Abstract
Deep neural networks have achieved impressive successes in fields ranging from object recognition to complex games such as Go1,2. Navigation, however, remains a substantial challenge for artificial agents, with deep neural networks trained by reinforcement learning3,4,5 failing to rival the proficiency of mammalian spatial behaviour, which is underpinned by grid cells in the entorhinal cortex6. Grid cells are thought to provide a multi-scale periodic representation that functions as a metric for coding space7,8 and is critical for integrating self-motion (path integration)6,7,9 and planning direct trajectories to goals (vector-based navigation)7,10,11. Here we set out to leverage the computational functions of grid cells to develop a deep reinforcement learning agent with mammal-like navigational abilities. We first trained a recurrent network to perform path integration, leading to the emergence of representations resembling grid cells, as well as other entorhinal cell types12. We then showed that this representation provided an effective basis for an agent to locate goals in challenging, unfamiliar, and changeable environments—optimizing the primary objective of navigation through deep reinforcement learning. The performance of agents endowed with grid-like representations surpassed that of an expert human and comparison agents, with the metric quantities necessary for vector-based navigation derived from grid-like units within the network. Furthermore, grid-like representations enabled agents to conduct shortcut behaviours reminiscent of those performed by mammals. Our findings show that emergent grid-like representations furnish agents with a Euclidean spatial metric and associated vector operations, providing a foundation for proficient navigation. As such, our results support neuroscientific theories that see grid cells as critical for vector-based navigation7,10,11, demonstrating that the latter can be combined with path-based strategies to support navigation in challenging environments.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Silver, D. et al. Mastering the game of Go with deep neural networks and tree search. Nature 529, 484–489 (2016).
Oh, J., Chockalingam, V., Singh, S. P. & Lee, H. Control of memory, active perception, and action in Minecraft. Proc. Intl Conf. Machine Learning 48 (2016).
Kulkarni, T. D., Saeedi, A., Gautam, S. & Gershman, S. J. Deep successor reinforcement learning. Preprint at https://arxiv.org/abs/1606.02396 (2016).
Mirowski, P. et al. Learning to navigate in complex environments. Intl Conf. Learning Representations (2017).
Hafting, T., Fyhn, M., Molden, S., Moser, M.-B. & Moser, E. I. Microstructure of a spatial map in the entorhinal cortex. Nature 436, 801–806 (2005).
Fiete, I. R., Burak, Y. & Brookings, T. What grid cells convey about rat location. J. Neurosci. 28, 6858–6871 (2008).
Mathis, A., Herz, A. V. & Stemmler, M. Optimal population codes for space: grid cells outperform place cells. Neural Comput. 24, 2280–2317 (2012).
McNaughton, B. L., Battaglia, F. P., Jensen, O., Moser, E. I. & Moser, M.-B. Path integration and the neural basis of the ‘cognitive map’. Nat. Rev. Neurosci. 7, 663–678 (2006).
Erdem, U. M. & Hasselmo, M. A goal-directed spatial navigation model using forward trajectory planning based on grid cells. Eur. J. Neurosci. 35, 916–931 (2012).
Bush, D., Barry, C., Manson, D. & Burgess, N. Using grid cells for navigation. Neuron 87, 507–520 (2015).
Barry, C. & Burgess, N. Neural mechanisms of self-location. Curr. Biol. 24, R330–R339 (2014).
Mittelstaedt, M.-L. & Mittelstaedt, H. Homing by path integration in a mammal. Naturwissenschaften 67, 566–567 (1980).
Bassett, J. P. & Taube, J. S. Neural correlates for angular head velocity in the rat dorsal tegmental nucleus. J. Neurosci. 21, 5740–5751 (2001).
Kropff, E., Carmichael, J. E., Moser, M.-B. & Moser, E. I. Speed cells in the medial entorhinal cortex. Nature 523, 419–424 (2015).
Srivastava, N., Hinton, G. E., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15, 1929–1958 (2014).
Wills, T. J., Cacucci, F., Burgess, N. & O’Keefe, J. Development of the hippocampal cognitive map in preweanling rats. Science 328, 1573–1576 (2010).
Langston, R. F. et al. Development of the spatial representation system in the rat. Science 328, 1576–1580 (2010).
Zhang, S.-J. et al. Optogenetic dissection of entorhinal-hippocampal functional connectivity. Science 340, 1232627 (2013).
Sargolini, F. et al. Conjunctive representation of position, direction, and velocity in entorhinal cortex. Science 312, 758–762 (2006).
