Abstract
G-protein-coupled receptors comprise the largest family of mammalian transmembrane receptors. They mediate numerous cellular pathways by coupling with downstream signalling transducers, including the hetrotrimeric G proteins Gs (stimulatory) and Gi (inhibitory) and several arrestin proteins. The structural mechanisms that define how G-protein-coupled receptors selectively couple to a specific type of G protein or arrestin remain unknown. Here, using cryo-electron microscopy, we show that the major interactions between activated rhodopsin and Gi are mediated by the C-terminal helix of the Gi α-subunit, which is wedged into the cytoplasmic cavity of the transmembrane helix bundle and directly contacts the amino terminus of helix 8 of rhodopsin. Structural comparisons of inactive, Gi-bound and arrestin-bound forms of rhodopsin with inactive and Gs-bound forms of the β2-adrenergic receptor provide a foundation to understand the unique structural signatures that are associated with the recognition of Gs, Gi and arrestin by activated G-protein-coupled receptors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
21 June 2018
In the PDF version of this Article, owing to a typesetting error, an incorrect figure was used for Extended Data Fig. 5; the correct figure was used in the HTML version. This has been corrected online.
References
Fredriksson, R., Lagerström, M. C., Lundin, L. G. & Schiöth, H. B. The G-protein-coupled receptors in the human genome form five main families. Phylogenetic analysis, paralogon groups, and fingerprints. Mol. Pharmacol. 63, 1256–1272 (2003).
Neves, S. R., Ram, P. T. & Iyengar, R. G protein pathways. Science 296, 1636–1639 (2002).
Gurevich, E. V. & Gurevich, V. V. Arrestins: ubiquitous regulators of cellular signaling pathways. Genome Biol. 7, 236 (2006).
Zhou, X. E., Melcher, K. & Xu, H. E. Understanding the GPCR biased signaling through G protein and arrestin complex structures. Curr. Opin. Struct. Biol. 45, 150–159 (2017).
Palczewski, K. et al. Crystal structure of rhodopsin: a G protein-coupled receptor. Science 289, 739–745 (2000).
Cherezov, V. et al. High-resolution crystal structure of an engineered human β2-adrenergic G protein-coupled receptor. Science 318, 1258–1265 (2007).
Rosenbaum, D. M. et al. GPCR engineering yields high-resolution structural insights into beta2-adrenergic receptor function. Science 318, 1266–1273 (2007).
Xu, F. et al. Structure of an agonist-bound human A2A adenosine receptor. Science 332, 322–327 (2011).
Lebon, G. et al. Agonist-bound adenosine A2A receptor structures reveal common features of GPCR activation. Nature 474, 521–525 (2011).
Choe, H. W. et al. Crystal structure of metarhodopsin II. Nature 471, 651–655 (2011).
Wang, C. et al. Structural basis for molecular recognition at serotonin receptors. Science 340, 610–614 (2013).
Standfuss, J. et al. The structural basis of agonist-induced activation in constitutively active rhodopsin. Nature 471, 656–660 (2011).
Rosenbaum, D. M. et al. Structure and function of an irreversible agonist–β2 adrenoceptor complex. Nature 469, 236–240 (2011).
Rasmussen, S. G. et al. Crystal structure of the β2 adrenergic receptor–Gs protein complex. Nature 477, 549–555 (2011).
Zhang, Y. et al. Cryo-EM structure of the activated GLP-1 receptor in complex with a G protein. Nature 546, 248–253 (2017).
Liang, Y. L. et al. Phase-plate cryo-EM structure of a class B GPCR-G-protein complex. Nature 546, 118–123 (2017).
Flock, T. et al. Selectivity determinants of GPCR–G-protein binding. Nature 545, 317–322 (2017).
Palczewski, K. G protein-coupled receptor rhodopsin. Annu. Rev. Biochem. 75, 743–767 (2006).
Zhou, X. E., Melcher, K. & Xu, H. E. Structure and activation of rhodopsin. Acta Pharmacol. Sin. 33, 291–299 (2012).
Hamm, H. E. How activated receptors couple to G proteins. Proc. Natl Acad. Sci. USA 98, 4819–4821 (2001).
Van Meurs, K. P. et al. Deduced amino acid sequence of bovine retinal Goα: similarities to other guanine nucleotide-binding proteins. Proc. Natl Acad. Sci. USA 84, 3107–3111 (1987).
