Abstract
Transdifferentiation is a complete and stable change in cell identity that serves as an alternative to stem-cell-mediated organ regeneration. In adult mammals, findings of transdifferentiation have been limited to the replenishment of cells lost from preexisting structures, in the presence of a fully developed scaffold and niche1. Here we show that transdifferentiation of hepatocytes in the mouse liver can build a structure that failed to form in development—the biliary system in a mouse model that mimics the hepatic phenotype of human Alagille syndrome (ALGS)2. In these mice, hepatocytes convert into mature cholangiocytes and form bile ducts that are effective in draining bile and persist after the cholestatic liver injury is reversed, consistent with transdifferentiation. These findings redefine hepatocyte plasticity, which appeared to be limited to metaplasia, that is, incomplete and transient biliary differentiation as an adaptation to cell injury, based on previous studies in mice with a fully developed biliary system3,4,5,6. In contrast to bile duct development7,8,9, we show that de novo bile duct formation by hepatocyte transdifferentiation is independent of NOTCH signalling. We identify TGFβ signalling as the driver of this compensatory mechanism and show that it is active in some patients with ALGS. Furthermore, we show that TGFβ signalling can be targeted to enhance the formation of the biliary system from hepatocytes, and that the transdifferentiation-inducing signals and remodelling capacity of the bile-duct-deficient liver can be harnessed with transplanted hepatocytes. Our results define the regenerative potential of mammalian transdifferentiation and reveal opportunities for the treatment of ALGS and other cholestatic liver diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Merrell, A. J. & Stanger, B. Z. Adult cell plasticity in vivo: de-differentiation and transdifferentiation are back in style. Nat. Rev. Mol. Cell Biol. 17, 413–425 (2016).
Vanderpool, C. et al. Genetic interactions between hepatocyte nuclear factor-6 and Notch signaling regulate mouse intrahepatic bile duct development in vivo. Hepatology 55, 233–243 (2012).
Tarlow, B. D. et al. Bipotential adult liver progenitors are derived from chronically injured mature hepatocytes. Cell Stem Cell 15, 605–618 (2014).
Sekiya, S. & Suzuki, A. Hepatocytes, rather than cholangiocytes, can be the major source of primitive ductules in the chronically injured mouse liver. Am. J. Pathol. 184, 1468–1478 (2014).
Kamimoto, K. et al. Heterogeneity and stochastic growth regulation of biliary epithelial cells dictate dynamic epithelial tissue remodeling. eLife 5, e15034 (2016).
Tanimizu, N. et al. Progressive induction of hepatocyte progenitor cells in chronically injured liver. Sci. Rep. 7, 39990 (2017).
Zong, Y. et al. Notch signaling controls liver development by regulating biliary differentiation. Development 136, 1727–1739 (2009).
Sparks, E. E., Huppert, K. A., Brown, M. A., Washington, M. K. & Huppert, S. S. Notch signaling regulates formation of the three-dimensional architecture of intrahepatic bile ducts in mice. Hepatology 51, 1391–1400 (2010).
Jeliazkova, P. et al. Canonical Notch2 signaling determines biliary cell fates of embryonic hepatoblasts and adult hepatocytes independent of Hes1. Hepatology 57, 2469–2479 (2013).
Thorel, F. et al. Conversion of adult pancreatic α-cells to β-cells after extreme β-cell loss. Nature 464, 1149–1154 (2010).
Stange, D. E. et al. Differentiated Troy+ chief cells act as reserve stem cells to generate all lineages of the stomach epithelium. Cell 155, 357–368 (2013).
Jain, R. et al. Plasticity of Hopx+ type I alveolar cells to regenerate type II cells in the lung. Nat. Commun. 6, 6727 (2015).
Tetteh, P. W. et al. Replacement of lost Lgr5-positive stem cells through plasticity of their enterocyte-lineage daughters. Cell Stem Cell 18, 203–213 (2016).
Van Eyken, P., Sciot, R. & Desmet, V. J. A cytokeratin immunohistochemical study of cholestatic liver disease: evidence that hepatocytes can express ‘bile duct-type’ cytokeratins. Histopathology 15, 125–135 (1989).