Barry, C., Hayman, R., Burgess, N. & Jeffery, K. J. Experience-dependent rescaling of entorhinal grids. Nat. Neurosci. 10, 682–684 (2007).
Stensola, H. et al. The entorhinal grid map is discretized. Nature 492, 72–78 (2012).
Stemmler, M., Mathis, A. & Herz, A. V. Connecting multiple spatial scales to decode the population activity of grid cells. Sci. Adv. 1, e1500816 (2015).
Doeller, C. F., Barry, C. & Burgess, N. Evidence for grid cells in a human memory network. Nature 463, 657–661 (2010).
Kanitscheider, I. & Fiete, I. Training recurrent networks to generate hypotheses about how the brain solves hard navigation problems. Preprint at https://arxiv.org/abs/1609.09059 (2016).
Milford, M. J. & Wyeth, G. F. Mapping a suburb with a single camera using a biologically inspired slam system. IEEE Trans. Robot. 24, 1038–1053 (2008).
Hardcastle, K., Ganguli, S. & Giocomo, L. M. Environmental boundaries as an error correction mechanism for grid cells. Neuron 86, 827–839 (2015).
Chen, G., King, J. A., Burgess, N. & O’Keefe, J. How vision and movement combine in the hippocampal place code. Proc. Natl Acad. Sci. USA 110, 378–383 (2013).
Sarel, A., Finkelstein, A., Las, L. & Ulanovsky, N. Vectorial representation of spatial goals in the hippocampus of bats. Science 355, 176–180 (2017).
Dissanayake, M. G., Newman, P., Clark, S., Durrant-Whyte, H. F. & Csorba, M. A solution to the simultaneous localization and map building (slam) problem. IEEE Trans. Robot. Autom. 17, 229–241 (2001).
Raudies, F. & Hasselmo, M. E. Modeling boundary vector cell firing given optic flow as a cue. PLOS Comput. Biol. 8, e1002553 (2012).
Hochreiter, S. & Schmidhuber, J. Long short-term memory. Neural Comput. 9, 1735–1780 (1997).
Bridle, J. S. in Touretzky, D. S. (ed.) Advances in Neural Information Processing Systems 2 211–217 (Morgan-Kaufmann, 1990).
Elman, J. L. & McClelland, J. L. Exploiting lawful variability in the speech wave. Invariance and Variability in Speech Processes 1, 360–380 (1986).
Tieleman, T. & Hinton, G. Lecture 6.5—RmsProp: Divide the gradient by a running average of its recent magnitude. COURSERA: Neural Networks for Machine Learning (2012).
MacKay, D. J. A practical bayesian framework for backpropagation networks. Neural Comput. 4, 448–472 (1992).
Pascanu, R., Mikolov, T. & Bengio, Y. On the difficulty of training recurrent neural networks. Proc. 30th ICML 28, 1310–1318 (2013).
Ackley, D. H., Hinton, G. E. & Sejnowski, T. J. A learning algorithm for Boltzmann machines. Cogn. Sci. 9, 147–169 (1985).
Beattie, C. et al. Deepmind lab. Preprint at https://arxiv.org/abs/1612.03801 (2016).
Doeller, C. F., Barry, C. & Burgess, N. Evidence for grid cells in a human memory network. Nature 463, 657–661 (2010).
Mnih, V. et al. Asynchronous methods for deep reinforcement learning. In Proc. 33nd Intl Conf. Machine Learning 1928–1937 (2016).
Touretzky, D. S. & Redish, A. D. Theory of rodent navigation based on interacting representations of space. Hippocampus 6, 247–270 (1996).
Foster, D. J., Morris, R. G. & Dayan, P. A model of hippocampally dependent navigation, using the temporal difference learning rule. Hippocampus 10, 1–16 (2000).
Graves, A. et al. Hybrid computing using a neural network with dynamic external memory. Nature 538, 471–476 (2016).
Mnih, V. et al. Human-level control through deep reinforcement learning. Nature 518, 529–533 (2015).
Lin, L.-J. Reinforcement learning for robots using neural networks. Technical Report (Carnegie-Mellon Univ. School of Computer Science, 1993).
Knight, R. et al. Weighted cue integration in the rodent head direction system. Phil. Trans. R. Soc. Lond. B 369, 20120512 (2013).
Solstad, T., Boccara, C. N., Kropff, E., Moser, M.-B. & Moser, E. I. Representation of geometric borders in the entorhinal cortex. Science 322, 1865–1868 (2008).
