Abstract
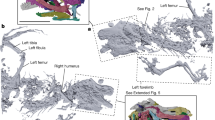
Modern squamates (lizards, snakes and amphisbaenians) are the world’s most diverse group of tetrapods along with birds1 and have a long evolutionary history, with the oldest known fossils dating from the Middle Jurassic period—168 million years ago2,3,4. The evolutionary origin of squamates is contentious because of several issues: (1) a fossil gap of approximately 70 million years exists between the oldest known fossils and their estimated origin5,6,7; (2) limited sampling of squamates in reptile phylogenies; and (3) conflicts between morphological and molecular hypotheses regarding the origin of crown squamates6,8,9. Here we shed light on these problems by using high-resolution microfocus X-ray computed tomography data from the articulated fossil reptile Megachirella wachtleri (Middle Triassic period, Italian Alps10). We also present a phylogenetic dataset, combining fossils and extant taxa, and morphological and molecular data. We analysed this dataset under different optimality criteria to assess diapsid reptile relationships and the origins of squamates. Our results re-shape the diapsid phylogeny and present evidence that M. wachtleri is the oldest known stem squamate. Megachirella is 75 million years older than the previously known oldest squamate fossils, partially filling the fossil gap in the origin of lizards, and indicates a more gradual acquisition of squamatan features in diapsid evolution than previously thought. For the first time, to our knowledge, morphological and molecular data are in agreement regarding early squamate evolution, with geckoes—and not iguanians—as the earliest crown clade squamates. Divergence time estimates using relaxed combined morphological and molecular clocks show that lepidosaurs and most other diapsids originated before the Permian/Triassic extinction event, indicating that the Triassic was a period of radiation, not origin, for several diapsid lineages.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Uetz, P. & Hošek, J. The Reptile Database http://www.reptile-database.org (2017).
Nessov, L. Late Mesozoic amphibians and lizards of Soviet Middle Asia. Acta Zool. Cracov. 31, 475–486 (1988).
Fedorov, P. & Nessov, L. A lizard from the boundary of the Middle and Late Jurassic of north-east Fergana. Bull. St Petersburg Univ. Geol. Geog. 3, 9–14 (1992).
Evans, S. Crown group lizards (Reptilia, Squamata) from the Middle Jurassic of Britain. Palaeontographica. A 250, 123–154 (1998).
Jones, M. E. H. et al. Integration of molecules and new fossils supports a Triassic origin for Lepidosauria (lizards, snakes, and tuatara). BMC Evol. Biol. 13, 208 (2013).
Pyron, R. A. Novel approaches for phylogenetic inference from morphological data and total-evidence dating in squamate reptiles (lizards, snakes, and amphisbaenians). Syst. Biol. 66, 38–56 (2017).
Irisarri, I. et al. Phylotranscriptomic consolidation of the jawed vertebrate timetree. Nat. Ecol. Evol. 1, 1370–1378 (2017).
Simões, T. R., Caldwell, M. W., Nydam, R. L. & Jiménez-Huidobro, P. Osteology, phylogeny, and functional morphology of two Jurassic lizard species and the early evolution of scansoriality in geckoes. Zool. J. Linn. Soc. 180, 216–241 (2017).
Reeder, T. W. et al. Integrated analyses resolve conflicts over squamate reptile phylogeny and reveal unexpected placements for fossil taxa. PLoS ONE 10, e0118199 (2015).
Renesto, S. & Posenato, R. A new lepidosauromorph reptile from the Middle Triassic of the Dolomites (Northern Italy). Riv. Ital. Paleontol. Stratigr. 109, 463–474 (2003).
Renesto, S. & Bernardi, M. Redescription and phylogenetic relationships of Megachirella wachtleri Renesto et Posenato, 2003 (Reptilia, Diapsida). Palaontol. Z. 88, 197–210 (2014).
Carroll, R. L. in Problems in Vertebrate Evolution Vol. 4 (eds Andrews, S. M. et al.) 1–28 (Academic Press, London, 1977).
Mateer, N. Osteology of the Jurassic lizard Ardeosaurus brevipes (Meyer). Palaeontology 25, 461–469 (1982).
Chen, X. H., Motani, R., Cheng, L., Jiang, D.-Y. & Rieppel, O. The enigmatic marine reptile Nanchangosaurus from the lower triassic of Hubei, China and the phylogenetic affinities of Hupehsuchia. PLoS ONE 9, e102361 (2014).
Müller, J. in Recent Advances in the Origin and Early Radiation of Vertebrates (eds Arratia, G. et al.) 379–408 (F. Pfeil, Münich, 2004).
Motani, R., Minoura, N. & Ando, T. Ichthyosaurian relationships illuminated by new primitive skeletons from Japan. Nature 393, 255–257 (1998).