Libbrecht, L. et al. Peripheral bile duct paucity and cholestasis in the liver of a patient with Alagille syndrome: further evidence supporting a lack of postnatal bile duct branching and elongation. Am. J. Surg. Pathol. 29, 820–826 (2005).
Ernst, L. M., Spinner, N. B., Piccoli, D. A., Mauger, J. & Russo, P. Interlobular bile duct loss in pediatric cholestatic disease is associated with aberrant cytokeratin 7 expression by hepatocytes. Pediatr. Dev. Pathol. 10, 383–390 (2007).
Fabris, L. et al. Analysis of liver repair mechanisms in Alagille syndrome and biliary atresia reveals a role for notch signaling. Am. J. Pathol. 171, 641–653 (2007).
Michalopoulos, G. K., Barua, L. & Bowen, W. C. Transdifferentiation of rat hepatocytes into biliary cells after bile duct ligation and toxic biliary injury. Hepatology 41, 535–544 (2005).
Yanger, K. et al. Robust cellular reprogramming occurs spontaneously during liver regeneration. Genes Dev. 27, 719–724 (2013).
Tanimizu, N., Nishikawa, Y., Ichinohe, N., Akiyama, H. & Mitaka, T. Sry HMG box protein 9-positive (Sox9+) epithelial cell adhesion molecule-negative (EpCAM-) biphenotypic cells derived from hepatocytes are involved in mouse liver regeneration. J. Biol. Chem. 289, 7589–7598 (2014).
Nagahama, Y. et al. Contributions of hepatocytes and bile ductular cells in ductular reactions and remodeling of the biliary system after chronic liver injury. Am. J. Pathol. 184, 3001–3012 (2014).
Li, L. et al. Alagille syndrome is caused by mutations in human Jagged1, which encodes a ligand for Notch1. Nat. Genet. 16, 243–251 (1997).
Oda, T. et al. Mutations in the human Jagged1 gene are responsible for Alagille syndrome. Nat. Genet. 16, 235–242 (1997).
McDaniell, R. et al. NOTCH2 mutations cause Alagille syndrome, a heterogeneous disorder of the notch signaling pathway. Am. J. Hum. Genet. 79, 169–173 (2006).
Poncy, A. et al. Transcription factors SOX4 and SOX9 cooperatively control development of bile ducts. Dev. Biol. 404, 136–148 (2015).
Walter, T. J., Vanderpool, C., Cast, A. E. & Huppert, S. S. Intrahepatic bile duct regeneration in mice does not require Hnf6 or Notch signaling through Rbpj. Am. J. Pathol. 184, 1479–1488 (2014).
Gong, A. Y. et al. Somatostatin stimulates ductal bile absorption and inhibits ductal bile secretion in mice via SSTR2 on cholangiocytes. Am. J. Physiol. Cell Physiol. 284, C1205–C1214 (2003).
Li, B. et al. Adult mouse liver contains two distinct populations of cholangiocytes. Stem Cell Reports 9, 478–489 (2017).
Peng, L. et al. RNA sequencing reveals dynamic changes of mRNA abundance of cytochromes P450 and their alternative transcripts during mouse liver development. Drug Metab. Dispos. 40, 1198–1209 (2012).
Rezvani, M. et al. In vivo hepatic reprogramming of myofibroblasts with AAV vectors as a therapeutic strategy for liver fibrosis. Cell Stem Cell 18, 809–816 (2016).
Fiorotto, R. et al. Notch signaling regulates tubular morphogenesis during repair from biliary damage in mice. J. Hepatol. 59, 124–130 (2013).
Clotman, F. et al. Control of liver cell fate decision by a gradient of TGFβ signaling modulated by Onecut transcription factors. Genes Dev. 19, 1849–1854 (2005).
Clotman, F. et al. The onecut transcription factor HNF6 is required for normal development of the biliary tract. Development 129, 1819–1828 (2002).
