Abstract
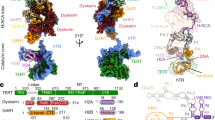
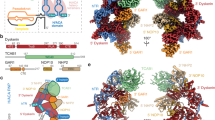
The enzyme telomerase adds telomeric repeats to chromosome ends to balance the loss of telomeres during genome replication. Telomerase regulation has been implicated in cancer, other human diseases, and ageing, but progress towards clinical manipulation of telomerase has been hampered by the lack of structural data. Here we present the cryo-electron microscopy structure of the substrate-bound human telomerase holoenzyme at subnanometre resolution, showing two flexibly RNA-tethered lobes: the catalytic core with telomerase reverse transcriptase (TERT) and conserved motifs of telomerase RNA (hTR), and an H/ACA ribonucleoprotein (RNP). In the catalytic core, RNA encircles TERT, adopting a well-ordered tertiary structure with surprisingly limited protein–RNA interactions. The H/ACA RNP lobe comprises two sets of heterotetrameric H/ACA proteins and one Cajal body protein, TCAB1, representing a pioneering structure of a large eukaryotic family of ribosome and spliceosome biogenesis factors. Our findings provide a structural framework for understanding human telomerase disease mutations and represent an important step towards telomerase-related clinical therapeutics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Arnoult, N. & Karlseder, J. Complex interactions between the DNA-damage response and mammalian telomeres. Nat. Struct. Mol. Biol. 22, 859–866 (2015).
Doksani, Y. & de Lange, T. The role of double-strand break repair pathways at functional and dysfunctional telomeres. Cold Spring Harb. Perspect. Biol. 6, a016576 (2014).
Levy, M. Z., Allsopp, R. C., Futcher, A. B., Greider, C. W. & Harley, C. B. Telomere end-replication problem and cell aging. J. Mol. Biol. 225, 951–960 (1992).
Holohan, B., Wright, W. E. & Shay, J. W. Cell biology of disease: telomeropathies: an emerging spectrum disorder. J. Cell Biol. 205, 289–299 (2014).
Wegman-Ostrosky, T. & Savage, S. A. The genomics of inherited bone marrow failure: from mechanism to the clinic. Br. J. Haematol. 177, 526–542 (2017).
Blackburn, E. H. & Collins, K. Telomerase: an RNP enzyme synthesizes DNA. Cold Spring Harb. Perspect. Biol. 3, a003558 (2011).
Shay, J. W. Role of telomeres and telomerase in aging and cancer. Cancer Discov. 6, 584–593 (2016).
Fu, D. & Collins, K. Distinct biogenesis pathways for human telomerase RNA and H/ACA small nucleolar RNAs. Mol. Cell 11, 1361–1372 (2003).
Podlevsky, J. D. & Chen, J. J. Evolutionary perspectives of telomerase RNA structure and function. RNA Biol. 13, 720–732 (2016).
Chan, H., Wang, Y. & Feigon, J. Progress in human and Tetrahymena telomerase structure determination. Annu. Rev. Biophys. 46, 199–225 (2017).
Wu, R. A., Upton, H. E., Vogan, J. M. & Collins, K. Telomerase mechanism of telomere synthesis. Annu. Rev. Biochem. 86, 439–460 (2017).
Gillis, A. J., Schuller, A. P. & Skordalakes, E. Structure of the Tribolium castaneum telomerase catalytic subunit TERT. Nature 455, 633–637 (2008).
Mitchell, M., Gillis, A., Futahashi, M., Fujiwara, H. & Skordalakes, E. Structural basis for telomerase catalytic subunit TERT binding to RNA template and telomeric DNA. Nat. Struct. Mol. Biol. 17, 513–518 (2010).
Robart, A. R. & Collins, K. Human telomerase domain interactions capture DNA for TEN domain-dependent processive elongation. Mol. Cell 42, 308–318 (2011).
Wu, R. A., Dagdas, Y. S., Yilmaz, S. T., Yildiz, A. & Collins, K. Single-molecule imaging of telomerase reverse transcriptase in human telomerase holoenzyme and minimal RNP complexes. eLife 4, 10.7554 (2015).
Wenz, C. et al. Human telomerase contains two cooperating telomerase RNA molecules. EMBO J. 20, 3526–3534 (2001).
Sauerwald, A. et al. Structure of active dimeric human telomerase. Nat. Struct. Mol. Biol. 20, 454–460 (2013).