Conrad, J. L. Phylogeny and systematics of Squamata (Reptilia) based on morphology. Bull. Am. Mus. Nat. Hist. 310, 1–182 (2008).
Simões, T. R., Caldwell, M. W., Palci, A. & Nydam, R. L. Giant taxon-character matrices: quality of character constructions remains critical regardless of size. Cladistics 33, 198–219 (2017).
Gauthier, J. A., Kearney, M., Maisano, J. A., Rieppel, O. & Behlke, A. D. B. Assembling the squamate tree of life: perspectives from the phenotype and the fossil record. Bull. Peabody Mus. Nat. Hist. 53, 3–308 (2012).
Pyron, R. A., Burbrink, F. T. & Wiens, J. J. A phylogeny and revised classification of Squamata, including 4161 species of lizards and snakes. BMC Evol. Biol. 13, 93 (2013).
Vidal, N. & Hedges, S. B. The phylogeny of squamate reptiles (lizards, snakes, and amphisbaenians) inferred from nine nuclear protein-coding genes. C. R. Biol. 328, 1000–1008 (2005).
Reynoso, V.-H. Huehuecuetzpalli mixtecus gen. et sp. nov: a basal squamate (Reptilia) from the Early Cretaceous of Tepexi de Rodríguez, Central México. Phil. Trans. R. Soc. Lond. B 353, 477–500 (1998).
Evans, S. E. The skull of a new eosuchian reptile from the Lower Jurassic of South Wales. Zool. J. Linn. Soc. 70, 203–264 (1980).
Zheng, Y. & Wiens, J. J. Combining phylogenomic and supermatrix approaches, and a time-calibrated phylogeny for squamate reptiles (lizards and snakes) based on 52 genes and 4162 species. Mol. Phylogenet. Evol. 94, 537–547 (2016).
Hugall, A. F., Foster, R. & Lee, M. S. Y. Calibration choice, rate smoothing, and the pattern of tetrapod diversification according to the long nuclear gene RAG-1. Syst. Biol. 56, 543–563 (2007).
Jiang, D.-Y. et al. The Early Triassic Eosauropterygian Majiashanosaurus discocoracoidis, gen. et sp. nov. (Reptilia, Sauropterygia), from Chaohu, Anhui Province, People’s Republic of China. J. Vertebr. Paleontol. 34, 1044–1052 (2014).
Butler, R. J. et al. The sail-backed reptile Ctenosauriscus from the latest Early Triassic of Germany and the timing and biogeography of the early archosaur radiation. PLoS ONE 6, e25693 (2011).
Bernardi, M., Klein, H., Petti, F. M. & Ezcurra, M. D. The origin and early radiation of archosauriforms: integrating the skeletal and footprint record. PLoS ONE 10, e0128449 (2015).
Ho, S. Y. & Phillips, M. J. Accounting for calibration uncertainty in phylogenetic estimation of evolutionary divergence times. Syst. Biol. 58, 367–380 (2009).
Chen, Z.-Q. & Benton, M. J. The timing and pattern of biotic recovery following the end-Permian mass extinction. Nat. Geosci. 5, 375–383 (2012).
Tuniz, C. et al. The ICTP-Elettra X-ray laboratory for cultural heritage and archaeology. Nucl. Instrum. Methods Phys. Res. A 711, 106–110 (2013).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Maddison, W. P. & Maddison, D. R. Mesquite: a Modular System for Evolutionary Analysis v.3.04 http://mesquiteproject.org (2015).
Lanfear, R., Frandsen, P. B., Wright, A. M., Senfeld, T. & Calcott, B. PartitionFinder 2: new methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol. Biol. Evol. 34, 772–773 (2017).
Goloboff, P. A., Farris, J. S. & Nixon, K. C. TNT, a free program for phylogenetic analysis. Cladistics 24, 774–786 (2008).
Goloboff, P. A. Analyzing large data sets in reasonable times: solutions for composite optima. Cladistics 15, 415–428 (1999).
Goloboff, P. A., Carpenter, J. M., Arias, J. S. & Esquivel, D. R. M. Weighting against homoplasy improves phylogenetic analysis of morphological data sets. Cladistics 24, 758–773 (2008).
Goloboff, P. A., Torres, A. & Arias, J. S. Weighted parsimony outperforms other methods of phylogenetic inference under models appropriate for morphology. Cladistics https://doi.org/10.1111/cla.12205 (2017).
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012).
Miller, M. A., Pfeiffer, W. & Schwartz, T. in Proceedings of the 2010 Gateway Computing Environments Workshop (GCE) (IEEE, 2010).
Lewis, P. O. A likelihood approach to estimating phylogeny from discrete morphological character data. Syst. Biol. 50, 913–925 (2001).
Harrison, L. B. & Larsson, H. C. Among-character rate variation distributions in phylogenetic analysis of discrete morphological characters. Syst. Biol. 64, 307–324 (2015).