Plumb-Rudewiez, N. et al. Transcription factor HNF-6/OC-1 inhibits the stimulation of the HNF-3α/Foxa1 gene by TGF-β in mouse liver. Hepatology 40, 1266–1274 (2004).
Tanimizu, N. et al. Intrahepatic bile ducts are developed through formation of homogeneous continuous luminal network and its dynamic rearrangement in mice. Hepatology 64, 175–188 (2016).
Rougemont, A. L. et al. Bile ducts in regenerative liver nodules of Alagille patients are not the result of genetic mosaicism. J. Pediatr. Gastroenterol. Nutr. 61, 91–93 (2015).
Raven, A. et al. Cholangiocytes act as facultative liver stem cells during impaired hepatocyte regeneration. Nature 547, 350–354 (2017).
Mu, X. et al. Epithelial transforming growth factor-β signaling does not contribute to liver fibrosis but protects mice from cholangiocarcinoma. Gastroenterology 150, 720–733 (2016).
Luche, H., Weber, O., Nageswara Rao, T., Blum, C. & Fehling, H. J. Faithful activation of an extra-bright red fluorescent protein in “knock-in” Cre-reporter mice ideally suited for lineage tracing studies. Eur. J. Immunol. 37, 43–53 (2007).
Yamamoto, M. et al. A multifunctional reporter mouse line for Cre- and FLP-dependent lineage analysis. Genesis 47, 107–114 (2009).
Lakso, M. et al. Efficient in vivo manipulation of mouse genomic sequences at the zygote stage. Proc. Natl Acad. Sci. USA 93, 5860–5865 (1996).
Snippert, H. J. et al. Intestinal crypt homeostasis results from neutral competition between symmetrically dividing Lgr5 stem cells. Cell 143, 134–144 (2010).
Levéen, P. et al. Induced disruption of the transforming growth factor beta type II receptor gene in mice causes a lethal inflammatory disorder that is transplantable. Blood 100, 560–568 (2002).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Shinkai, Y. et al. RAG-2-deficient mice lack mature lymphocytes owing to inability to initiate V(D)J rearrangement. Cell 68, 855–867 (1992).
Cao, X. et al. Defective lymphoid development in mice lacking expression of the common cytokine receptor γ chain. Immunity 2, 223–238 (1995).
Malato, Y. et al. Fate tracing of mature hepatocytes in mouse liver homeostasis and regeneration. J. Clin. Invest. 121, 4850–4860 (2011).
Raymond, C. S. & Soriano, P. High-efficiency FLP and PhiC31 site-specific recombination in mammalian cells. PLoS One 2, e162 (2007).
Wieser, R., Wrana, J. L. & Massagué, J. GS domain mutations that constitutively activate TβR-I, the downstream signaling component in the TGF-β receptor complex. EMBO J. 14, 2199–2208 (1995).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Dorrell, C. et al. Prospective isolation of a bipotential clonogenic liver progenitor cell in adult mice. Genes Dev. 25, 1193–1203 (2011).
Chen, Z., Wang, M., Xiang, Q., Sun, Z. & Zhang, R. The lectin of Dolichos biflorus agglutinin recognises glycan epitopes on the surface of a subset of cardiac progenitor cells. Cell Biol. Int. 37, 1238–1245 (2013).
Orellana, D. et al. Calmodulin controls liver proliferation via interactions with C/EBPβ-LAP and C/EBPβ-LIP. J. Biol. Chem. 285, 23444–23456 (2010).
Siggs, O. M., Schnabl, B., Webb, B. & Beutler, B. X-linked cholestasis in mouse due to mutations of the P4-ATPase ATP11C. Proc. Natl Acad. Sci. USA 108, 7890–7895 (2011).