Barry, C. & Burgess, N. To be a grid cell: Shuffling procedures for determining gridness. Preprint at https://www.biorxiv.org/content/early/2017/12/08/230250 (2017).
Acknowledgements
We thank M. Jaderberg, V. Mnih, A. Santoro, T. Schaul, K. Stachenfeld and J. Yosinski for discussions, and M. Botvinick and J. Wang for comments on an earlier version of the manuscript. C.Ba. funded by Royal Society and Wellcome Trust.
Reviewer information
Nature thanks J. Conradt and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Conceived project: A.B., D.K., C.Ba., R.H., P.M. and B.U.; contributed ideas to experiments: A.B., D.K., C.Ba., B.U., R.H., T.L., C.Bl., P.M., A.P., T.D., J.M., K.K., N.R., G.W., R.G., M.J.C., D.H. and R.P.; performed experiments and analysis: A.B., C.Ba., B.U., M.J.C., T.L., H.S., A.P., B.Z. and F.V.; development of testing platform and environments: C.Be., S.P., R.H., T.L., G.W., D.K., A.B., B.U. and D.H.; human expert tester: A.S.; managed project: D.K., R.H., A.B., H.K., S.G. and D.H.; wrote paper; D.K., A.B., C.Ba., T.L., C.Bl., B.U., M.C., A.P., R.H., N.R., K.K. and D.H.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Network architecture in the supervised learning experiment.
The recurrent layer of the grid cell network is an LSTM with 128 hidden units. The recurrent layer receives as input the vector \(\left[\overrightarrow{v},{\rm{\sin }}\left(\mathop{\varphi }\limits^{^\circ }\right),{\rm{\cos }}\left(\mathop{\varphi }\limits^{^\circ }\right)\right]\). The initial cell state and hidden state of the LSTM, \(\vec{{l}_{0}}\) and \(\vec{{m}_{0}}\), respectively, are initialized by computing a linear transformation of the ground truth place \(\vec{{c}_{0}}\) and head-direction activity \(\vec{{h}_{0}}\) at time 0. The output of the LSTM is followed by a linear layer on which dropout is applied. The output of the linear layer, \(\vec{{g}_{t}}\), is linearly transformed and passed to two softmax functions that calculate the predicted head direction cell activity, \(\vec{{z}_{t}}\), and place cell activity, \(\vec{{y}_{t}}\). We found evidence of grid-like and head direction-like units in the linear layer activations \(\vec{{g}_{t}}\).
Extended Data Fig. 2 Linear layer spatial activity maps from the supervised learning experiment.
Spatial activity plots for all 512 units in the linear layer \(\vec{{g}_{t}}\). Units exhibit spatial activity patterns resembling grid cells, border cells, and place cells. Head direction tuning was also present but is not shown.
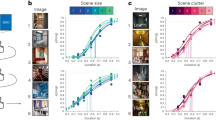
Extended Data Fig. 3 Characterization of grid-like units in square environment and circular environment.
a, The scale (assessed from the spatial autocorrelogram of the ratemaps) of grid-like units exhibited a tendency to cluster at specific values. The number of distinct scale clusters was assessed by sequentially fitting Gaussian mixture models with one to eight components. In each case, the efficiency of the fit (likelihood versus number of parameters) was assessed using Bayesian information criterion (BIC). BIC was minimized with three Gaussian components, indicating the presence of three distinct scale clusters. b, Spatial stability of units in the linear layer of the supervised network was assessed using spatial correlations— bin-wise Pearson product moment correlation between spatial activity maps (32 spatial bins in each map) generated at two different points in training, t = 2 × 105 and t′ = 3 × 105 training steps (two-thirds of the way through training and at the end of training, respectively). This separation was imposed to minimize the effect of temporal correlations and to provide a conservative test of stability. Grid-like units (gridness > 0.37), blue; directionally modulated units (resultant vector length > 0.47, green. Grid-like units exhibit high spatial stability, while directionally modulated units do not. c, Robustness of the grid representation to starting conditions. The network was retrained 100 times with the same hyperparameters but different random seeds controlling the initialization of network weights, \(\overrightarrow{c}\) and \(\overrightarrow{h}\). Populations of grid-like units (gridness > 0.37) were found to appear in all cases, with the average proportion of grid-like units being 23% (s.d. 2.8%). d, The supervised network was also trained in a circular environment (diameter 2.2 m). As before, units in the linear layer exhibited spatially tuned responses resembling grid, border, and head direction cells. Eight units are shown. Top, ratemap displaying activity binned over location. Middle, spatial autocorrelogram of the ratemap; gridness20 is indicated above. Bottom, polar plot of activity binned over head direction. e, Spatial scale of grid-like units (n = 56 (21.9%)) is clustered. Distribution is best fit by a mixture of two Gaussians (centres 0.58 and 0.96 m, ratio 1.66). f, Distribution of directional tuning for 31 most directionally active units; single line for each unit indicates length and orientation of resultant vector47. g, Distribution of gridness and directional tuning. Dashed lines indicate 95% confidence interval derived from shuffling procedure (500 permutations); five grid units (9%) exhibit significant directional modulation.