Lerea, C. L., Somers, D. E., Hurley, J. B., Klock, I. B. & Bunt-Milam, A. H. Identification of specific transducin α subunits in retinal rod and cone photoreceptors. Science 234, 77–80 (1986).
Kang, Y. et al. Crystal structure of rhodopsin bound to arrestin by femtosecond X-ray laser. Nature 523,561–567 (2015).
Zhou, X. E. et al. Identification of phosphorylation codes for arrestin recruitment by G protein-coupled receptors. Cell 170, 457–469 (2017).
Maeda, S. et al. Crystallization scale preparation of a stable GPCR signaling complex between constitutively active rhodopsin and G-protein. PLoS One 9, e98714 (2014).
Liu, P. et al. The structural basis of the dominant negative phenotype of the Gαi1β1γ2 G203A/A326S heterotrimer. Acta Pharmacol. Sin. 37, 1259–1272 (2016).
Van Eps, N. et al. Conformational equilibria of light-activated rhodopsin in nanodiscs. Proc. Natl Acad. Sci. USA 114, E3268–E3275 (2017).
Van Eps, N. et al. Gi- and Gs-coupled GPCRs show different modes of G-protein binding. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.1721896115 (2018).
Oldham, W. M., Van Eps, N., Preininger, A. M., Hubbell, W. L. & Hamm, H. E. Mechanism of the receptor-catalyzed activation of heterotrimeric G proteins. Nat. Struct. Mol. Biol. 13, 772–777 (2006).
Oldham, W. M. & Hamm, H. E. Heterotrimeric G protein activation by G-protein-coupled receptors. Nat. Rev. Mol. Cell Biol. 9, 60–71 (2008).
Marin, E. P., Krishna, A. G. & Sakmar, T. P. Rapid activation of transducin by mutations distant from the nucleotide-binding site: evidence for a mechanistic model of receptor-catalyzed nucleotide exchange by G proteins. J. Biol. Chem. 276, 27400–27405 (2001).
Marin, E. P., Krishna, A. G. & Sakmar, T. P. Disruption of the α5 helix of transducin impairs rhodopsin-catalyzed nucleotide exchange. Biochemistry 41, 6988–6994 (2002).
Garcia, P. D., Onrust, R., Bell, S. M., Sakmar, T. P. & Bourne, H. R. Transducin-α C-terminal mutations prevent activation by rhodopsin: a new assay using recombinant proteins expressed in cultured cells. EMBO J. 14, 4460–4469 (1995).
Onrust, R. et al. Receptor and βγ binding sites in the alpha subunit of the retinal G protein transducin. Science 275, 381–384 (1997).
Skiba, N. P., Bae, H. & Hamm, H. E. Mapping of effector binding sites of transducin alpha-subunit using Gαt/Gαi1 chimeras. J. Biol. Chem. 271, 413–424 (1996).
Sun, D. et al. Probing Gαi1 protein activation at single-amino acid resolution. Nat. Struct. Mol. Biol. 22, 686–694 (2015).
Itoh, Y., Cai, K. & Khorana, H. G. Mapping of contact sites in complex formation between light-activated rhodopsin and transducin by covalent crosslinking: use of a chemically preactivated reagent. Proc. Natl Acad. Sci. USA 98, 4883–4887 (2001).
Subramaniam, S., Gerstein, M., Oesterhelt, D. & Henderson, R. Electron diffraction analysis of structural changes in the photocycle of bacteriorhodopsin. EMBO J. 12, 1–8 (1993).
Subramaniam, S. & Henderson, R. Molecular mechanism of vectorial proton translocation by bacteriorhodopsin. Nature 406, 653–657 (2000).
Slessareva, J. E. et al. Closely related G-protein-coupled receptors use multiple and distinct domains on G-protein α-subunits for selective coupling. J. Biol. Chem. 278, 50530–50536 (2003).
Kling, R. C., Lanig, H., Clark, T. & Gmeiner, P. Active-state models of ternary GPCR complexes: determinants of selective receptor–G-protein coupling. PloS One 8, (2013).
DeVree, B. T. et al. Allosteric coupling from G protein to the agonist-binding pocket in GPCRs. Nature 535, 182–186 (2016).
