Abstract
A central question in genetics and evolution is the extent to which the outcomes of mutations change depending on the genetic context in which they occur1,2,3. Pairwise interactions between mutations have been systematically mapped within4,5,6,7,8,9,10,11,12,13,14,15,16,17,18 and between19 genes, and have been shown to contribute substantially to phenotypic variation among individuals20. However, the extent to which genetic interactions themselves are stable or dynamic across genotypes is unclear21, 22. Here we quantify more than 45,000 genetic interactions between the same 87 pairs of mutations across more than 500 closely related genotypes of a yeast tRNA. Notably, all pairs of mutations interacted in at least 9% of genetic backgrounds and all pairs switched from interacting positively to interacting negatively in different genotypes (false discovery rate < 0.1). Higher-order interactions are also abundant and dynamic across genotypes. The epistasis in this tRNA means that all individual mutations switch from detrimental to beneficial, even in closely related genotypes. As a consequence, accurate genetic prediction requires mutation effects to be measured across different genetic backgrounds and the use of higher-order epistatic terms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lehner, B. Genotype to phenotype: lessons from model organisms for human genetics. Nat. Rev. Genet. 14, 168–178 (2013).
de Visser, J. A. & Krug, J. Empirical fitness landscapes and the predictability of evolution. Nat. Rev. Genet. 15, 480–490 (2014).
Starr, T. N. & Thornton, J. W. Epistasis in protein evolution. Protein Sci. 25, 1204–1218 (2016).
Fowler, D. M. et al. High-resolution mapping of protein sequence–function relationships. Nat. Methods 7, 741–746 (2010).
Araya, C. L. et al. A fundamental protein property, thermodynamic stability, revealed solely from large-scale measurements of protein function. Proc. Natl Acad. Sci. USA 109, 16858–16863 (2012).
Diss, G, Lehner, B. The genetic landscape of a physical interaction. eLife 7, e32472 (2018).
Melamed, D., Young, D. L., Gamble, C. E., Miller, C. R. & Fields, S. Deep mutational scanning of an RRM domain of the Saccharomyces cerevisiae poly(A)-binding protein. RNA 19, 1537–1551 (2013).
Gong, L. I., Suchard, M. A. & Bloom, J. D. Stability-mediated epistasis constrains the evolution of an influenza protein. eLife 2, e00631 (2013).
Olson, C. A., Wu, N. C. & Sun, R. A comprehensive biophysical description of pairwise epistasis throughout an entire protein domain. Curr. Biol. 24, 2643–2651 (2014).
Gong, L. I. & Bloom, J. D. Epistatically interacting substitutions are enriched during adaptive protein evolution. PLoS Genet. 10, e1004328 (2014).
Bank, C., Hietpas, R. T., Jensen, J. D. & Bolon, D. N. A systematic survey of an intragenic epistatic landscape. Mol. Biol. Evol. 32, 229–238 (2015).
Hayden, E. J., Bendixsen, D. P. & Wagner, A. Intramolecular phenotypic capacitance in a modular RNA molecule. Proc. Natl Acad. Sci. USA 112, 12444–12449 (2015).
Bank, C., Matuszewski, S., Hietpas, R. T. & Jensen, J. D. On the (un)predictability of a large intragenic fitness landscape. Proc. Natl Acad. Sci. USA 113, 14085–14090 (2016).
Puchta, O. et al. Network of epistatic interactions within a yeast snoRNA. Science 352, 840–844 (2016).
Li, C., Qian, W., Maclean, C. J. & Zhang, J. The fitness landscape of a tRNA gene. Science 352, 837–840 (2016).
Julien, P., Miñana, B., Baeza-Centurion, P., Valcárcel, J. & Lehner, B. The complete local genotype–phenotype landscape for the alternative splicing of a human exon. Nat. Commun. 7, 11558 (2016).
Sarkisyan, K. S. et al. Local fitness landscape of the green fluorescent protein. Nature 533, 397–401 (2016).
Guy, M. P. et al. Identification of the determinants of tRNA function and susceptibility to rapid tRNA decay by high-throughput in vivo analysis. Genes Dev. 28, 1721–1732 (2014).
Costanzo, M. et al. A global genetic interaction network maps a wiring diagram of cellular function. Science 353, https://doi.org/10.1126/science.aaf1420 (2016).
