Abstract
Although plasma proteins have important roles in biological processes and are the direct targets of many drugs, the genetic factors that control inter-individual variation in plasma protein levels are not well understood. Here we characterize the genetic architecture of the human plasma proteome in healthy blood donors from the INTERVAL study. We identify 1,927 genetic associations with 1,478 proteins, a fourfold increase on existing knowledge, including trans associations for 1,104 proteins. To understand the consequences of perturbations in plasma protein levels, we apply an integrated approach that links genetic variation with biological pathway, disease, and drug databases. We show that protein quantitative trait loci overlap with gene expression quantitative trait loci, as well as with disease-associated loci, and find evidence that protein biomarkers have causal roles in disease using Mendelian randomization analysis. By linking genetic factors to diseases via specific proteins, our analyses highlight potential therapeutic targets, opportunities for matching existing drugs with new disease indications, and potential safety concerns for drugs under development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Albert, F. W. & Kruglyak, L. The role of regulatory variation in complex traits and disease. Nat. Rev. Genet. 16, 197–212 (2015).
Liu, Y. et al. Quantitative variability of 342 plasma proteins in a human twin population. Mol. Syst. Biol. 11, 786 (2015).
Suhre, K. et al. Connecting genetic risk to disease end points through the human blood plasma proteome. Nat. Commun. 8, 14357 (2017).
Yao, C. et al. Genome-wide association study of plasma proteins identifies putatively causal genes, proteins, and pathways for cardiovascular disease. Preprint at https://www.biorxiv.org/content/early/2017/05/12/136523 (2017).
de Vries, P. S. et al. Whole-genome sequencing study of serum peptide levels: the Atherosclerosis Risk in Communities study. Hum. Mol. Genet. 26, 3442–3450 (2017).
Wu, L. et al. Variation and genetic control of protein abundance in humans. Nature 499, 79–82 (2013).
Battle, A. et al. Impact of regulatory variation from RNA to protein. Science 347, 664–667 (2015).
Rohloff, J. C. et al. Nucleic acid ligands with protein-like side chains: modified aptamers and their use as diagnostic and therapeutic agents. Mol. Ther. Nucleic Acids 3, e201 (2014).
Di Angelantonio, E. et al. Efficiency and safety of varying the frequency of whole blood donation (INTERVAL): a randomised trial of 45 000 donors. Lancet 390, 2360–2371 (2017).
Astle, W. J. et al. The allelic landscape of human blood cell trait variation and links to common complex disease. Cell 167, 1415–1429.e19 (2016).
Burgess, S., Scott, R. A., Timpson, N. J., Davey Smith, G. & Thompson, S. G. Using published data in Mendelian randomization: a blueprint for efficient identification of causal risk factors. Eur. J. Epidemiol. 30, 543–552 (2015).
Stranger, B. E. et al. Patterns of cis regulatory variation in diverse human populations. PLoS Genet. 8, e1002639 (2012).
Lundberg, M., Eriksson, A., Tran, B., Assarsson, E. & Fredriksson, S. Homogeneous antibody-based proximity extension assays provide sensitive and specific detection of low-abundant proteins in human blood. Nucleic Acids Res. 39, e102 (2011).
Walter, K. et al. The UK10K project identifies rare variants in health and disease. Nature 526, 82–90 (2015).
Westra, H.-J. et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat. Genet. 45, 1238–1243 (2013).
Lappalainen, T. et al. Transcriptome and genome sequencing uncovers functional variation in humans. Nature 501, 506–511 (2013).
Schadt, E. E. et al. Mapping the genetic architecture of gene expression in human liver. PLoS Biol. 6, e107 (2008).
Zeller, T. et al. Genetics and beyond—the transcriptome of human monocytes and disease susceptibility. PLoS ONE 5, e10693 (2010).
Nai, A. et al. TMPRSS6 rs855791 modulates hepcidin transcription in vitro and serum hepcidin levels in normal individuals. Blood 118, 4459–4462 (2011).
Carrasquillo, M. M. et al. Genome-wide screen identifies rs646776 near sortilin as a regulator of progranulin levels in human plasma. Am. J. Hum. Genet. 87, 890–897 (2010).