Dror, R. O. et al. Signal transduction. Structural basis for nucleotide exchange in heterotrimeric G proteins. Science 348, 1361–1365 (2015).
Dickson, B. M., de Waal, P. W., Ramjan, Z. H., Xu, H. E. & Rothbart, S. B. A fast, open source implementation of adaptive biasing potentials uncovers a ligand design strategy for the chromatin regulator BRD4. J. Chem. Phys. 145, 154113 (2016).
Huang, W. J. et al. Structural insights into μ-opioid receptor activation. Nature 524, 315–321 (2015).
Rose, A. S. et al. Position of transmembrane helix 6 determines receptor G protein coupling specificity. J. Am. Chem. Soc. 136, 11244–11247 (2014).
Caro, L. N. et al. Rapid and facile recombinant expression of bovine rhodopsin in HEK293S GnTI− cells using a PiggyBac inducible system. Methods Enzymol. 556, 307–330 (2015).
Hornsby, M. et al. A high through-put platform for recombinant antibodies to folded proteins. Mol. Cell Proteomics 14, 2833–2847 (2015).
Paduch, M. & Kossiakoff, A. A. Generating conformation and complex-specific synthetic antibodies. Methods Mol. Biol. 1575, 93–119 (2017).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Brilot, A. F. et al. Beam-induced motion of vitrified specimen on holey carbon film. J. Struct. Biol. 177, 630–637 (2012).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Wang, R. Y. et al. Automated structure refinement of macromolecular assemblies from cryo-EM maps using Rosetta. eLife 5, e17219 (2016).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Manglik, A. et al. Structural insights into the dynamic process of β2-adrenergic receptor signaling. Cell 161, 1101–1111 (2015).
Šali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Mahalingam, M., Martínez-Mayorga, K., Brown, M. F. & Vogel, R. Two protonation switches control rhodopsin activation in membranes. Proc. Natl Acad. Sci. USA 105, 17795–17800 (2008).
Ranganathan, A., Dror, R. O. & Carlsson, J. Insights into the role of Asp792.50 in β2 adrenergic receptor activation from molecular dynamics simulations. Biochemistry 53, 7283–7296 (2014).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Zoete, V., Cuendet, M. A., Grosdidier, A. & Michielin, O. SwissParam: a fast force field generation tool for small organic molecules. J. Comput. Chem. 32, 2359–2368 (2011).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Dickson, B. M., de Waal, P. W., Ramjan, Z. H., Xu, H. E. & Rothbart, S. B. A fast, open source implementation of adaptive biasing potentials uncovers a ligand design strategy for the chromatin regulator BRD4. J. Chem. Phys. 145, 154113 (2016).
Dickson, B. M. Overfill protection and hyperdynamics in adaptively biased simulations. J. Chem. Theory Comput. 13, 5925–5932 (2017).
Kohlhoff, K. J. et al. Cloud-based simulations on Google Exacycle reveal ligand modulation of GPCR activation pathways. Nat. Chem. 6, 15–21 (2014).
Alexander, S. P. H. et al. The Concise Guide To Pharmacology 2017/18: G protein-coupled receptors. Br. J. Pharmacol. 174 (Suppl. 1), S17–S129 (2017).
Acknowledgements
Cryo-EM data were collected at the David Van Andel Advanced Cryo-Electron Microscopy Suite in the Van Andel Research Institute. This work was supported in part by the National Institutes of Health grant, DK071662, American Asthma Foundation, Jay and Betty Van Andel Foundation, Ministry of Science and Technology (China) grants 2012ZX09301001 and 2012CB910403, 2013CB910600, XDB08020303, 2013ZX09507001 (to H.E.X.), GM117372 (to A.K.), GM0875119 (to A.A.K.), grant from Pfizer (to A.A.K.), the National Natural Science Foundation 31770796 (to Y.J.), the Canada Excellence Research Chairs program (to O.P.E.), the Canadian Institute for Advanced Research (to O.P.E.), the Anne and Max Tanenbaum Chair in Neuroscience (to O.P.E.), by funds from the Center for Cancer Research, National Cancer Institute, NIH, Bethesda, MD (to S.S.), and by federal funds from the Frederick National Laboratory for Cancer Research, National Institutes of Health, under contract HHSN261200800001E. We thank H. Li and W. Lü for help with analysing the cryo-EM data and for advice on refinement, L. Bai and Z. Yuan for advice on 3D reconstruction, V. Falconieri for assistance with figure preparation, the HPC team at VARI for computational support, D. Nadziejka for manuscript editing, and B. Dickson for consultation on molecular dynamics simulation.
