting protein and n numbers for all comparisons were shown.
Supplementary Table 7
Statistical analysis and sleep experiment figure layouts. The complete sample size, statistical test method and precise value results for each comparison were reported. The organization of sleep experiments related sleep phenotype results were listed.
Rights and permissions
About this article
Cite this article
Wang, Z., Ma, J., Miyoshi, C. et al. Quantitative phosphoproteomic analysis of the molecular substrates of sleep need. Nature 558, 435–439 (2018). https://doi.org/10.1038/s41586-018-0218-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0218-8
This article is cited by
-
nNOS in Erbb4-positive neurons regulates GABAergic transmission in mouse hippocampus
Cell Death & Disease (2024)
-
Knockdown of SIK3 in the CA1 Region can Reduce Seizure Susceptibility in Mice by Inhibiting Decreases in GABAAR α1 Expression
Molecular Neurobiology (2024)
-
SIK3-HDAC4 signaling pathway: the switch for transition between sleep and wakefulness
Molecular Biomedicine (2023)
-
Abnormal patterns of sleep and waking behaviors are accompanied by neocortical oscillation disturbances in an Ank3 mouse model of epilepsy-bipolar disorder comorbidity
Translational Psychiatry (2023)
-
Intracellular chloride regulation mediates local sleep pressure in the cortex
Nature Neuroscience (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.