Acknowledgements
The authors received the following support: H.W.: NIH R01 DK107553, CIRM DISC1-08792, NIH P30 DK026743. S.S.H.: NIH R01 DK078640, NIH R01 DK107553, NIH P30 DK078392. J.R.S.: NIH T32 DK060414, A. P. Giannini Foundation. S.N.T.K. and B.Y.H.: NIH T32 GM008568. M.R.: Deutsche Forschungsgemeinschaft RE 3749/1-1. F.C.: NIH T32 DK060414, Jane Coffin Childs Memorial Fund. H.Y.L.: Eli and Edythe Broad Regeneration Medicine and Stem Cell Fellowship. The authors thank Donghui Wang (UCSF Preclinical Therapeutics Core), Chris Her (UCSF Liver Center Cell Biology Core), Vinh Nguyen (UCSF Flow Cytometry Core) and Matt Kofron and Mike Muntifering (CCHMC Nikon Center of Excellence Confocal Imaging Core) for technical support, Nick Timchenko, Maria Shrower and Abigail Roebker (CCHMC) for reagents and technical assistance, Rik Derynck (UCSF) for advice and Pamela Derish (UCSF) for manuscript editing.
Reviewer information
Nature thanks S. Forbes, W. Goessling and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
H.W. and S.S.H. conceived the study. The authors contributed to designing and executing the experiments and data analysis as follows: J.R.S.: Figs. 1–4, Extended Data Figs. 1–6. K.A.H.: Figs. 1, 3, 4, Extended Data Figs. 3, 5, 6. S.N.T.K.: Figs. 1–4, Extended Data Fig. 1, 3–6, Extended Data Table 1. B.Y.H.: Fig. 4. A.E.C.: Extended Data Fig. 6. B.D.: Extended Data Fig. 6. R.A.K.: Fig. 2, Extended Data Fig. 4. F.C.: Extended Data Fig. 2. M.R.: Fig. 4. H.Y.L.: Fig. 3, Extended Data Fig. 3. A.N.M.: Fig. 4, Extended Data Table 1. A.-L.R.: Fig. 4, Extended Data Table 1. P.R.: Fig. 4, Extended Data Table 1. S.S.H.: Figs. 1–4, Extended Data Figs. 3, 5. H.W.: Fig. 2, Extended Data Fig. 4, Extended Data Table 1. H.W., S.S.H., J.R.S. and S.N.T.K. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Flp-based hepatocyte fate tracing.
a, R26ZG allele. b, Experimental design for establishing efficient, specific and constitutive labelling of hepatocytes in normal adult R26ZG+/+ mice. c–f, Immunofluorescence of R26ZG+/+ mouse liver (n = 2) for GFP and the hepatocyte marker major urinary protein (MUP) (c), peripheral and hilar cholangiocyte marker CK19 (d, e), hilar cholangiocyte marker DBA (e), hepatic stellate cell marker desmin (DES), macrophage marker F4/80 and endothelial cell marker LYVE1 (f) 2 weeks after intravenous injection of 1 × 1012 viral genomes (vg) of AAV8-Ttr-Flp. g, Reporter activation in R26ZG+/+ mice 1 and 2 weeks after intravenous injection of the indicated dose of AAV8-Ttr-Flp (n = 1 for each dose and time point). Scale bars, 100 µm.
Extended Data Fig. 2 Efficiency of hepatocyte fate tracing in mice born with or without pBDs.
a, b, Experimental design for hepatocyte fate tracing at P17 and immunofluorescence at P120 in Rbpjf/fHnf6f/fR26ZG+/+ mice (n = 4). c, Correlation of GFP labelling efficiency between hepatocytes and peripheral cholangiocytes in hepatocyte-fate-traced P120 Alb-cre+/−Rbpjf/fHnf6f/fR26ZG+/+ mice (n = 5; top and bottom). Horizontal line denotes the mean value. d, e, Experimental design for hepatocyte fate tracing at P39 and immunofluorescence at P120 in Alb-cre+/−Rbpjf/fHnf6f/fR26ZG+/+ mice (n = 3). Scale bars, 100 µm.