Author information
Authors and Affiliations
Contributions
Y.K. initiated the project, prepared samples, performed data acquisition and structure determination, and prepared the figures and manuscript writing; H.E.X. and K.M. conceived the project and designed the research, and wrote the paper with contributions from all authors; O.K., X.E.Z., A.B. and S.S. performed image processing, structure determination, figure preparation, and manuscript writing; P.W.d.W. performed computational experiments, analysed the structure, prepared figures, and manuscript writing; P.D., S.M., S.E. and A.A.K. designed and performed Fab selection; N.V.E., T.M. and O.P.E. designed and performed DEER experiments; X.G., Y.Y., P.L. and Y.J. performed cell-based assays; G.Z. and X.M. helped with data collection.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Purification, characterization and cryo-EM images of the Rho–Gi–Fab complex.
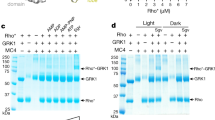
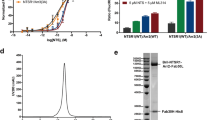
a, Representative elution profile of the purified Rho–Gi–Fab_G50 complex on Superdex 200 10/300 gel filtration. b, SDS–PAGE analysis of the complex after gel filtration. c, The inability of rhodopsin to stimulate the Gs-mediated signalling as assayed by the cAMP-driven luciferase reporter assays. The glucagon-like peptide 1 receptor (GLP-1R) shows stronger Gs-meditated signalling with the agonist GLP-1 (n = 3 independent experiments). Data are mean ± s.d. d, An overall view of rhodopsin showing the three intramolecular distances between two nitroxide N–O bonds based on the models of the R1 nitroxide pairs Y74R1-Q225R1, Y74R1-R252R1 and Y74R1-M308R1, respectively (Y742.41, Q2255.60, R2526.35, M3087.55; superscripts denote Ballesteros–Weinstein numbering). R1 side-chain modelling details have been described previously27. e, Similar DEER distance distributions of TM6 and TM7 to TM2 of rhodopsin bound to Gi and Gt. f, Time domain data of DEER measurements.
Extended Data Fig. 2 Cryo-EM images and single-particle analysis of the Rho–Gi–Fab complex.
a, Representative cryo-EM micrograph of Rho–Gi–Fab complex. Examples of particle projections are circled. b, Reference-free two-dimensional class averages of the complex in digitonin micelles. c, Half-map Fourier shell correlation (FSC) plots as produced by RELION with the mask used shown as an inset. d, FSC curve of model versus the full map, as well as FSC curves obtained for a model refined against a half-map and compared to the two half-maps as well as the full model. The r.m.s.d. between the model refined against half-map and compared to the full map, and the one refined against the full map is 0.984 Å, and their corresponding FSCs against the final map show a resolution difference at the 0.5-cutoff of approximately 0.1 Å. e, Particle classification and refinement. f, Local resolution map of the rhodopsin–Gi complex.
Extended Data Fig. 3 Electron microscopy density map of rhodopsin–Gi complex.
a–c, Three views of the electron microscopy density map of the rhodopsin–Gαi interface. d, Electron microscopy density map of all rhodopsin transmembrane helices and helix 8. e–g, An overall view of the rhodoposin–Gαi interface (e), and electron microscopy density map for the TM6 of rhodopsin (f) and the α5-helix of Gαi (g).
Extended Data Fig. 4 The rhodopsin–Gi interface and disulfide crosslinking of rhodopsin with Gαi.
a, The rhodopsin–Gi interface surrounding the G352 residue of Gαi α5-helix. Not all side chains shown are visible in the map but shown here for illustrating their Cα positions to facilitate understanding of data in panel b. b, Lack of disulfide crosslinking of G352C of Gi with surrounding residues from rhodopsin (compare with d; n = 3 independent experiments). c, Interactions at the interface between ICL2 of rhodopsin and αN helix of Gαi. The side chains are not visible in the map but shown here for illustrating their Cα positions. d, Demonstration that E28C of Gαi can be disulfide cross-linked to rhodopsin residues N145CICL2 and F146C ICL2 (n = 3 independent experiments).