Egan, E. D. & Collins, K. Specificity and stoichiometry of subunit interactions in the human telomerase holoenzyme assembled in vivo. Mol. Cell. Biol. 30, 2775–2786 (2010).
Jiang, J. et al. Structure of Tetrahymena telomerase reveals previously unknown subunits, functions, and interactions. Science 350, aab4070 (2015).
MacNeil, D. E., Bensoussan, H. J. & Autexier, C. Telomerase regulation from beginning to the end. Genes (Basel) 7, (64 (2016).
Hamma, T. & Ferré-D’Amaré, A. R. The box H/ACA ribonucleoprotein complex: interplay of RNA and protein structures in post-transcriptional RNA modification. J. Biol. Chem. 285, 805–809 (2010).
Yu, Y.-T. & Meier, U. T. RNA-guided isomerization of uridine to pseudouridine–pseudouridylation. RNA Biol. 11, 1483–1494 (2014).
Cohen, S. B. et al. Protein composition of catalytically active human telomerase from immortal cells. Science 315, 1850–1853 (2007).
Li, L. & Ye, K. Crystal structure of an H/ACA box ribonucleoprotein particle. Nature 443, 302–307 (2006).
Fu, D. & Collins, K. Purification of human telomerase complexes identifies factors involved in telomerase biogenesis and telomere length regulation. Mol. Cell 28, 773–785 (2007).
Cristofari, G. et al. Human telomerase RNA accumulation in Cajal bodies facilitates telomerase recruitment to telomeres and telomere elongation. Mol. Cell 27, 882–889 (2007).
Wu, R. A. & Collins, K. Human telomerase specialization for repeat synthesis by unique handling of primer-template duplex. EMBO J. 33, 921–935 (2014).
Vogan, J. M. et al. Minimized human telomerase maintains telomeres and resolves endogenous roles of H/ACA proteins, TCAB1, and Cajal bodies. eLife 5, 10.7554 (2016).
Bai, X.-C., Rajendra, E., Yang, G., Shi, Y. & Scheres, S. H. W. Sampling the conformational space of the catalytic subunit of human γ-secretase. eLife 4, e11182 (2015).
Jiang, J. et al. The architecture of Tetrahymena telomerase holoenzyme. Nature 496, 187–192 (2013).
Bajon, E., Laterreur, N. & Wellinger, R. J. A single templating RNA in yeast telomerase. Cell Reports 12, 441–448 (2015).
Alves, D. et al. Single-molecule analysis of human telomerase monomer. Nat. Chem. Biol. 4, 287–289 (2008).
Huang, J. et al. Structural basis for protein-RNA recognition in telomerase. Nat. Struct. Mol. Biol. 21, 507–512 (2014).
Jacobs, S. A., Podell, E. R. & Cech, T. R. Crystal structure of the essential N-terminal domain of telomerase reverse transcriptase. Nat. Struct. Mol. Biol. 13, 218–225 (2006).
Petrova, O. A. et al. Structure and function of the N-terminal domain of the yeast telomerase reverse transcriptase. Nucleic Acids Res. 46, 1525–1540 (2018).
Kim, N.-K. et al. Solution structure and dynamics of the wild-type pseudoknot of human telomerase RNA. J. Mol. Biol. 384, 1249–1261 (2008).
Zhang, Q., Kim, N.-K. & Feigon, J. Architecture of human telomerase RNA. Proc. Natl Acad. Sci. USA 108, 20325–20332 (2011).
Wang, Y., Yesselman, J. D., Zhang, Q., Kang, M. & Feigon, J. Structural conservation in the template/pseudoknot domain of vertebrate telomerase RNA from teleost fish to human. Proc. Natl Acad. Sci. USA 113, E5125–E5134 (2016).
Theimer, C. A. et al. Structural and functional characterization of human telomerase RNA processing and cajal body localization signals. Mol. Cell 27, 869–881 (2007).
Cash, D. D. & Feigon, J. Structure and folding of the Tetrahymena telomerase RNA pseudoknot. Nucleic Acids Res. 45, 482–495 (2017).
Jansson, L. I. et al. Structural basis of template-boundary definition in Tetrahymena telomerase. Nat. Struct. Mol. Biol. 22, 883–888 (2015).
Mitchell, J. R. & Collins, K. Human telomerase activation requires two independent interactions between telomerase RNA and telomerase reverse transcriptase. Mol. Cell 6, 361–371 (2000).