Extended Data Fig. 4 Grid-like units did not emerge in the linear layer when dropout was not applied.
Linear layer spatial activity maps (n = 512) generated from a supervised network trained without dropout. The maps do not exhibit the regular periodic structure diagnostic of grid cells.
Extended Data Fig. 5 Architecture of the grid cell agent.
The architecture of the supervised network (grid network, light blue dashed) was incorporated into a larger deep reinforcement learning network, including a visual module (green dashed) and an actor–critic learner (based on A3C41; dark blue dashed). In this case the supervised learner does not receive the ground truth \(\vec{{c}_{0}}\) and \(\vec{{h}_{0}}\) to signal its initial position, but uses input from the visual module to self-localize after placement at a random position within the environment. Visual module: since experimental evidence suggests that place cell input to grid cells functions to correct for drift and anchor grids to environmental cues21,27, visual input was processed by a convolutional network to produce place cell (and head direction cell) activity patterns which were used as input to the grid network. The output of the vision module was only provided 5% of the time to the grid network (see Methods for implementational details), akin to occasional observations of salient environmental cues made by behaving animals27. The output of the vision module was concatenated with \(\overrightarrow{u},\overrightarrow{v},\vec{sin\mathop{\varphi }\limits^{^\circ }},\vec{cos\mathop{\varphi }\limits^{^\circ }}\) to form the input to the grid LSTM, which is the same network as in the supervised case (see Methods and Extended Data Fig. 1). The actor–critic learner (light blue dashed) receives as input the concatenation of \(\vec{{e}_{t}^{^{\prime} }}\) produced by a convolutional network with the reward rt, the previous action at−1, the linear layer activations of the grid cell network \(\vec{{g}_{t}}\) (current grid-code), and the linear layer activations observed last time the goal was reached, \(\vec{{g}_{\ast }}\)(goal grid-code), which is set to zero if the goal has not been reached in the episode. The fully connected layer was followed by an LSTM with 256 units. The LSTM has two different outputs. The first output, the actor, is a linear layer with six units followed by a softmax activation function, which represents a categorical distribution over the agent’s next action \(\vec{{\pi }_{t}}\). The second output, the critic, is a single linear unit that estimates the value function vt.
Extended Data Fig. 6 Characterization of grid-like representations and robustness of performance for the grid cell agent in the square land maze environment.
a, Spatial activity plots for the 256 linear layer units in the agent exhibit spatial patterns similar to grid, border, and place cells. b, Cumulative reward indexing goal visits per episode (goal, 10 points) when distal cues are removed (dark blue) and when distal cues are present (light blue). Performance is unaffected, hence dark blue largely obscures light blue. Average of 50% best agent replicas (n = 32) plotted (see Methods). The grey band displays the 68% CI based on 5,000 bootstrapped samples. c, Cumulative reward per episode when no goal code was provided (light blue) and when goal code was provided (dark blue). When no goal code was provided the agent performance fell to that of the baseline deep reinforcement learning agent (A3C) (100 episodes average score no goal code, 123.22 versus A3C, 112.06; effect size, 0.21; 95% CI, 0.18–0.28). Average of 50% best agent replicas (n = 32) plotted (see Methods). The grey band displays the 68% CI based on 5,000 bootstrapped samples. d, After locating the goal for the first time during an episode, the agent typically returned directly to it from each new starting position, showing decreased latencies for subsequent visits, paralleling the behaviour exhibited by rodents.