Forsberg, S. K., Bloom, J. S., Sadhu, M. J., Kruglyak, L. & Carlborg, Ö. Accounting for genetic interactions improves modeling of individual quantitative trait phenotypes in yeast. Nat. Genet. 49, 497–503 (2017).
Tischler, J., Lehner, B. & Fraser, A. G. Evolutionary plasticity of genetic interaction networks. Nat. Genet. 40, 390–391 (2008).
Weinreich, D. M., Lan, Y., Wylie, C. S. & Heckendorn, R. B. Should evolutionary geneticists worry about higher-order epistasis? Curr. Opin. Genet. Dev. 23, 700–707 (2013).
Palmer, A. C. et al. Delayed commitment to evolutionary fate in antibiotic resistance fitness landscapes. Nat. Commun. 6, 7385 (2015).
Sailer, Z. R. & Harms, M. J. Detecting high-order epistasis in nonlinear genotype-phenotype maps. Genetics 205, 1079–1088 (2017).
Wu, N. C., Dai, L., Olson, C. A., Lloyd-Smith, J. O. & Sun, R. Adaptation in protein fitness landscapes is facilitated by indirect paths. eLife 5, 16965 (2016).
Marcet-Houben, M. & Gabaldón, T. Beyond the whole-genome duplication: phylogenetic evidence for an ancient interspecies hybridization in the baker’s yeast lineage. PLoS Biol. 13, e1002220 (2015).
Hopf, T. A. et al. Mutation effects predicted from sequence co-variation. Nat. Biotechnol. 35, 128–135 (2017).
Ferretti, L. et al. Measuring epistasis in fitness landscapes: The correlation of fitness effects of mutations. J. Theor. Biol. 396, 132–143 (2016).
Weinreich, D. M., Watson, R. A. & Chao, L. Perspective: Sign epistasis and genetic constraint on evolutionary trajectories. Evolution 59, 1165–1174 (2005).
Chan, P. P. & Lowe, T. M. GtRNAdb: a database of transfer RNA genes detected in genomic sequence. Nucleic Acids Res. 37, D93–D97 (2009).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
McWilliam, H. et al. Analysis tool web services from the EMBL-EBI. Nucleic Acids Res. 41, W597–W600 (2013).
Sikorski, R. S. & Hieter, P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122, 19–27 (1989).
Matuszewski, S., Hildebrandt, M. E., Ghenu, A. H., Jensen, J. D. & Bank, C. A statistical guide to the design of deep mutational scanning experiments. Genetics 204, 77–87 (2016).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12, (2011).
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30, 614–620 (2014).
Crawley, M. J. The R Book. (Wiley, Chichester, 2007).
Rubin, A. F. et al. A statistical framework for analyzing deep mutational scanning data. Genome Biol. 18, 150 (2017).
Poelwijk, F. J., Krishna, V. & Ranganathan, R. The context-dependence of mutations: a linkage of formalisms. PLOS Comput. Biol. 12, e1004771 (2016).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Series B Stat. Methodol. 57, 289–300 (1995).
Poelwijk, F. J., Kiviet, D. J., Weinreich, D. M. & Tans, S. J. Empirical fitness landscapes reveal accessible evolutionary paths. Nature 445, 383–386 (2007).
Szendro, I. G., Schenk, M. F., Franke, J., Krug, J. & de Visser, J. A. Quantitative analyses of empirical fitness landscapes. J. Stat. Mech. 2013, P01005 (2013).
Acknowledgements
We thank J. Schmiedel for statistical guidance. This work was supported by a European Research Council Consolidator grant (616434), the Spanish Ministry of Economy and Competitiveness (BFU2011-26206 and SEV-2012-0208), the AXA Research Fund, the Bettencourt Schueller Foundation, Agència de Gestió d’Ajuts Universitaris i de Recerca (AGAUR), the EMBL-CRG Systems Biology Program, and the CERCA Program/Generalitat de Catalunya. Deep sequencing was performed in the EMBL Heidelberg GeneCore Genomics Core Facility.
Reviewer information
Nature thanks Z. Blount, D. Marks, A. Wagner and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
J.D. performed all experiments and analyses. J.D., G.D., and B.L. designed the experiments and analyses. B.L. and J.D. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Experimental design.