Chen, J.-L., Opperman, K. K. & Greider, C. W. A critical stem-loop structure in the CR4-CR5 domain of mammalian telomerase RNA. Nucleic Acids Res. 30, 592–597 (2002).
Bley, C. J. et al. RNA–protein binding interface in the telomerase ribonucleoprotein. Proc. Natl Acad. Sci. USA 108, 20333–20338 (2011).
Chen, J.-L., Blasco, M. A. & Greider, C. W. Secondary structure of vertebrate telomerase RNA. Cell 100, 503–514 (2000).
Egan, E. D. & Collins, K. An enhanced H/ACA RNP assembly mechanism for human telomerase RNA. Mol. Cell. Biol. 32, 2428–2439 (2012).
Kittur, N., Darzacq, X., Roy, S., Singer, R. H. & Meier, U. T. Dynamic association and localization of human H/ACA RNP proteins. RNA 12, 2057–2062 (2006).
Venteicher, A. S. et al. A human telomerase holoenzyme protein required for Cajal body localization and telomere synthesis. Science 323, 644–648 (2009).
Tycowski, K. T., Shu, M. D., Kukoyi, A. & Steitz, J. A. A conserved WD40 protein binds the Cajal body localization signal of scaRNP particles. Mol. Cell 34, 47–57 (2009).
Sarek, G., Marzec, P., Margalef, P. & Boulton, S. J. Molecular basis of telomere dysfunction in human genetic diseases. Nat. Struct. Mol. Biol. 22, 867–874 (2015).
Reeves, P. J., Callewaert, N., Contreras, R. & Khorana, H. G. Structure and function in rhodopsin: high-level expression of rhodopsin with restricted and homogeneous N-glycosylation by a tetracycline-inducible N-acetylglucosaminyltransferase I-negative HEK293S stable mammalian cell line. Proc. Natl Acad. Sci. USA 99, 13419–13424 (2002).
Durocher, Y., Perret, S. & Kamen, A. High-level and high-throughput recombinant protein production by transient transfection of suspension-growing human 293-EBNA1 cells. Nucleic Acids Res. 30, E9 (2002).
Kim, N. W. et al. Specific association of human telomerase activity with immortal cells and cancer. Science 266, 2011–2015 (1994).
Schirmer, E. C., Yates, J. R. III & Gerace, L. MudPIT: A powerful proteomics tool for discovery. Discov. Med. 3, 38–39 (2003).
Sexton, A. N. et al. Genetic and molecular identification of three human TPP1 functions in telomerase action: recruitment, activation, and homeostasis set point regulation. Genes Dev. 28, 1885–1899 (2014).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Scheres, S. H. W. Semi-automated selection of cryo-EM particles in RELION-1.3. J. Struct. Biol. 189, 114–122 (2015).
Voss, N. R., Yoshioka, C. K., Radermacher, M., Potter, C. S. & Carragher, B. DoG Picker and TiltPicker: software tools to facilitate particle selection in single particle electron microscopy. J. Struct. Biol. 166, 205–213 (2009).
Lander, G. C. et al. Appion: an integrated, database-driven pipeline to facilitate EM image processing. J. Struct. Biol. 166, 95–102 (2009).
Nguyen, T. H. D. et al. The architecture of the spliceosomal U4/U6.U5 tri-snRNP. Nature 523, 47–52 (2015).
Tivol, W. F., Briegel, A. & Jensen, G. J. An improved cryogen for plunge freezing. Microsc. Microanal. 14, 375–379 (2008).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Kimanius, D., Forsberg, B. O., Scheres, S. H. W. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, 18722 (2016).
Zheng, S. Q., Palovcak, E., Armache, J.-P., Cheng, Y. & Agard, D. A. MotionCor2: anisotropic correction of beam-induced motion for improved single particle electron cryo-microscopy. Nat. Methods 14, 331–332 (2017).
Biyani, N. et al. Focus: The interface between data collection and data processing in cryo-EM. J. Struct. Biol. 198, 124–133 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Scheres, S. H. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Scheres, S. H. W., Núñez-Ramírez, R., Sorzano, C. O. S., Carazo, J. M. & Marabini, R. Image processing for electron microscopy single-particle analysis using XMIPP. Nat. Protocols 3, 977–990 (2008).
Goddard, T. D., Huang, C. C. & Ferrin, T. E. Visualizing density maps with UCSF Chimera. J. Struct. Biol. 157, 281–287 (2007).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protocols 10, 845–858 (2015).