Extended Data Fig. 3 HpBDs relieve cholestasis and liver injury.
a, Serum total bilirubin levels in P20–P29 (n = 6), P30–P39 (n = 33), P40–P49 (n = 35), P50–P59 (n = 8), P60–P69 (n = 46), P70–P79 (n = 22), P80–P89 (n = 13), P90–P119 (n = 20), P120–P149 (n = 52) and ≥P150 (n = 40) Alb-cre+/−Rbpjf/fHnf6f/f and P20–P29 (n = 5), P30–P39 (n = 24), P40–P49 (n = 19), P50–P59 (n = 7), P60–P69 (n = 27), P70–P79 (n = 10), P80–P89 (n = 11), P90–P119 (n = 10), P120–P149 (n = 41) and ≥P150 (n = 25) Rbpjf/fHnf6f/f mice. **P = 0.0011 at P20–P29, ****P = 1.7 × 10−13 at P30–P39, ****P = 8.2 × 10−12 at P40–P49, ***P = 0.00019 at P50–P59, ****P = 8.4 × 10−10 at P60–P69, ***P = 0.00055 at P70–P79, not significant (ns; P = 0.090) at P80–P89, not significant (P = 0.050) at P90–P119, not significant (P = 0.052) at P120–P149 and not significant (P = 0.064) at ≥P150, two-sided Welch’s t-test. b, Serial measurements of serum total bilirubin levels in Alb-cre+/−Rbpjf/fHnf6f/f (n = 14) and Rbpjf/fHnf6f/f (n = 5) mice. P = 0.11, two-sided Welch’s t-test. c–e, Serum alkaline phosphatase (ALP), alanine aminotransferase (ALT) and aspartate aminotransferase (AST) levels in P43–P45 (n = 13), P69–P82 (n = 13), P120 (n = 14) and P150 (n = 11) Alb-cre+/−Rbpjf/fHnf6f/f and P43–P45 (n = 5), P69–P82 (n = 4), P120 (n = 9) and P150 (n = 6) Rbpjf/fHnf6f/f mice. In c: ****P = 0.000058 at P43–P45, ****P = 0.000019 at P69–P82, **P = 0.0064 at P120, not significant (P = 0.22) at P150 and ***P = 0.00010 at P120 versus P69–P82, two-way ANOVA followed by Holm–Sidak multiple comparison test. In d: **P = 0.0050 at P43–P45, not significant (P = 0.47) at P69–P82, not significant (P = 0.30) at P120 and not significant (P = 0.47) at P150, two-way ANOVA followed by Holm–Sidak multiple comparison test. In e: **P = 0.0024 at P43–P45, *P = 0.015 at P69–P82, ****P = 0.000073 at P120 and not significant (P = 0.061) at P150, two-way ANOVA followed by Holm–Sidak multiple comparison test. f, Sirius red staining in P15 (n = 9), P70–P90 (n = 7) and P120 (n = 6) Alb-cre+/−Rbpjf/fHnf6f/f and P15 (n = 5), P70–P90 (n = 3) and P120 (n = 3) Rbpjf/fHnf6f/f mice with quantification. Not significant (P = 0.94) at P15, ****P = 0.000095 at P70–P90, **P = 0.0074 at P120 and *P = 0.027 at P120 versus P70–90, two-way ANOVA followed by Holm–Sidak multiple comparison test. g, Immunohistochemistry and Sirius red staining in P313 Alb-cre+/−Rbpjf/fHnf6f/f mice with persistent or resolved cholestasis (n = 1 each). Horizontal lines in a–f denote mean values. Scale bars, 100 µm.
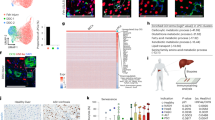
Extended Data Fig. 4 Isolation and gene expression profiling of hepatocyte-derived peripheral cholangiocytes.