Wagner, P. J. Modelling rate distributions using character compatibility: implications for morphological evolution among fossil invertebrates. Biol. Lett. 8, 143–146 (2012).
Nylander, J. A. A., Ronquist, F., Huelsenbeck, J. P. & Nieves-Aldrey, J. L. Bayesian phylogenetic analysis of combined data. Syst. Biol. 53, 47–67 (2004).
Kass, R. E. & Raftery, A. E. Bayes factors. J. Am. Stat. Assoc. 90, 773–795 (1995).
Xie, W., Lewis, P. O., Fan, Y., Kuo, L. & Chen, M.-H. Improving marginal likelihood estimation for Bayesian phylogenetic model selection. Syst. Biol. 60, 150–160 (2011).
Ronquist, F. et al. A total-evidence approach to dating with fossils, applied to the early radiation of the hymenoptera. Syst. Biol. 61, 973–999 (2012).
Zhang, C., Stadler, T., Klopfstein, S., Heath, T. A. & Ronquist, F. Total-evidence dating under the fossilized birth–death process. Syst. Biol. 65, 228–249 (2016).
Stadler, T. Sampling-through-time in birth–death trees. J. Theor. Biol. 267, 396–404 (2010).
Lepage, T., Bryant, D., Philippe, H. & Lartillot, N. A general comparison of relaxed molecular clock models. Mol. Biol. Evol. 24, 2669–2680 (2007).
Höhna, S., Stadler, T., Ronquist, F. & Britton, T. Inferring speciation and extinction rates under different sampling schemes. Mol. Biol. Evol. 28, 2577–2589 (2011).
Bouckaert, R. et al. BEAST 2: a software platform for Bayesian evolutionary analysis. PLOS Comput. Biol. 10, e1003537 (2014).
O’Reilly, J. E., Dos Reis, M. & Donoghue, P. C. J. Dating tips for divergence-time estimation. Trends Genet. 31, 637–650 (2015).
O’Reilly, J. E. & Donoghue, P. C. Tips and nodes are complementary not competing approaches to the calibration of molecular clocks. Biol. Lett. 12, 20150975 (2016).
Ogg, J. G., Ogg, G. & Gradstein, F. M. A Concise Geologic Time Scale (Elsevier, Amsterdam, 2016).
Benton, M. J. et al. Constraints on the timescale of animal evolutionary history. Palaeontol. Electronica 18, 1–106 (2015).
Gelman, A. & Rubin, D. B. Inference from iterative simulation using multiple sequences. Stat. Sci. 7, 457–472 (1992).
Aberer, A. J., Krompass, D. & Stamatakis, A. Pruning rogue taxa improves phylogenetic accuracy: an efficient algorithm and webservice. Syst. Biol. 62, 162–166 (2013).
Wilkinson, M. Identifying stable reference taxa for phylogenetic nomenclature. Zool. Scr. 35, 109–112 (2006).
Acknowledgements
We are grateful for funding from the Vanier Canada and the Izaak Walton Killam Memorial PhD scholarships to T.R.S.; Euregio Science Fund (call 2014, IPN16) to M.B.; Midwestern University Intramural Funds to R.L.N.; Natural Science and Engineering Research Council of Canada Discovery Grant to M.W.C. (23458); Alberta Ukrainian Centennial Scholarship to O.V.; and National Science Centre grant 2014/13/N/NZ8/02467 to M.T.; and E. Kustatscher for access to the holotype of M. wachtleri.
Reviewer information
Nature thanks M. Baron, J.-C. Rage and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
T.R.S. conducted phylogenetic data collection and analyses; M.W.C. conceived the project; F.B., L.M., M.B., T.R.S. and A.P. conducted micro-CT scans and computed tomography segmentations; T.R.S. and O.V. performed molecular sequence alignment; T.R.S., M.B., M.T., A.P. and R.L.N. performed morphological description; all authors contributed to writing and discussions.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Cranial anatomy of M. wachtleri (PZO 628) based on personal examination and micro-CT scan data.
a, Skull in dorsal view. b, Skull in posteroventral view. c, Skull in anteroventral view. d, Skull in right ventrolateral view. e, Skull in left dorsal lateral view. f, Line drawing of the skull in dorsal view. g, Reconstruction of the skull in dorsal view. h, Detailed view of right lateral side of the skull. i, Drawing of the view in h. San, surangular. Scale bars, 5 mm (a–g).