Dodonova, S. O. et al. Vesicular transport. A structure of the COPI coat and the role of coat proteins in membrane vesicle assembly. Science 349, 195–198 (2015).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Acknowledgements
We thank P. Grob, J. Fang, A. Chintangal, P. Tobias and H. Upton for technical support; X. Zhang and J. Vogan for establishing the suspension cell transfection protocol used in early stages of this work and sharing cell lines for in vivo studies and CAB-box mutant hTR; L. Kohlstaedt and QB3 mass spectrometry facility for analysis of purified telomerase; Y. Wang, J. Jiang and J. Feigon for sharing the coordinates of the Tetrahymena catalytic core and the modelled human t/PK; E. Rodina for sharing the Hansenula polymorpha TEN coordinates before publication; and H. Upton and A. Deshpande for comments on the manuscript. We thank the National Energy Research Scientific Computing Center supported by the Office of Science of the US Department of Energy for providing computing resources under contract number DE-AC02-05CH11231. This work was funded by N.I.H. grant GM054198 to K.C. T.H.D.N. is a Fellow of the University of California, Berkeley Miller Institute for Basic Research in Science. B.J.G. was supported by fellowships from the Swiss National Science Foundation (projects P300PA_160983, P300PA_174355). E.N. is a Howard Hughes Medical Investigator.
Reviewer information
Nature thanks S. Scheres and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
K.C. and E.N. directed the study. T.H.D.N. developed the telomerase purification procedure and performed in vitro biochemical assays, negative-stain and cryo-EM specimen preparation, data collection, data processing and model fitting. J.T. performed in vivo experiments and assisted T.H.D.N with cell culture and biochemistry. R.A.W guided T.H.D.N with telomerase biochemistry and purification at the initial stage of the project. B.J.G. and D.T. helped T.H.D.N with data collection. T.H.D.N, K.C. and E.N. wrote the paper with contributions from all other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Protein identification by immunoblotting, enriching active telomerase, substrate pre-binding, and comparison of intact, ΔTCAB1, and TERT–hTRmin RNPs.
a, Immunoblotting of TERT, TCAB1, dyskerin, GAR1, NHP2 and NOP10 in telomerase purified after CHAPS lysis protocol as shown in Fig. 1b. We used primary antibodies against each protein, except ZZ-SS-TERT, for which we used rabbit IgG. Owing to the wide range of molecular weights of the proteins in our sample, TERT, TCAB1, dyskerin and GAR1 were detected in one blot, while NHP2 and NOP10 were detected in a separate blot. The use of the same sample to probe all proteins was performed only once, but TERT, dyskerin and TCAB1 were also probed individually twice. b, Silver-stained SDS–PAGE gel of purified telomerase fractions obtained from adherent cells lysed using the hypotonic lysis method, which enriches active telomerase. This experiment was repeated more than five times with similar results. c, Direct primer-extension assays of the purified telomerase fractions shown in b, confirming that E1 is no longer inactive (left), and of the substrate-bound purified telomerase fractions with additional DNA substrate omitted from the assays (right). The activity observed confirmed that purified telomerase contains the DNA substrate. The activity assays with substrate added were repeated over five times and the activity assays with substrate pre-bound were repeated twice. All repeats showed similar results. d, Silver-stained SDS–PAGE gel of purified intact and ΔTCAB1 telomerase and TERT–hTRmin telomerase prepared for subunit assignments. This experiment was done only once to provide a direct comparison between these different purified telomerase complexes. e–g, Negative-stained 2D class averages of intact and ΔTCAB1 telomerase and TERT–hTRmin, respectively. h, Comparison of representative 2D class averages of intact and ΔTCAB1 telomerase and TERT–hTRmin showing the inferred localization of TCAB1 and TERT. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 2 Cellular function of the tagged TERT used for structural analysis.