Extended Data Fig. 5 Structural comparison of Gi-bound rhodopsin, Gs-bound GLP-1R, and Gs-bound CTR, and the role of α4-helix of Gα in receptor selectivity.
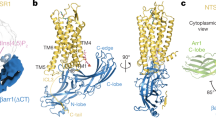
a, b, Side and cytoplasmic views of Gi-bound rhodopsin (orange) overlaid with Gs-bound GLP-1R (PDB code 5VAI, light blue, black arrows indicate differences in helix positions). c, d, Side and cytoplasmic views of Gi-bound rhodopsin (orange) overlaid with Gs-bound CTR (PDB code 5UZ7, grey). e, f, Side-by-side comparison of the rhodopsin–Gi complex (e) with the β2AR–Gs complex (f). g. An overlay of the rhodopsin–Gi complex with the β2AR–Gs complex reveals possible collision of TM5 of β2AR with α4-helix of Gαi.
Extended Data Fig. 6 The mechanism of rhodopsin-mediated Gi activation.
a, b, Superposition of the rhodopsin–Gi complex with the inactive GDP-bound Gi (PDB code 1GG2) reveals separation of the AHD from the Ras domain of Gαi (a) and conformational changes in the α5-helix (b). c, d, Side-by-side comparison of the GDP-binding site of the Gαi Ras domain in the inactive GDP-bound Gαi (c) and nucleotide-free state Gαi with GDP added for comparison (d).
Extended Data Fig. 7 Collective variables for mABP simulations and free-energy landscapes of mABP simulations.
a, To bias movement between TM6 relative to that of the receptor bundle, two centre-of-geometry (COG) distance collective variables (CVs) were implemented into fABMACS66. CV1 and CV2 are COG distances between selected atoms of TM6 to TM1/2 and TM6 to TM3/4 respectively. Collective variable atoms for the rhodopsin simulation are highlighted. b, COG collective variable formula and the CV1 and CV2 distances. c, Potential energy surface reveals that CV1 and CV2 distances are larger in the Gs-coupled receptors (A2AR and β2AR) than those in the Gi-coupled receptors (mOR1 and rhodopsin).
Extended Data Fig. 8 Enrichment profiles for Gi and Gs coupling receptors.
a–c, Relative probability of hydrophobic and polar residues for Gi (n = 76) and Gs (n = 25) coupling receptors. Residues with relative enrichments over 20% were mapped onto the structures of Gs-bound β2AR (b) and Gi-bound rhodopsin (c). GPCR principal coupling was previously defined68. d–f, Interaction network of TM6.36 of β2AR, A2AR and rhodopsin with the G protein α5-helix. g, Hydrogen bonding between TM3.36 and the backbone of TM6.
Supplementary information
Video 1: Structural difference between inactive Rho and Gi-bound Rho.
Conformational changes in the transmembrane helices of rhodopsin illustrated by morphing from inactive state to Gi-bound state.
Video 2: Structural difference between arrestin-bound Rho and Gi-bound Rho.
Conformational changes in the transmembrane helices of rhodopsin illustrated by morphing from arrestin-bound state to Gi-bound state.
Video 3: Structural difference between Gs-bound β2-AR and Gi-bound Rho.
Conformational changes in the transmembrane helices of GPCR illustrated by morphing from β2-AR in Gs-bound state to rhodopsin in Gi-bound state.
Rights and permissions
About this article
Cite this article
Kang, Y., Kuybeda, O., de Waal, P.W. et al. Cryo-EM structure of human rhodopsin bound to an inhibitory G protein. Nature 558, 553–558 (2018). https://doi.org/10.1038/s41586-018-0215-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0215-y
This article is cited by
-
Constitutive activation mechanism of a class C GPCR
Nature Structural & Molecular Biology (2024)
-
Cryo-electron microscopy for GPCR research and drug discovery in endocrinology and metabolism
Nature Reviews Endocrinology (2024)
-
Binding kinetics drive G protein subtype selectivity at the β1-adrenergic receptor
Nature Communications (2024)
-
Scaling up Functional Analyses of the G Protein-Coupled Receptor Rhodopsin
Journal of Molecular Evolution (2024)
-
Structural insights into neurokinin 3 receptor activation by endogenous and analogue peptide agonists
Cell Discovery (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.