a, FACS gates for peripheral cholangiocyte (pC; EPCAM+DBA–) and hilar cholangiocyte (hC; EPCAM+DBA+) isolation from Alb-cre+/−Rbpjf/fHnf6f/f and Rbpjf/fHnf6f/f mice. b, qPCR analysis of floxed Rbpj (Rbpjf/f) genomic DNA in hepatocyte-derived pC (HpC) and hC isolated from Alb-cre+/−Rbpjf/fHnf6f/f mice relative to hepatocytes isolated from Rbpjf/fHnf6f/f mice (dashed line; n = 3 each). Data were normalized to a downstream genomic region of Rbpj to control for gene copy number. Data are mean ± s.e.m. c, d, RNA-seq analysis of normal pC (n = 3 mice), HpC (n = 4 mice) and RBPJ- and HNF6-deficient hepatocytes (H; n = 3 mice). c, Heat map of genes reflecting deletion of Rbpj and Hnf6 (also known as Onecut1). Rbpj mRNA is present in this knockout mouse as a truncated transcript that does not produce a functional protein26. d, Heat map of all differentially expressed CYP genes distinguishing genes associated with mature (M), adolescent (A) and immature (I) hepatocyte differentiation or low expression in the liver (L)29. P < 0.05, one-way ANOVA, FDR-corrected; fold change > 3 (c, d, except Rbpj and Notch1–Notch4). Bold genes denote P < 0.05 for HpC versus pC, two-sided Student’s t-test (c, d).
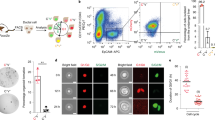
Extended Data Fig. 5 Proliferation in HpBDs and reactive ductules.
a, Size distribution of wsCK-positive DBA-positive hilar cholangiocyte clones in P90 Alb-cre+/−Rbpjf/fHnf6f/fR26R-Confetti+/− (n = 2) and Alb-cre+/−R26R-Confetti+/− (n = 2) mouse livers. **P = 0.0079 for three cells and ***P = 0.00092 for seven cells, two-sided Student’s t-test. b, Immunofluorescence of reactive ductules in hepatocyte-fate-traced P32 Alb-cre+/−Rbpjf/fHnf6f/fR26ZG+/+ mouse liver (n = 3). c, Size distribution of wsCK-positive DBA-negative peripheral cholangiocyte clones in P90 Alb-cre+/−Rbpjf/fHnf6f/fR26R-Confetti+/− (n = 2) and Alb-cre+/−R26R-Confetti+/− (n = 2) mouse livers. *P = 0.032 for one cell, *P = 0.024 for two cells, *P = 0.020 for three cells and *P = 0.014 for four cells, two-sided Student’s t-test. Horizontal lines in a and c denote mean values. d, Immunofluorescence and breakdown of OPN-positive KI67-positive cells based on CK19 expression in P54 Alb-cre+/−Rbpjf/fHnf6f/f mouse liver (n = 4). Arrowheads indicate OPN-positive KI67-positive CK19-negative cells. e, Immunofluorescence of liver of >P120 Alb-cre+/−Rbpjf/fHnf6f/f and Rbpjf/fHnf6f/f mice after DDC diet feeding for 2 (n = 1 each), 4 (n = 3 each) and 6 (n = 1 each) weeks. f, Immunofluorescence of liver and breakdown of OPN-positive cells based on hepatocyte fate tracing in >P120 Alb-cre+/−Rbpjf/fHnf6f/fR26ZG+/+ (n = 4) and Rbpjf/fHnf6f/fR26ZG+/+ (n = 3) mice fed DDC diet for 5 weeks starting 1 week after hepatocyte fate tracing was induced. Data in d and f are mean ± s.e.m. Scale bars, 100 µm (b, e, f), 50 µm (d).
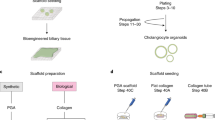
Extended Data Fig. 6 TGFβ signalling in hepatocyte transdifferentiation.