Gooptu, B., Dickens, J. A. & Lomas, D. A. The molecular and cellular pathology of α1-antitrypsin deficiency. Trends Mol. Med. 20, 116–127 (2014).
Stacey, D. et al. ProGeM: A framework for the prioritisation of candidate causal genes at molecular quantitative trait loci. https://doi.org/10.1101/230094 (2017).
Liu, J. Z. et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat. Genet. 47, 979–986 (2015).
Di Narzo, A. F. et al. High-throughput characterization of blood serum proteomics of IBD patients with respect to aging and genetic factors. PLoS Genet. 13, e1006565 (2017).
Huang, H. et al. Fine-mapping inflammatory bowel disease loci to single-variant resolution. Nature 547, 173–178 (2017).
Lyons, P. A. et al. Genetically distinct subsets within ANCA-associated vasculitis. N. Engl. J. Med. 367, 214–223 (2012).
Merkel, P. A. et al. Identification of functional and expression polymorphisms associated with risk for anti-neutrophil cytoplasmic autoantibody-associated vasculitis. Arthritis Rheumatol. 69, 1054–1066 (2017).
Battle, A., Brown, C. D., Engelhardt, B. E. & Montgomery, S. B. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Grönberg, H. et al. Prostate cancer screening in men aged 50-69 years (STHLM3): a prospective population-based diagnostic study. Lancet Oncol. 16, 1667–1676 (2015).
Eeles, R. A. et al. Multiple newly identified loci associated with prostate cancer susceptibility. Nat. Genet. 40, 316–321 (2008).
Paternoster, L. et al. Multi-ancestry genome-wide association study of 21,000 cases and 95,000 controls identifies new risk loci for atopic dermatitis. Nat. Genet. 47, 1449–1456 (2015).
Dahl, R. et al. Effects of an oral MMP-9 and -12 inhibitor, AZD1236, on biomarkers in moderate/severe COPD: a randomised controlled trial. Pulm. Pharmacol. Ther. 25, 169–177 (2012).
Ganz, P. et al. Development and validation of a protein-based risk score for cardiovascular outcomes among patients with stable coronary heart disease. J. Am. Med. Assoc. 315, 2532–2541 (2016).
Traylor, M. et al. A novel MMP12 locus is associated with large artery atherosclerotic stroke using a genome-wide age-at-onset informed approach. PLoS Genet. 10, e1004469 (2014).
Nelson, M. R. et al. The support of human genetic evidence for approved drug indications. Nat. Genet. 47, 856–860 (2015).
Albagha, O. M. E. et al. Genome-wide association study identifies variants at CSF1, OPTN and TNFRSF11A as genetic risk factors for Paget’s disease of bone. Nat. Genet. 42, 520–524 (2010).
Schwarz, P., Rasmussen, A. Q., Kvist, T. M., Andersen, U. B. & Jørgensen, N. R. Paget’s disease of the bone after treatment with Denosumab: a case report. Bone 50, 1023–1025 (2012).
Moore, C. et al. The INTERVAL trial to determine whether intervals between blood donations can be safely and acceptably decreased to optimise blood supply: study protocol for a randomised controlled trial. Trials 15, 363 (2014).
Gold, L. et al. Aptamer-based multiplexed proteomic technology for biomarker discovery. PLoS ONE 5, e15004 (2010).
Sattlecker, M. et al. Alzheimer’s disease biomarker discovery using SOMAscan multiplexed protein technology. Alzheimers Dement. 10, 724–734 (2014).
McLaren, W. et al. Deriving the consequences of genomic variants with the Ensembl API and SNP Effect Predictor. Bioinformatics 26, 2069–2070 (2010).
UniProt Consortium. UniProt: a hub for protein information. Nucleic Acids Res. 43, D204–D212 (2015).
Ashburner, M. et al.; The Gene Ontology Consortium. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Menni, C. et al. Circulating proteomic signatures of chronological age. J. Gerontol. A Biol. Sci. Med. Sci. 70, 809–816 (2015).
Ngo, D. et al. Aptamer-based proteomic profiling reveals novel candidate biomarkers and pathways in cardiovascular disease. Circulation 134, 270–285 (2016).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012).