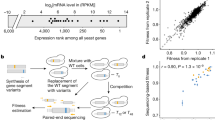
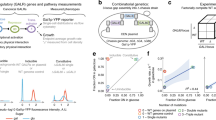
a, Maximum growth rate (measured in a plate reader using spectrophotometry) of tRNA-Arg(CCU) (HSX1) deletion strain carrying either an empty plasmid (red) or a single-copy plasmid expressing wild-type tRNA-Arg(CCU) (blue) at high temperature, high salt, and high temperature with high salt (n = 3 independent colonies from the plasmid transformation). b, Distribution of number of mutations per genotype in the library relative to the sequence of the tRNA from each species. c, Genotype network of the 4,176 tRNA-Arg(CCU) variants. Each node is one genotype. Colour indicates the ln(fitness) relative to S. cerevisiae. Edges connect genotypes differing by a single substitution, acquisition of a U2C mutation is highlighted in yellow as example. Genotypes are arranged in concentric circles according to the total number of substitutions (one to ten) from the S. cerevisiae tRNA, which is the central node. Highlighted nodes indicate the genotypes of the seven extant species. d, Table showing the possible number of mutation combinations from order one to eight, with or without a complete genotype space (whether all intermediate genotypes are measured in the library or not) when using S. cerevisiae as a reference or any other background (the effect of a given combination of mutations can be measured from at least one genetic background). The total number of unique backgrounds is also indicated, together with the minimum, median and maximum number of backgrounds in which these mutations can be found.
Extended Data Fig. 2 Mutations have varying fitness effects in different backgrounds.
a, Single mutations (columns) have effects that differ significantly between genetic backgrounds from different species (rows). Paired two-sided t-test between fitness effects of mutations of tRNAs from different species (145 tests of n = 6). Significant fitness effects differences (FDR < 0.1) shown in blue (positive) or red (negative), non-significant differences (FDR ≥ 0.1) coloured in white. Mutations that were not shared are coloured in grey (that is, a substitution that would result in a mutation in one species but is part of the wild-type background in another). Bar plots show the percentage (absolute numbers on top) of species comparisons or shared mutations between species in which the effect of the mutation significantly changes in magnitude (light grey) or switches sign (dark grey). b, Proportion of genetic backgrounds in which each mutation has a beneficial (blue) or detrimental (red) fitness effect at different FDRs for backgrounds with −0.3 < ln(fitness) < −0.15 (left), backgrounds with −0.15 < ln(fitness) < 0.15 (middle left), genotypes with no more than four mutations from the S. cerevisiae sequence (middle right) and genotypes with average input read counts of more than 100 (right). q values were obtained after adjusting for FDR across the total number of single mutations with unique background after filtering (n = 10,746, 6,129, 3,568, 6,338 tests respectively). c, Fitness effect of single mutations plotted against the ln(fitness) of the backgrounds in which the mutation are made; for all genetic backgrounds (left), backgrounds with −0.3 < ln(fitness) < −0.15 (middle) and backgrounds with −0.15 < ln(fitness) < 0.15 (right).
Extended Data Fig. 3 Comparison of epistasis scores between all pairs of species.
a, Comparison of epistasis scores for species pairs not shown in Fig. 3c. Pairs of species that share less than three mutations are not shown. b, Decline of correlation between epistasis scores and Hamming distance between the tRNA genotypes from different species (inset). The left plot shows how this negative correlation holds when restricting the minimum number of shared pairs of mutations between the two species to compute the correlation.
Extended Data Fig. 4 Changes in pairwise epistasis between mutations across the seven extant species.
a, Comparison of pairwise epistasis (rows) between different species (columns) (1,000 paired two-sided t-tests of n = 6). Differences in epistasis are only shown for comparisons with FDR < 0.1 in orange or green for positive or negative differences respectively. Comparisons with FDR ≥ 0.1 are coloured in white. Pairs of mutations that are not shared between species are coloured in grey. Bar plots show the percentage of species comparisons (right) or shared pairs of mutations between species (top) that significantly change (light grey) or switch (dark grey). b, Interaction networks of four extant species not shown in Fig. 3b. Colours indicate epistasis sign (orange for positive, green for negative and grey for not significant at FDR < 0.1) and edge width indicates epistasis magnitude.