a, Western blot detection of ZZ-SS-TERT in TERT knockout (KO) cells rescued by untagged TERT or ZZ-SS-TERT expression. Whole-cell extracts were probed using Strep antibody. HCT116 is the parental cell line. Lysate prepared from HEK 293T cells transiently transfected with ZZ-SS-TERT and hTR was used as a positive control (Ctrl). Tubulin was detected as a loading control. This experiment was performed only once to confirm the success of ZZ-SS-TERT incorporation into the HCT116 TERT KO cells. b, Telomeric restriction fragment analysis of HCT116 parental cells, TERT KO cells (before senescence), and TERT KO cells rescued with untagged or ZZ-SS-TERT transgene. Transgene-expressing cells were sampled at 31, 62 and 98 days post-transfection with transgene vectors. This experiment was performed twice with similar results. c, TRAP assay detection of telomerase activity in HCT116 parental cells, TERT KO cells, and TERT KO cells rescued with untagged TERT or ZZ-SS-TERT transgene. Whole-cell extracts were normalized by total protein concentration and assayed at 100, 30 or 10 ng of total protein per reaction. IC, internal control. This experiment was repeated four times with similar results. d, Quantification of Q-TRAP assay detection of telomerase activity in HCT parental cells, TERT KO cells and TERT KO cells rescued with untagged TERT or ZZ-SS-TERT transgene. Error bars were calculated by taking the s.d. of the average ΔCt from four time points. Data points were shown as overlays. e, Direct primer-extension assay of telomerase after template-complementary oligonucleotide purification from extracts of TERT KO cells rescued by untagged TERT or ZZ-SS-TERT transgene. Assays were performed on clarified cell lysate (crude), flow-through (O-FT) and elution (OE) using equivalent amounts of cell extract. This experiment was performed only once to re-confirm the results in c. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 3 Image processing procedures.
a, Representative raw micrograph. We collected a total of 11,654 micrographs for this study. b, Representative 2D class averages obtained from reference-free 2D classification. c, Data processing strategy used in this study.
Extended Data Fig. 4 Resolution estimation and analysis of the flexibility of the complex.
a, Representative 2D class averages obtained from 2D classification without alignment of particles that were aligned on either the catalytic core or the H/ACA lobe. For both cases, the other lobe adopts a wide range of conformations, as illustrated by the blurriness of the density. b, FSC curves for the overall map and the maps of the catalytic core and H/ACA lobe resulting from focused classification with signal subtraction and gold-standard refinement. c, Model versus map FSC curves for the catalytic core and the H/ACA RNP. We fitted only homology models as rigid bodies into the map and did not perform model coordinate refinement owing to the limited resolutions of the maps. Therefore, we used a lower FSC threshold of 0.25 for resolution estimates. d, e, Local resolution for the catalytic lobe (d) and the H/ACA lobe (e) estimated by RELION 2.064. Most of the catalytic core is resolved at 6–8 Å while most of the H/ACA lobe is resolved at 7–9 Å. f, Front (left) and back (right) views of the reconstruction showing modelled (grey) and unmodelled (gold) density. Most of the unmodelled density corresponds to single-stranded RNA regions or RNA bulges, and human protein extensions that cannot be built de novo at this resolution.
Extended Data Fig. 5 Fittings of proteins and RNA into the cryo-EM map.
a–d, Domains of TERT. a, The TEN domain from Tetrahymena34 (PDB 2B2A). b, The truncated medaka TRBD domain33 (PDB 4O26). c, d, The RT and CTE domains from Tribolium13 (PDB 3KYL). e, Front (top) and back (bottom) views of the 5′ hairpin set of H/ACA proteins (dyskerin, red; GAR1, cyan; NOP10, wheat; NHP2, pink) bound to P4 stem (dark blue) fit by the archaeal H/ACA RNP24 (PDB 2HVY). f, Front (top) and back (bottom) views of the 3′ hairpin set of H/ACA proteins using the same model and colour scheme as e. g, Homology model of TCAB1 WD40 domain. h, Front (top) and bottom (bottom) views of hTR in the catalytic core. i, hTR in the H/ACA lobe.
Extended Data Fig. 6 Sequence alignment of TERT with secondary structure assignments based on known structures.
a, Sequence alignment of Tetrahymena and human TEN domains. The secondary structure assignments of the Tetrahymena TEN domain34 (PDB 2B2A) are shown above the aligned sequences. Regions removed before fitting are indicated with dashed lines below the sequences. b, Sequence alignment of the Tribolium, human and Tetrahymena TERT, with the latter two N-terminally truncated to match Tribolium. Secondary structure assignments of the Tribolium TERT are shown on top, with conserved motifs labelled in blue. Throughout the figure, the η symbol refers to a 310-helix. Strict β-turns and strict α-turns are displayed as TT and TTT. The three catalytic aspartic acids are indicated with black arrowheads. ESpript was used to generate this figure76.