a, Ink visualization of biliary tree of P32 Alb-cre+/−Tgfbr2f/f mouse (n = 2). b, Immunofluorescence of P60 Alb-cre+/−Rbpjf/fHnf6f/f and Rbpjf/fHnf6f/f mouse livers (n = 2 each). Arrowheads indicate pSMAD3-positive HNF1-positive nuclei. c, Western blot with quantification of pSMAD3 in nuclear extracts from Alb-cre+/−Rbpjf/fHnf6f/f, Rbpjf/fHnf6f/f and Alb-cre+/−Rbpjf/fHnf6f/fTgfbr2f/f mouse livers (n = 2 each). Source data are shown in Supplementary Fig. 1. d–f, Experimental design (d) and results of analysis of the effect of TGFβ signalling on biliary differentiation of adult RBPJ- and HNF6-deficient hepatocytes cultured in 3D for the indicated number of days (d). e, Phase-contrast images of RPBJ- and HNF6-deficient hepatocyte spheroids embedded in collagen gels and cultured in the presence or absence of the TGFβ inhibitor SB-431542 (SB) for the indicated number of days. f, Relative expression levels of cholangiocyte and hepatocyte genes in freshly isolated hepatocytes and spheroids before and after embedding in collagen gels. Gene expression in the liver of a mouse fed choline-deficient ethionine-supplemented (CDE) diet was used as a positive control. Data are from three independent cultures per treatment in a representative experiment (n = 2). *P = 0.038 for Sox9 5 days, *P = 0.044 for Sox9 10 days, **P = 0.0034 for Krt19 5 days and **P = 0.0071 for Spp1 5 days, two-sided Welch’s t-test. g, Serum total bilirubin levels in P34–P53 Alb-cre+/−Rbpjf/fHnf6f/fTgfbr2f/f (n = 16) and Rbpjf/fHnf6f/fTgfbr2f/f (n = 7) mice. ****P = 0.000024, two-sided Welch’s t-test. h, Quantification of Sirius red staining in P58–P100 Rbpjf/fHnf6f/f mice after intravenous injection of AAV8-Eef1a1-caTgfbr1 at P20 (n = 4). Grey area represents the range of liver collagen in the indicated Rbpjf/fHnf6f/f mice from Extended Data Fig. 3f. Horizontal lines in c, g and h denote mean values. Data in f are mean ± s.e.m. Scale bars, 2 mm (a), 100 µm (e), 50 µm (b) and 10 µm (b, inset).
Supplementary information
Supplementary Information
This file contains Supplementary Tables 2–4 and Supplementary Figures 1–2
Supplementary Table 1
RNA-seq analysis. Excel file showing differentially expressed genes, including analysis of pathway and biological process enrichment, between normal peripheral cholangiocytes (pC; n = 3 mice), hepatocytes (H; n = 3 mice) and hepatocyte-derived peripheral cholangiocytes (HpC; n = 4 mice) as defined by fold-change > 3 for at least 1 of the 3 pairwise comparisons and FDR-corrected P < 0.05 (one-way ANOVA).
Video 1: HpBD connecting to a preexisting hBD
3D projection and surface rendering of a confocal z-stack from a hepatocyte-fate-traced P120 Alb-cre+/-Rbpjf/fHnf6f/fR26ZG+/+ mouse liver (n = 2) showing hepatocyte fate tracing (GFP, green) and labelling of both peripheral and hilar cholangiocytes (wsCK, red) and specifically hilar cholangiocytes (DBA, blue).
Video 2: Clones of hepatocyte-derived peripheral cholangiocytes
3D projection of a confocal z-stack from a P150 Alb-cre+/-Rbpjf/fHnf6f/fR26R-Confetti+/- mouse liver (n = 4) showing clonally labeled (yellow, cyan and red) peripheral cholangiocytes (wsCK, purple) and hepatocytes.
Rights and permissions
About this article
Cite this article
Schaub, J.R., Huppert, K.A., Kurial, S.N.T. et al. De novo formation of the biliary system by TGFβ-mediated hepatocyte transdifferentiation. Nature 557, 247–251 (2018). https://doi.org/10.1038/s41586-018-0075-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0075-5
This article is cited by
-
A MTA2-SATB2 chromatin complex restrains colonic plasticity toward small intestine by retaining HNF4A at colonic chromatin
Nature Communications (2024)
-
Cellular reprogramming in vivo initiated by SOX4 pioneer factor activity
Nature Communications (2024)
-
A spatiotemporal atlas of cholestatic injury and repair in mice
Nature Genetics (2024)
-
Current view of liver cancer cell-of-origin and proposed mechanisms precluding its proper determination
Cancer Cell International (2023)
-
Atypical cholangiocytes derived from hepatocyte-cholangiocyte transdifferentiation mediated by COX-2: a kind of misguided liver regeneration
Inflammation and Regeneration (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.