Extended Data Fig. 5 Pairwise epistatic interactions switch from positive to negative.
a, Epistasis scores between pairs of mutations plotted against the ln(fitness) of the genetic background. Scatter plots are divided into double mutants that restore WCBPs (left, n = 1,883), other double mutants in which both mutation are in facing base pair positions (middle left, n = 1,739), in base pair positions but not facing each other (middle right, n = 28,622), and the rest (right, n = 17,144). b, Proportion of genetic backgrounds in which each pair of mutations interacts with positive (orange) or negative (green) epistasis at different FDRs restricted to genetic backgrounds with −0.3 < fitness < −0.15 (top), with −0.15 < fitness <0.15 (top middle), with additive expected fitness outcome greater than−0.2 and less than 0.1 (middle bottom) or when excluding all genotypes with average input counts less than 100 (bottom). 23,128, 23,652, 29,628 and 15,306 one sample two-sided t-tests (n = 6). c, A small fraction of tRNA-Arg(CCU) from other eukaryotic species have lost the base pairing in positions 1–71, 2–70 and 6–66 of the tRNA (multiple sequence alignment (MSA) across 1,614 species was taken from previously published work27; sequences with indels were excluded). d, Number of positive, negative or not significant pairwise interactions at FDR < 0.1 within the acceptor stem of the tRNA (n = 23,237) when both mutations are found in the same helix strand or when each mutation is located in a different strand (n = 13,615). log2 odds ratio shown below together with two-sided Fisher’s exact test P values. e, Number of positive, negative and non-significant background-averaged pairwise interactions between pairs of mutations in the acceptor stem that are found in the same RNA strand and between mutations that are in positions that base pair with each other. log2 odds ratio and two-sided Fisher’s exact test P values are shown below. f, Distribution of pairwise epistasis values of mutation pairs that restore a canonical WCBP depending on the location of their background mutations in the acceptor stem (P values from Welch’s two-sided t-test, n = 263 or n = 1,368 when more than one background mutations are in the same strand or not, respectively). The same result is obtained when epistasis values are corrected for the ln(fitness) of the background (residuals of a linear model using background ln(fitness) to predict epistasis, data not shown).
Extended Data Fig. 6 Changes in base pairing partially explain the consequences on fitness of single mutations.
a, A single mutation can either disrupt or restore a canonical WCBP depending on the background context. b, Percentage of deleterious or beneficial single mutations (at FDR < 0.1) that restore or disturb a canonical WCBP in any base pairing position of the tRNA. From a total of 4,300 mutations that restore WCBP, 721 are beneficial and 498 deleterious. 13,195 mutations result in the loss of a canonical pair (n = 6,806 mutations that create a wobble base pair and n = 6,389 that completely break the base pair interaction), of these 3,030 and 721 have significant deleterious and beneficial effects, respectively. WC, Watson–Crick, W, wobble and L, lost base pair. c, Same as b but split by mutation identity. d, Distribution of the effects of mutations in the tRNA acceptor stem that break a base pairing (left, n = 1,356 single mutations with higher background fitness than −0.15) have more deleterious effects when the neighbour base-pairing positions are composed of one or more wobble interactions (n = 921), instead of all canonical WCBP (n = 435, average fitness effect difference = 0.028, Welch’s two-sided t-test P value shown). Right plot illustrates the context of the base pairing of the stem.
Extended Data Fig. 7 Background-averaged third and higher-order interactions.
a, The most significant background-averaged third-order interactions (8 out of 74, FDR < 0.1, n = 3,691 tests for all interactions across all orders). The first three plots of each row show how the distribution of pairwise epistasis of two mutations across different genetic backgrounds (each double mutation can be found in a median of 506 different genetic backgrounds) changes in the presence or absence of a third mutation. The paired differences between pairwise interactions in those three cases correspond to third order epistatic coefficients. Distributions of third-order epistasis for the same three mutations are shown to the right. Horizontal lines correspond to the background-averaged third-order epistatic term, coloured by sign (orange or green for positive or negative respectively). b, Number of significantly positive and negative background-averaged epistatic interactions of order one to eight (at FDR < 0.1). c, Distribution of the absolute magnitude of averaged third-order interactions plotted against the mean nucleotide distance between the three mutations (n = 316 triple mutations). Welch's two-sided t-test P values for differences between the groups are shown. Significant interactions (one-sample two-sided t-test at FDR < 0.1) are coloured in orange or green for positive or negative epistasis respectively. d, Top, Number of positive, negative or non-significant background-averaged third-order interactions (FDR < 0.1) within the acceptor stem of the tRNA when both mutations are found in the same helix strand or not (n = 129). Bottom, the log2 odds ratios (when all three mutations are found in the same strand of the tRNA acceptor stem) of significantly positive interactions versus others (negative or not significant interactions) and significantly negative interactions versus other double mutants. P values reported from the two-sided Fisher’s exact test.