Extended Data Fig. 7 Selected protein–protein and protein–RNA interactions in telomerase holoenzyme and comparisons between human and Tetrahymena TERT.
a, Interactions between the RT and CTE domains of TERT and the substrate–template duplex. The RT domain is divided into two subdomains, the palm (green) and fingers (orange), that are commonly observed in retroviral reverse transcriptases. The CTE (cyan) is the putative thumb. The IFD insertion that is missing in the Tribolium TERT is indicated. b, Region of the cryo-EM reconstruction shown in a. Unassigned density close to the IFD insertion is highlighted in magenta. c, Cryo-EM density of the TEN domain in the same view as that in Fig. 4b. Connecting density is observed between the template region and the P2a.1 stem. d, Map of the CR4/5 three-way junction (wheat) and the nearby TERT domains highlighting the position of the P6.1 loop near the interface of the CTE (cyan) and TRBD (blue) domains of TERT. This loop was not ordered in medaka CR4/5 bound to the TRBD alone33. e, Comparison of the Tribolium (left) and medaka (right) TRBD with the medaka CR4/5 domain of hTR13,33. Extensions of the medaka TRBD that did not fit the map were truncated for visualization. f, Cryo-EM map with H/ACA components fitted. g, h, Detailed views of regions boxed in f show TCAB1 interactions with dyskerin, GAR1 and the P8 stem-loop (g), and interactions between the two dyskerin molecules (h), where a cluster of DC mutations are found (Fig. 5d). i, Comparison of the human and Tetrahymena TERT superposed on the RT domain. Domains of human TERT are coloured as in Fig. 1a, while Tetrahymena TERT is coloured grey. The bound human and Tetrahymena templates are coloured dark and light red, respectively. j, Comparison of human and Tetrahymena19 catalytic cores fitted into the corresponding cryo-EM maps. Domains of TERT were coloured as in Fig. 1a and TER is coloured yellow. We used the catalytic core and H/ACA lobe densities resulting from our focused classification/refinement for the human telomerase and the overall 9.4 Å Tetrahymena telomerase map (EMD-6442).
Extended Data Fig. 8 Sequence alignments of H/ACA proteins with secondary structure assignments based on known structures.
a–d, Sequence alignments of Pyrococcus furiosus (archaeal) and human Cbf5/dyskerin (a), GAR1 (b), NOP10 (c), and L7Ae/NHP2 (d). Secondary structure assignments displayed on the top are from the archaeal H/ACA RNP structure24 (PDB 2HVY). The η symbol refers to a 310-helix. Strict β-turns and strict α-turns are displayed as TT and TTT, respectively. Known human dyskeratosis congenita and Hoyeraal–Hreidarsson disease mutations50 in H/ACA proteins are indicated with arrowheads. Blue arrowheads indicate residues that can be mapped onto the archaeal structure and black arrowheads indicate residues that were not mapped. ESpript was used to generate this figure76.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1, which shows source images for all data obtained by gel electrophoresis in indicated figures.
Supplementary Data
This file contains Pymol session containing all models used for fitting in this study
Video 1: Cryo-EM structure of human telomerase holoenzyme
The video first shows the cryo-EM densities of the H/ACA lobe and the catalytic core of the human telomerase at 8.2 Å and 7.7 Å, respectively, and fitting of the protein and RNA subunits into the cryo-EM densities. It is followed by the close-up views of the catalytic core and the H/ACA lobe highlighting how the subunits are assembled in our human telomerase structure
Rights and permissions
About this article
Cite this article
Nguyen, T.H.D., Tam, J., Wu, R.A. et al. Cryo-EM structure of substrate-bound human telomerase holoenzyme. Nature 557, 190–195 (2018). https://doi.org/10.1038/s41586-018-0062-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0062-x
This article is cited by
-
The regulations of telomerase reverse transcriptase (TERT) in cancer
Cell Death & Disease (2024)
-
Favipiravir, an antiviral drug, in combination with tamoxifen exerts synergistic effect in tamoxifen-resistant breast cancer cells via hTERT inhibition
Scientific Reports (2024)
-
2.7 Å cryo-EM structure of human telomerase H/ACA ribonucleoprotein
Nature Communications (2024)
-
Methods that shaped telomerase research
Biogerontology (2024)
-
A CRISPR base editing approach for the functional assessment of telomere biology disorder-related genes in human health and aging
Biogerontology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.