Welter, D. et al. The NHGRI GWAS Catalog, a curated resource of SNP-trait associations. Nucleic Acids Res. 42, D1001–D1006 (2014).
Malone, J. et al. Modeling sample variables with an Experimental Factor Ontology. Bioinformatics 26, 1112–1118 (2010).
Javierre, B. M. et al. Lineage-specific genome architecture links enhancers and non-coding disease variants to target gene promoters. Cell 167, 1369–1384. e19 (2016).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLOS Comput. Biol. 9, e1003118 (2013).
Amberger, J. S., Bocchini, C. A., Schiettecatte, F., Scott, A. F. & Hamosh, A. OMIM.org: Online Mendelian Inheritance in Man (OMIM®), an online catalog of human genes and genetic disorders. Nucleic Acids Res. 43, D789–D798 (2015).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457–D462 (2016).
Szklarczyk, D. et al. STRINGv10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–D452 (2015).
Smith, R. N. et al. InterMine: a flexible data warehouse system for the integration and analysis of heterogeneous biological data. Bioinformatics 28, 3163–3165 (2012).
Franceschini, A. et al. STRINGv9.1: protein–protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 41, D808–D815 (2013).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Iotchkova, V. et al. GARFIELD—GWAS Analysis of Regulatory or Functional Information Enrichment with LD correction. Preprint at https://www.biorxiv.org/content/early/2016/11/07/085738 (2016).
Staley, J. R. et al. PhenoScanner: a database of human genotype–phenotype associations. Bioinformatics 32, 3207–3209 (2016).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Hingorani, A. & Humphries, S. Nature’s randomised trials. Lancet 366, 1906–1908 (2005).
Nikpay, M. et al. A comprehensive 1,000 Genomes-based genome-wide association meta-analysis of coronary artery disease. Nat. Genet. 47, 1121–1130 (2015).
Burgess, S., Butterworth, A. & Thompson, S. G. Mendelian randomization analysis with multiple genetic variants using summarized data. Genet. Epidemiol. 37, 658–665 (2013).
Burgess, S., Dudbridge, F. & Thompson, S. G. Combining information on multiple instrumental variables in Mendelian randomization: comparison of allele score and summarized data methods. Stat. Med. 35, 1880–1906 (2016).
Burgess, S. & Thompson, S. G. Multivariable Mendelian randomization: the use of pleiotropic genetic variants to estimate causal effects. Am. J. Epidemiol. 181, 251–260 (2015).
Burgess, S., Dudbridge, F. & Thompson, S. G. Re: “Multivariable Mendelian randomization: the use of pleiotropic genetic variants to estimate causal effects”. Am. J. Epidemiol. 181, 290–291 (2015).
Acknowledgements
A. Day-Williams, J. McElwee, D. Diogo, W. Astle, E. Di Angelantonio, E. Birney, A. Richard, J. Mason and M. Inouye commented on the manuscript, and M. Sharp helped with mapping drug indications to GWAS traits. We thank INTERVAL study participants; staff at recruiting NHSBT blood donation centres; and the INTERVAL Study Co-ordination team, Operations Team (led by R. Houghton and C. Moore) and Data Management Team (led by M. Walker). Funding sources are listed in the Supplementary Information.