Extended Data Fig. 2 Cranial and postcranial anatomy of M. wachtleri (PZO 628) based on personal examination and micro-CT scan data.
a, Cross-section of the skull at the level of the frontals in anterior view. b, Details of the anterior end of the left dentary in occlusal view. c, Left quadrate. d, Whole body of the holotype as preserved in the slab (dorsal view). e, Anterior cervical vertebrae in left lateral view. f, Longitudinal section of the anterior cervicals in ventral view. g, Last cervicals and anterior dorsals in dorsal view. h, Pectoral girdle in ventral view. i, Pectoral girdle in left ventrolateral view. j, Right humerus in ventral view. k, Right manus in dorsal view. l, Line drawing of right manus in dorsal view. Ax.R., axis rib; Ce.Pl., cervical, pleurocentrum; Co, cotyle; C.V.3, third cervical vertebra; dc2–5, distal carpals 2–5; DPC, deltopectoral crest; D.R., dorsal rib; D.T., dentary teeth; Epi.St., epiphysial suture; H.Epi., humeral epiphysis; i, intermedium; lc, lateral centrale; McI–V, metacarpals I–V; N.A., neural arch; Olf.Tr., olfactory tract; Po.Co., posterior cotyle; Qj.Fr., quadratojugal foramen; Qj.St., quadratojugal suture; r, radiale; Sbd.Sh., subdentary shelf; Sof.Pr., subolfactory processes; u, ulnare. Scale bars, 1 mm (a, b), 5 mm (c, e–h, j–l), 10 mm (d, i).
Extended Data Fig. 3 Equal weights maximum parsimony analysis, morphological data only.
Strict consensus of 621 most parsimonious trees (2,268 steps each). Numbers at nodes indicate Bremer indices.
Extended Data Fig. 4 Implied weighting maximum parsimony analysis, morphological data only.
Strict consensus of the five best feet trees (fit = 91.768892).
Extended Data Fig. 5 Bayesian inference analysis, morphological data only.
Bayesian majority-rule consensus tree. Numbers at nodes indicate posterior probabilities.
Extended Data Fig. 6 Bayesian inference analysis, combined morphological and molecular data.
Bayesian majority-rule consensus tree. Numbers at nodes indicate posterior probabilities.
Extended Data Fig. 7 Relaxed-clock Bayesian inference analysis with total-evidence tip dating using the fossilized birth–death tree model, combined morphological and molecular data.
Bayesian majority-rule consensus tree. Numbers at nodes indicate posterior probabilities.
Extended Data Fig. 8 Relaxed-clock Bayesian inference analysis with total-evidence tip and node dating using the fossilized birth–death tree model, combined morphological and molecular data.
Bayesian majority-rule consensus tree. Numbers at nodes indicate median estimates for the divergence times, and node bars indicate the 95% highest posterior density for divergence times.
Extended Data Fig. 9 Taxon stability plotted against taxon completeness in the analysis combining both morphological and molecular data.
a, Taxon stability in uncalibrated Bayesian inference analysis. b, Taxon stability in relaxed-clock Bayesian inference analysis with tip dating. Taxon stability increases directly proportional to taxon completeness. M. wachtleri (taxon 67, in red) has a stability slightly above average for uncalibrated Bayesian inference, and well above average for Bayesian inference with tip dating. All taxa are identified in Supplementary Table 3 (n = 129 taxa). Regression line in blue and 95% confidence interval in grey. Labels for extant taxa (~100% completeness) are omitted for simplicity.
Supplementary information
Supplementary Information
This file contains Supplementary Discussions on the morphology of M. wachtleri, Supplementary Methods including taxon sampling and character list and Supplementary Tables 1-3
Supplementary Data 1
Morphological phylogenetic dataset
Supplementary Data 2
Molecular phylogenetic dataset
Supplementary Data 3
Combined phylogenetic dataset with Mr. Bayes commands used for uncalibrated Bayesian analysis.
Supplementary Data 4
Combined phylogenetic dataset with Mr. Bayes commands used for relaxed clock Bayesian analysis calibrated with tip dating.
Supplementary Data 5
Combined phylogenetic dataset with Mr. Bayes commands used for relaxed clock Bayesian analysis calibrated with tip and node dating.
Rights and permissions
About this article
Cite this article
Simões, T.R., Caldwell, M.W., Tałanda, M. et al. The origin of squamates revealed by a Middle Triassic lizard from the Italian Alps. Nature 557, 706–709 (2018). https://doi.org/10.1038/s41586-018-0093-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0093-3
This article is cited by
-
A late-surviving phytosaur from the northern Atlantic rift reveals climate constraints on Triassic reptile biogeography
BMC Ecology and Evolution (2023)
-
Extended embryo retention and viviparity in the first amniotes
Nature Ecology & Evolution (2023)
-
Lepidosauromorphs and associated vertebrate fauna from the Late Triassic Tiki Formation, South Rewa, Gondwana basin, India: implication for paleoenvironment and paleobiogeography
Proceedings of the Indian National Science Academy (2023)
-
High morphological disparity in a bizarre Paleocene fauna of predatory freshwater reptiles
BMC Ecology and Evolution (2022)
-
Evolutionary origins of the prolonged extant squamate radiation
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.