Extended Data Fig. 8 Genetic prediction.
a, Mean RMSE of the fitness prediction for tenfold cross-validation held-out genotypes (purple, test set) or genotypes included in the training set (yellow) for each of the eight-mutation sub-landscapes when progressively adding the 100 most significant epistatic coefficients out of the 256 possible coefficients. Highlighted in red is the average number of epistatic coefficients to obtain the lowest RMSE across all the sub-landscapes. b, Histogram of the minimum number of epistatic coefficients that give the minimum RMSE when predicting the fitness of the test genotypes by tenfold cross-validation in all complete eight-mutation sub-landscapes (top). Histogram of the median number of coefficients for each sub-landscape (bottom).
Extended Data Fig. 9 Comparison of the combinatorially-complete tRNA sub-landscapes to theoretical fitness landscapes.
a, Expected pattern of the average correlation of fitness effects γd at different mutational distances for theoretical di-allelic fitness landscapes with three to eight mutated positions. The average γd behaviour is highlighted in bold for each theoretical landscape (n = 250 simulated landscapes for each theoretical model). The NK landscape was modelled with K = L/2 (L, number of mutated positions) and the RMF as a mixture of 50% additive and 50% HoC. b, Decay of γd with mutational distance for all tRNA complete di-allelic sub-landscapes containing the S. cerevisiae parental genotype of three to eight loci (mean behaviour of γd in bold). c, Mean euclidean distance between the γd for the tRNA sub-landscapes and the γd of theoretical landscapes (each tRNA landscape was compared to the 250 simulations of each theoretical landscape, n = 73,250, 142,000, 159,500, 100,750, 33,000 and 4,500 for tRNA landscapes from three to eight mutations respectively). d, e, Mean roughness-to-slope ratio (r/s) (d) and epistasis classes (e) for all combinatorially-complete tRNA di-allelic landscapes from three to eight mutations, as well as for all theoretical landscape models (n = 250 for each theoretical landscape models and 293, 568, 638, 403, 132 and 18 tRNA landscapes from three to eight mutations respectively). Error bars are s.d.
Extended Data Fig. 10 Direct paths accessibility between extant species.
Shortest paths between some pairs of extant species (top) together with the proportion of them that are accessible (bottom; yellow, accessible; purple, inaccessible). Nodes are the ln(fitness) of the species genotypes and the intermediate genotypes between them. Edge colours indicate the frequency at which a one-step mutation belongs to an accessible path (completely accessible, yellow; completely inaccessible, purple). Error bars are ln(fitness) s.e.m. of each genotype (propagated error from the n = 6 replicates).
Supplementary information
Supplementary Table 1
Read counts and fitness measurements for all 4,176 genotypes. Standard error reported is the propagated error from the 6 combined replicates.
Rights and permissions
About this article
Cite this article
Domingo, J., Diss, G. & Lehner, B. Pairwise and higher-order genetic interactions during the evolution of a tRNA. Nature 558, 117–121 (2018). https://doi.org/10.1038/s41586-018-0170-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0170-7
This article is cited by
-
Evolvability-enhancing mutations in the fitness landscapes of an RNA and a protein
Nature Communications (2023)
-
ACIDES: on-line monitoring of forward genetic screens for protein engineering
Nature Communications (2023)
-
Pervasive epistasis exposes intramolecular networks in adaptive enzyme evolution
Nature Communications (2023)
-
Experimental exploration of a ribozyme neutral network using evolutionary algorithm and deep learning
Nature Communications (2022)
-
The structure of genotype-phenotype maps makes fitness landscapes navigable
Nature Ecology & Evolution (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.