Reviewer information
Nature thanks T. Lappalainen, M. McCarthy and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Conceptualization and experimental design: J.D., A.S.B., B.B.S., H.R., R.M.P.; methodology: B.B.S., A.S.B., J.C.M., J.E.P., H.R., S.B.; conducted experimental work: N.J., S.K.W., E.S.Z.; analysis: B.B.S., J.C.M., J.E.P., D.S., J.B., J.R.S., T.J., E.P., P.S., C.O.-W., M.A.K., S.K.W., A.C., N.B., S.L.S.; contributed reagents, materials, protocols or analysis tools: N.J., S.K.W., E.S.Z., J.B., M.A.K., J.R.S., B.P.P.; supervision: A.S.B., H.R., J.D., R.M.P., C.S.F., D.S.P., A.M.W.; writing: A.S.B., J.E.P., B.B.S., J.C.M., H.R., J.D., J.A.T., N.S., K.S.; creation of the INTERVAL BioResource: J.R.B., D.J.R.,W.H.O., N.W.M., J.D.; funding: N.W.M., J.R.B., D.J.R., W.H.O., H.R., R.M.P., J.D.; all authors critically reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare the following competing interests: A.C., CSF-Merck employee; N.J., S.K.W., SomaLogic Inc employees and stakeholders; E.S.Z., SomaLogic Inc employee; J.C.M., R.M.P., Merck employees during this study, now Celgene employees; H.R., Merck employee during this study; J.E.P., travel and accommodation expenses and hospitality from Olink to speak at Olink-sponsored academic meetings; A.S.B., grants from Merck, Pfizer, Novartis, Biogen and Bioverativ and personal fees from Novartis; J.D., sits on the Novartis Cardiovascular and Metabolic Advisory Board, had grant support from Novartis.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
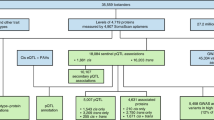
Extended Data Fig. 1
Flowchart of sample processing and quality control stages for proteomic and genetic measurements before genetic analyses.
Extended Data Fig. 2 Examples of protein targets for which the SOMAmer is highly specific.
SDS–PAGE with Alexa-647-labelled proteins captured by the IL1RL2 SOMAmer (a) or GP1BA SOMAmer (b). For each protein target, the protein captured by the SOMAmer is compared to the standard. The cognate targets are the only ones with protein visible in the capture lanes, whereas the proteins homologous to the target proteins show no evidence of binding. These experiments were performed once. MW markers, molecular weight markers.
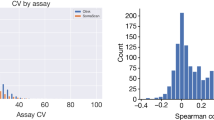
Extended Data Fig. 3 Evidence for the reliability of protein measurements made using the SOMAscan assay.
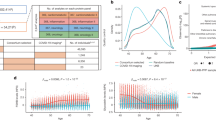
a, Distribution of coefficients of variation of all proteins on the SOMAscan assay in each subcohort. b, Spearman’s correlations for all proteins passing QC derived from contemporaneous assay of baseline and two-year samples from 60 participants. c, Scatterplot of pQTL effect size estimates from SOMAscan versus Olink showing all 163 pQTLs tested (top) and the 106 that replicated (bottom). r is Pearson’s correlation coefficient. d, Distribution of inflation factors across proteins that underwent genome-wide association testing, stratified by subcohort and allele frequency (MAF ≥ 5%, MAF < 5%).
Extended Data Fig. 4 The WFIKKN2 region is a trans pQTL for GDF11/8 plasma levels.
a, Regional association plots of the trans pQTL (sentinel variant rs11079936) for GDF11/8 before and after adjusting for levels of WFIKKN2 (upper panels), and the WFIKKN2 cis pQTL after adjusting for GDF11/8 levels (bottom panel). A similar pattern of association for WFIKKN2 was seen before GDF11/8 adjustment (not shown). b, Attenuation of the GDF11/8 trans pQTL upon adjustment for plasma levels of the cis protein WFIKKN2.
Extended Data Fig. 5 Genetic architecture of the pQTLs.
pQTL mapping in n = 3,301 individuals. a, Distribution of the predicted consequences of the sentinel pQTL variants compared to matched permuted null sets of variants, stratified by cis and trans. Asterisks indicate empirical enrichment using a permutation test (10,000 permuted sets of non-associated variants) at a Bonferroni-corrected significance value (P < 0.005). Bar height represents the mean proportion of variants within each class and error bars reflect one standard deviation from the mean. b, Number of proteins associated (P < 1.5 × 10−11) with each sentinel variant across the genome.
Extended Data Fig. 6 Enrichment of pQTLs at DNase I hypersensitive sites by tissue or cell type.
Circle shows enrichment for DNase I hypersensitive sites (‘hotspots’) for each of 55 tissues (183 cell types) available from the ENCODE and Roadmap Epigenomics projects, with tissues or cell types clustered and coloured by anatomical grouping. Some tissues have multiple values due to availability of multiple cell types or multiple tests per cell type. Radial lines show fold-enrichment, while dots around the inside edge of the circle denote statistically significant enrichment at a Bonferroni-corrected significant threshold P < 5 × 10−5. Enrichment testing performed using GARFIELD (which tests enrichment against permuted sets of variants matched for MAF, distance to TSS and LD). pQTL data from n = 3,301 individuals.