Extended Data Fig. 7 Robustness of grid cell agent and performance of other agents.
a–c, AUC performance gives robustness to hyperparameters (that is, learning rate, baseline cost, entropy cost; see Supplementary Table 2 in Supplementary Methods for details of the range) and seeds (see Methods). For each environment we run 60 agent replicas (see Methods). Light purple is the grid agent, blue is the place cell agent and dark purple is A3C. a, Square arena. b, Goal-driven. c, Goal doors. In all cases the grid cell agent shows higher robustness to variations in hyperparameters and seeds. d–i, Performance of place cell prediction, NavMemNet and DNC agents (see Methods) against grid cell agent. Dark blue is the grid cell agent (Extended Data Fig. 5), green is the place cell prediction agent (Extended Data Fig. 9a), purple is the DNC agent, light blue is the NavMemNet agent (Extended Data Fig. 9b). The grey band displays the 68% CI based on 5,000 bootstrapped samples. d–f, Performance in goal-driven. g–i, Performance in goal-doors. Note that the performance of the place cell agent (Extended Data Fig. 8b, lower panel) is shown in Fig. 3.
Extended Data Fig. 8 Architecture of the A3C and place cell agent.
a, The A3C implementation is as described41. b, The place cell agent was provided with the ground-truth place, \(\vec{{c}_{t}}\), and head-direction, \(\vec{{h}_{t}}\), cell activations (as described above) at each time step. The output of the fully connected layer of the convolutional network \(\vec{{e}_{t}}\) was concatenated with the reward rt, the previous action at−1, the ground-truth current place code, \(\vec{{c}_{t}}\), and current head-direction code, \(\vec{{h}_{t}}\), together with the ground truth goal place code, \(\vec{{c}_{\ast }}\), and ground truth head direction code, \(\vec{{h}_{\ast }}\), observed the last time the agent reached the goal (see Methods).
Extended Data Fig. 9 Architecture of the place cell prediction agent and of the NavMemNet agent.
a, The architecture of the place cell prediction agent is similar to the grid cell agent, having a grid cell network with the same parameters as that of the grid cell agent. The key difference is the nature of the input provided to the policy LSTM. Instead of using grid codes from the linear layer of the grid network \(\overrightarrow{g}\), we used the predicted place cell population activity vector \(\overrightarrow{y}\), and the predicted head direction population activity vector \(\overrightarrow{z}\), (the activations present on the output place and head direction unit layers of the grid cell network, corresponding to the current and goal position, respectively) as input for the policy LSTM. As in the grid cell agent, the output of the fully connected layer of the convolutional network, \(\overrightarrow{e}\), the reward rt, and the previous action at−1, were also input to the policy LSTM. The convolutional network had the same architecture as described for the grid cell agent. b, NavMemNet agent. The architecture implemented is as described3, specifically FRMQN, but the A3C algorithm was used in place of Q-learning. The convolutional network had the same architecture described for the grid cell agent and the memory was formed of two banks (keys and values), each composed of 1,350 slots.
Extended Data Fig. 10 Flexible use of shortcuts.
a, Overhead view of the linear sunburst maze in initial configuration, with only door 5 open. Example trajectory from grid cell agent during training (green line, icon indicates start location). b, Test configuration with all doors open; grid cell agent uses the newly available shortcuts (multiple episodes shown). c, Histogram showing proportion of times the agent uses each of the doors during 100 test episodes. The agent shows a clear preference for the shortest paths. d, Performance of grid cell agent and comparison agents during test episodes. e, f, Example grid cell agent (e) and example place cell agent (f) trajectory during training in the double E-maze (corridor 1 doors closed). g, h, In the test phase, with all doors open, the grid cell agent exploits the available shortcut (g), while the place cell agent does not (h). i, j, Performance of agents during training (i) and test (j). k, l, The proportion of times the grid (k) and place (l) cell agents used the doors on the first to third corridors during test. The grid cell agent shows a clear preference for available shortcuts, while the place cell agent does not.
Supplementary information
Supplementary Information
This file contains Supplementary Results, Supplementary Discussion, Supplementary Methods and Supplementary Tables 1-2
Rights and permissions
About this article
Cite this article
Banino, A., Barry, C., Uria, B. et al. Vector-based navigation using grid-like representations in artificial agents. Nature 557, 429–433 (2018). https://doi.org/10.1038/s41586-018-0102-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0102-6
This article is cited by
-
The role of strategic visibility in shaping wayfinding behavior in multilevel buildings
Scientific Reports (2024)
-
Spontaneous emergence of rudimentary music detectors in deep neural networks
Nature Communications (2024)
-
Brain Cognition Mechanism-Inspired Hierarchical Navigation Method for Mobile Robots
Journal of Bionic Engineering (2024)
-
Emergent behaviour and neural dynamics in artificial agents tracking odour plumes
Nature Machine Intelligence (2023)
-
Future directions in human mobility science
Nature Computational Science (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.