Extended Data Fig. 7 Scheme outlining the combined ‘bottom-up’ and ‘top-down’ process used for candidate gene annotation of trans pQTL regions.
See Methods. GbA, guilt-by-association; KEGG, Kyoto Encyclopedia of Genes and Genomes; OMIM, Online Mendelian Inheritance in Man; STRINGdb, STRING database.
Extended Data Fig. 8 Follow-up of PR3 SOMAmers.
These experiments were repeated three times independently with similar results. a, SOMAmer pulldowns with purified PR3, A1AT, and PR3–A1AT complex. SOMAmer PRTN3.3514.49.2 enriched the PR3–A1AT complex to a much greater degree than free PR3. Conversely, SOMAmer PRTN3.13720.95.3 enriched free PR3 to a greater degree than the PR3–A1AT complex. b, Solution affinity of PRTN3.3514.49.2 and PRTN3.13720.95.3 for PR3, A1AT, and the PR3–A1AT complex. SOMAmer PRTN3.3514.49.2 has a higher affinity for the PR3–A1AT complex than for free PR3. SOMAmer PRTN3.13720.95.3, on the other hand, has a higher affinity for free PR3 than SOMAmer PRTN3.3514.49.2. c, Competitive binding of SOMAmers PRTN3.13720.95.3 and PRTN3.3514.49.2 to PR3. A limiting amount of radiolabelled PRTN3.13720.95.3 was incubated with 1 nM proteinase-3 and a titration of either cold PRTN3.13720.95.3 or cold PRTN3.3514.49.2.
Extended Data Fig. 9
Comparison between a randomized controlled trial and Mendelian randomization to assess the causal effect of changes in protein biomarker levels on disease risk.
Extended Data Fig. 10 Characterization of protein targets measured using the SOMAscan assay.
a, Compartment distribution with annotations of all proteins in the Human Protein Atlas for comparison. b, GO molecular functions.
Supplementary information
Supplementary Information
This file contains funding details, full Supplementary Table Legends, Supplementary Notes and Supplementary References
Supplementary Figure 1
A three-dimensional interactive plot of sentinel variant-protein associations (red-cis, blue-trans). X-axis (“pQTL position”) represents position of the sentinel variant along chromosomes 1-22. Y-axis (“Protein position”) represents the start position of the gene encoding the protein. Z-axis represents the –log10(p) of the association. Additional details can be viewed when hovering over the points. Clicking on cis/trans in the legend toggles display of points by cis/trans. Additional viewing controls are available at the top right of the window. For clarity, associations with p<10-300 (diamonds) are plotted at -log10(p)=300. The plot is generated using “plotly” R package v4.5.6 (Plotly Technologies Inc., Montréal, Canada)
Supplementary Tables
Supplementary Tables 1-21 – see Supplementary Information file for full descriptions
Rights and permissions
About this article
Cite this article
Sun, B.B., Maranville, J.C., Peters, J.E. et al. Genomic atlas of the human plasma proteome. Nature 558, 73–79 (2018). https://doi.org/10.1038/s41586-018-0175-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0175-2
This article is cited by
-
Proteomic associations with forced expiratory volume: a Mendelian randomisation study
Respiratory Research (2024)
-
Clinically relevant plasma proteome for adiposity depots: evidence from systematic mendelian randomization and colocalization analyses
Cardiovascular Diabetology (2024)
-
Associations between genetically predicted plasma protein levels and Alzheimer’s disease risk: a study using genetic prediction models
Alzheimer's Research & Therapy (2024)
-
Investigation of the acute pathogenesis of spondyloarthritis/HLA-B27-associated anterior uveitis based on genome-wide association analysis and single-cell transcriptomics
Journal of Translational Medicine (2024)
-
Peripheral blood indicators and COVID-19: an observational and bidirectional Mendelian randomization study
BMC Medical Genomics (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.